Published online Dec 6, 2021. doi: 10.12998/wjcc.v9.i34.10400
Peer-review started: February 7, 2021
First decision: March 14, 2021
Revised: March 26, 2021
Accepted: August 5, 2021
Article in press: August 5, 2021
Published online: December 6, 2021
Processing time: 296 Days and 2.2 Hours
The incidence of gastrointestinal cancers has increased significantly over the past decade and gastrointestinal malignancies now rank among the leading causes of mortality globally. Although newer therapeutic strategies such as targeted therapies have greatly improved patient outcomes, their clinical success is limited by drug resistance, treatment failure and recurrence of metastatic disease. Therefore, there is an urgent need for further research identifying accurate and reliable biomarkers for precise treatment strategies. Circular RNAs (circRNAs) exhibit a covalently closed structure, high stability and biological conservation, and their expression is associated with the occurrence and development of gastrointestinal tumors. Moreover, circRNAs may significantly influence drug resistance of gastrointestinal cancers. In this article, we review the role of circRNAs in the occurrence and development of gastrointestinal cancer, their association with drug resistance, and potential application for early diagnosis, treatment and prognosis in gastrointestinal malignancies. Furthermore, we summarize characteristics of circRNA, including mechanism of formation and biological effects via mRNA sponging, chromatin replication, gene regulation, translational modification, signal transduction, and damage repair. Finally, we discuss whether circRNA-related noninvasive testing may be clinically provided in the future. This review provides new insights for the future development of diagnostics and therapeutics based on circRNAs in gastrointestinal tumors.
Core Tip: Circular RNAs (circRNAs) were first discovered in 1976, and then more related biological functions were discovered. For example, as a miRNA sponge, circRNA participates in tumor-related processes, translation modification, signal transduction, and DNA damage repair. Gastrointestinal tumors have a high incidence, and some patients also show drug resistance during treatment. On this basis, we reviewed the relationship between circRNA and gastrointestinal tumors and the relevance of drug resistance, hoping to conduct more in-depth research on circRNA. At the same time, it is hoped that circRNA will provide new directions and strategies for future cancer treatment.
- Citation: Xi SJ, Cai WQ, Wang QQ, Peng XC. Role of circular RNAs in gastrointestinal tumors and drug resistance. World J Clin Cases 2021; 9(34): 10400-10417
- URL: https://www.wjgnet.com/2307-8960/full/v9/i34/10400.htm
- DOI: https://dx.doi.org/10.12998/wjcc.v9.i34.10400
The incidence of gastrointestinal tumors has increased significantly in recent years[1]. Gastrointestinal tumors, one of the most common cancer types, are increasingly treated with novel treatment strategies including targeted therapies and immunotherapies[2]. In clinical practice, pemetrexed, paclitaxel, cisplatin, sorafenib, and 5-fluorouracil (5-FU), among other drugs, are commonly used therapeutics, but the frequent emergence of drug resistance renders treatment extremely challenging. Radiotherapy and chemotherapy resistance are typically caused by both tumor-cell intrinsic and extrinsic mechanisms, including epithelial–mesenchymal transition (EMT), alterations in tumor microenvironment, and hypoxia signaling[3].
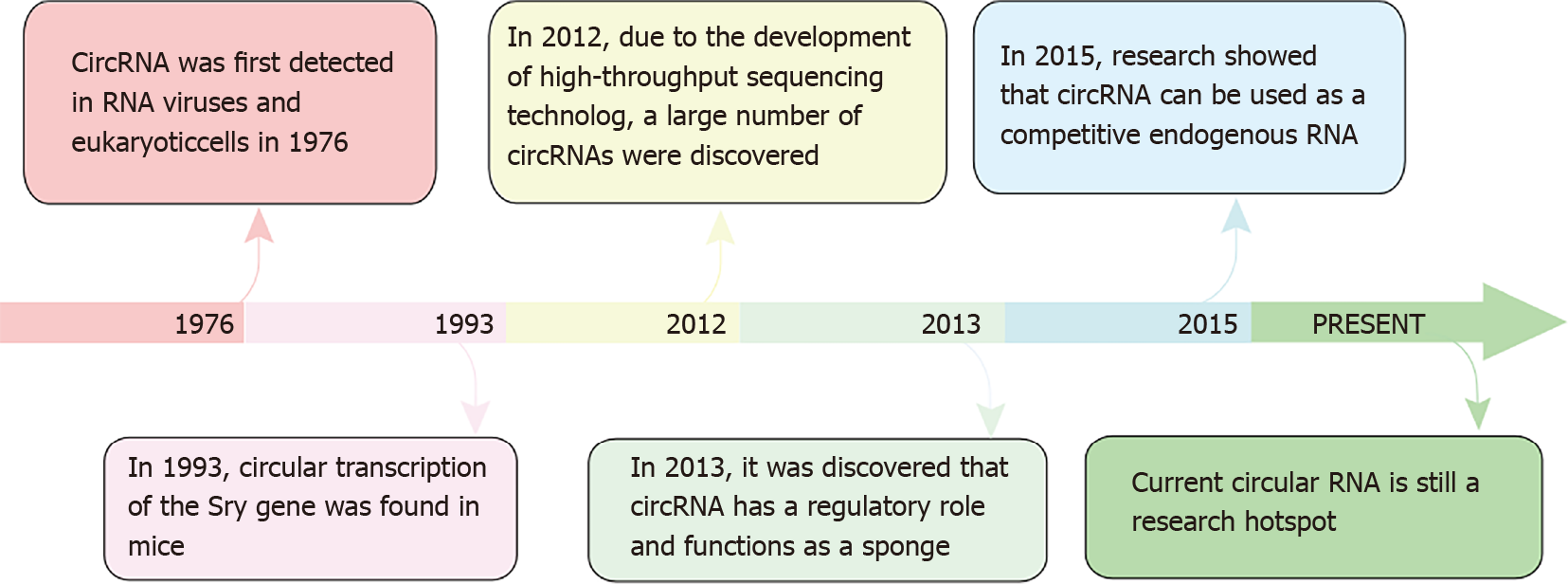
Circular RNAs (circRNAs) have been found to be associated with the occurrence and development of a variety of cancers. They were first detected in 1976 in both RNA viruses and eukaryotic cells[4], but were considered meaningless and an erroneous splicing structure at the time[5]. However, in 1993, Capel et al[6] found that circRNAs accounted for > 90% of adult testicular SRY transcription. In 2012, Salzman et al[7] reported circRNA transcription of hundreds of human genes. Work by Memczak et al[8] in 2013 revealed that circRNA exhibit regulatory effects, and later work demon
Figure 1 highlights the timeline of breakthrough research on circRNAs and their potential application in the field of oncology.
circRNAs have complex biological functions in gastrointestinal tumors. Here, we discuss the sponging effect of circRNAs in tumors as well as their effects on chromatin replication, gene regulation, translational modification, signal transduction, and damage repair.
circRNAs are circular RNA molecules formed by covalent sealing of single-stranded RNA molecules. The majority of circRNAs is located in the cytoplasm. circRNAs are highly stable and biologically conserved[10]. They have an uncapped 5' end and no polyA structure at the 3' end[11]. circRNAs usually occur during the splicing process after transcription[12], and the majority of circRNAs can be degraded by RNase L endonuclease while they cannot be easily degraded by exonuclease[13]. circRNAs are considered functional noncoding transcripts that can produce functional peptides or proteins[14]: studies have shown that many circRNAs are translated and therefore circRNAs cannot be defined as entirely noncoding RNAs. For example, the 220-nucleotide CCC RNA in the wheat germ extraction system provided the first proof of natural circRNA translation in 2014[15]. Moreover, some circRNAs have an N6-methyladenosine (m6A) modification that promotes the translation of circRNAs and plays a role in either inhibition or promotion of tumor progression[16,17]. circRNAs are suggested to be involved in the progression and drug resistance of a variety of cancers, although further studies need to be conducted to confirm the exact mechanisms of action[18]. By understanding the properties and functions of circRNAs, we can further explore the mechanisms of their involvement in gastrointestinal tumors as well as potential future applications for diagnosis and treatment.
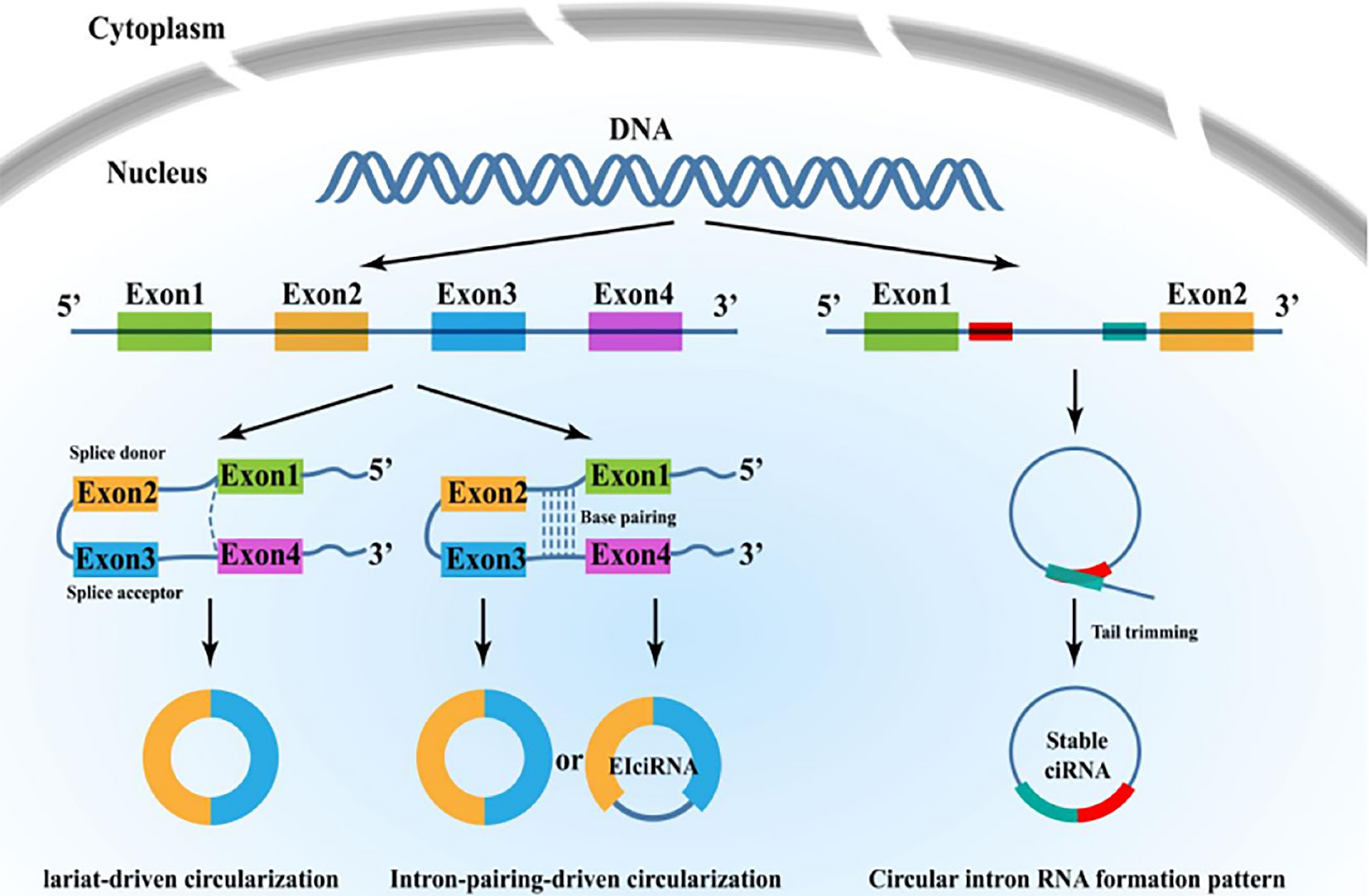
In recent years, there have been several studies investigating the formation of circRNAs. circRNAs are divided into four types: exonic, intronic, exon-intron, and intergenic[19]. At present, the mechanisms of circRNA processing and maturation are still largely unclear[20], but recent studies suggest that 80% of circRNAs are exonic circRNAs (ecircRNA), which have two key formation mechanisms: lasso-driven circularization (exon skipping) and intron pair-driven circularization (direct reverse splicing)[21] (Figure 2). mRNA intron lassos can evade degradation during splicing, forming circRNA with 2′, 5′phosphodiester bonds, and the same sequence can form different circRNAs via different types of splicing. Not all exons can be spliced to form circRNAs, however the more spliced an RNA is, the less fully processed mRNA exists, for example in the rat P4502C24 gene[22].
There are also other theories that state that circRNAs can be formed by RNA-binding proteins (RBPs) or flanking introns that interact with each intron through RNA–RNA interaction to promote the combination of both ends of the head and tail, ultimately leading to the formation of intronic circRNAs (ciRNAs) or exon-intron circRNAs (EIciRNAs). More research investigating the formation mechanisms of circRNA in-depth will thus be required in the future.
circRNA sponge effects: circRNAs are associated with various conditions and diseases including aging, defect insulin secretion, Alzheimer’s disease, gastrointestinal tumors, as well as cardiovascular and cerebrovascular diseases, to name a few[23-26]. Generally, circRNAs bind to mRNAs by base complementary pairing, thereby potentially participating in the tumor related processes[27].
CDR1as is the product of cerebellar degeneration-related protein 1 (CDR1), also known as ciRS-7[28]. Knockdown of CDR1as can promote the degradation of miR-7-targeted mRNA and can act as a miR-7 sponge[29,30]. It has been reported that in addition to ciRS-7, circPVT1 may also promote cell proliferation by acting as a sponge for miR-125 family members[31]. circRNAs can also act as sponges for RBPs or miR-138 family members[32,33].
circRNAs exhibit a sponging activity for specific miRNAs, thereby inhibiting miRNA-based inhibition of gene expression.
Impact of circRNAs on chromatin replication: Increasing numbers of studies have shown that circRNAs are involved in chromatin replication. Chromatin replication includes DNA and nucleosome replication, during which nucleosomes interact with nucleoproteins that are important for protein recruitment, DNA replication, transcription, and damage repair[34]. DNA replication occurs in the nucleus[35]. During chromatin replication, certain proteins may be recruited which help to organize chromatin regions and regulate gene expression. Chromatin immunoprecipitation (ChIP) sequencing technology has detected the presence of nuclear circRNAs during chromatin replication, which indicates that circRNAs may play a role in this process[36]. Moreover, ChIP studies have also found that overexpression of circRNAs inhibits tumor proliferation in bladder cancer[37]. Clarifying the role of circRNAs in chromatin replication may therefore provide future insights for inhibiting tumor proliferation.
CircRNA regulation of gene transcription: circRNAs regulate gene transcription function and play an important role in gastrointestinal tumors. Studies have shown that addition of m6A is a reversible transcriptomic modification found in many eukaryotic mRNAs[38]. m6A modification of circRNAs aids their translation to proteins[39,40]. For example, circ-FBXW7 can be translated into the FBXW7-185aa protein after m6A modification[14]. Circular structures may even be translated into more proteins than linear structure in the ribosome[41]. circRNAs are transcribed by RNA polymerase (Pol) II and can be produced by spliceosomes, including heterogeneous nuclear ribonucleoproteins (hnRNPs)[42]. For example, when precipitating circEIF3J or circPAIP2, PolⅡ, U1A, U1c and U1 small nuclear RNAs are detected. Similarly, EIciRNAs can interact with Pol II, U1 hnRNP and parental gene promoters to enhance the transcription of parent genes, thus affecting protein translation[43]. The peptides encoded by circRNA can interfere with the metabolism or metastasis of tumors, thereby exerting an antitumor effect[44]. Nuclear circRNAs act as transcriptional or splicing regulators that interfere with gene expression and participate in selective splicing and transcription processes[45]. For instance, circITGA7 plays a negative regulatory role in the Ras pathway and inhibits RREB1 upregulation through the Ras pathway, which typically results in ITGA7 transcription[46]. circSHKBP1 has been shown to act as a miR-582-3p sponge and regulates the HUR/VEGF pathway in the promotion of the progression of gastric cancer[47].
To summarize, circRNAs play many biological roles and are involved in splicing, transcriptional or translational regulation, sequestering of miRNA, and protein binding. Although the mechanisms are largely unclear, circRNAs are likely to play a vital role in tumor development via these processes.
circRNA post-translational modifications (PTMs): Modifications can be regulated by the structure of chromatin, and it can also use ATP hydrolysis to generate energy to relocate the recombinase, and post-translational histone modifications, such as acetylation, phosphorylation, methylation and many other forms[48]. PTMs often occur after protein translation. The most common PTMs influenced by circRNAs are methylation and acetylation. In the tumor microenvironment, circRNAs can promote acetylation and kill tumor cells by regulation of gene expression. For example, circSETD3 can inhibit α-tubulin acetylation by upregulation of MAPER1 expression in nasopharyngeal carcinoma, thereby promoting the invasion and migration of tumor cells, confirming a potential for circRNAs as a biomarker[49]. In gastric cancer, circMRPS35 can inhibit the progression of tumors via the circMRPS35/KAT7/ foxo1/3a pathway, influencing acetylation via the histone acetyltransferase KAT7[50]. In liver cancer, circRNAs have been shown to modify methylation, with most circRNAs with methylation sites having only one methylation peak[51]. Moreover, circSOD2 increases the DNA methylation of the SOCS3 promoter, thereby playing a role in liver cancer tumors[52].
circRNAs and signal transduction: Signal transduction pathways often undergo abnormal changes in the occurrence and development of tumors. In osteosarcoma, circTCF25 has been shown to promote phosphorylation of signal transducers and produce carcinogenic effects[53]. In lung cancer, circ-ZKSCAN1/miR-330-5p/FAM83A can regulate the signal transduction pathway, and the reduction of circ_ZKSCAN1 can inhibit tumor cell migration and proliferation[54]. In breast cancer, hsa-circ-0061825 (circ-TFF1) may promote progression by affecting miR-326/TFF1 signaling[55]. circFMN2 may promote colorectal cancer progression via the miR-1182/hTERT signaling pathway[56]. circ0000190 has been shown inhibit proliferation and metastasis of gastric cancer cells by regulation of caspase-3 expression and inhibition of the miR-1252/PAK3 pathway[57]. Exploring diverse signaling pathways influenced by circRNA may provide potential therapeutic targets relating to tumor development and progression in the future.
circRNAs and damage repair: DNA breaks are characterized as single- or double strand [58]. In vivo repair mechanisms support the precise repair of single-strand breaks, while double-strand breaks often lead to cell death[59]. The occurrence of tumors is sometimes related to radiation, which causes DNA double-strand breaks. In noncoding RNA, the signal ionizing radiation response can be adjusted by targeting[60]. In breast cancer, circSMARCA5 can participate in the remodeling of damaged chromatin DNA[61]. Assessing the influence of circRNAs on DNA damage repair mechanisms in different cells may provide valuable information for DNA repair in the future.
Tumor development is a complicated pathological process. Molecules such as noncoding RNAs and cell-free DNA may be used as potential biomarkers but also therapeutic targets for benign and malignant diseases of the gastrointestinal tract[62]. The mechanism of anticancer drug resistance is multifaceted. Tumor growth, tumor microenvironment, immune system, and selective treatment pressure all affect tumor resistance[63]. Unfortunately, some patients develop drug resistance that requires exploration of other treatment directions. Therefore, drug resistance research is a key area of investigation. Several studies have suggested that circRNAs may play a key role not only as therapeutic targets for digestive system tumors but targeting them may also help to overcome drug resistance.
Owing to their special circular structure, circRNAs can be stably present in peripheral bodily fluids such as saliva and plasma, and can remain stable in serum for > 24 h[64]. Research has found that out of 11 circRNAs assessed, five were upregulated and promoted cancer, while six were downregulated and exhibited cancer-suppressing effects[65].
circRNA100876 is highly expressed in gastric cancer (GC) tissues and influences the miR-665/YAP1 signaling pathway. It can inhibit the proliferation, invasion and migration of GC cells by downregulation of the EMT pathway[66]. Conversely, studies have shown that circCUL3 can promote extracellular acidification rate, lactic acid production, and GC cell proliferation. This circRNA is highly expressed in GC tumor cells and has been shown to promote the expression of signal transducer and activator of transcription (STAT) 3 via sponging of miR-515-5p[67]. Similarly, circTMEM87A is also highly expressed in GC and can sponge miR-142-5p, regulating ULK1 expression and thus promoting GC proliferation and metastasis[68]. circRNA-0044516 is likewise upregulated in GC and acts as a miR-149 sponge. circRNA-0044516 silencing results in reduced Wnt1 and β-catenin protein levels and inhibits GC progression[69]. circ0000039 is also upregulated in GC, adsorbs miR-1292-5p, and upregulates DEK expression to promote GC proliferation and progression as an oncogene[70]. Circ-0004872 is significantly downregulated in GC tumor cells and has a tumor-suppressive effect. It acts as a miR-224 sponge and upregulates expression of miR-224 downstream targets p21 and Smad4[71]. circ-ITCH is also lowly expressed in GC. It negatively regulates the Wnt/β-catenin signaling pathway as a tumor suppressor, inhibiting tumor occurrence and development[72]. circPDZD8 is an oncogene circRNA that is upregulated in GC tissues and cells and acts as a miR-197-5p sponge. Its knockout is beneficial for suppression of tumor occurrence[73].
circRNAs participate in various pathological processes. However, their participation in processes such as drug resistance is not mediated by simple up- or downregulation. Their special covalent closed-loop structures play biological roles in transcription and translation and these mechanisms need to be further explored and characterized. Studies have shown that circMTHFD2 can promote GC cell resistance to pemetrexed via molecular sponging of miR-124, thereby increasing the patient’s medication time for pemetrexed, reducing the occurrence of drug resistance, and prolonging patient survival[74]. circ-PVT1 enhances the sensitivity of GC cells to paclitaxel by negatively regulating miR-124-3p, and its knockdown increases the sensitivity of gastric cancer to paclitaxel[75]. circAKT3 promotes DNA damage repair both in vivo and in vitro and inhibits apoptosis in GC patients. Moreover, circAKT3 promotes PIK3R1 expression via miR-198 targeting, and circAKT3 downregulation, which is conducive to the cisplatin sensitivity of GC cells, indicating that circAKT3 knockdown inhibits further growth of GC, improves GC sensitivity and prolongs survival of patients using cisplatin drugs[76]. circFN1 (hsa-circ-0058147) is predominantly located in the cytoplasm and regulates cisplatin activity via miR-182-5p targeting, preventing the ability of miR-182-5p to activate caspase-3 via phosphorylation and inhibition of apoptosis in GC patients. Several studies point towards an involvement of circFN1 in cisplatin resistance and it may represent a future therapeutic target for cisplatin resistance in patients with GC[77]. circRNA may play a role in GC through regulating signaling pathways, and have an impact on the development of drug resistance in these GC patients. The specific mechanisms need to be further studied, but given the evidence so far, the role of circRNAs in GC should not be underestimated. Thus, circRNAs may play an important role as a potential diagnostic and prognostic indicators of GC in the future (Table 1).
| Cancer species | circRNA | Gene expression | Mechanism | Drug | Ref. | |
| Up | Down | |||||
| Gastric cancer | circRNA100876 | ↑ | Via miR-665/YAP1 signaling pathway | NR | [66] | |
| Gastric cancer | circCUL3 | ↑ | Acts as a sponge for miR-515-5p | NR | [67] | |
| Gastric cancer | circTMEM87A | ↑ | Acts as a sponge for miR-142-5p | NR | [68] | |
| Gastric cancer | circRNA0044516 | ↑ | Acts as a sponge for miR-149 | NR | [69] | |
| Gastric cancer | circ0000039 | ↑ | Acts as a sponge for miR-1292-5p | NR | [70] | |
| Gastric cancer | has-circ-0004872 | ↓ | Acts as a sponge for miR-224 | NR | [71] | |
| Gastric cancer | circ-ITCH | ↓ | Negative regulation of Wnt/β-catenin signaling pathway | NR | [72] | |
| Gastric cancer | circPDZD8 | ↑ | Acts as a sponge for miR-197-5p | NR | [73] | |
| Gastric cancer | circMTHFD2 | ↑ | Combined with miR-124 to exert sponge effect and induce MDR-1 protein | Pemetrexed | [74] | |
| Gastric cancer | circ-PVT1 | ↑ | By negatively regulating miR-124-3p | Paclitaxel | [75] | |
| Gastric cancer | circAKT3 | ↑ | Promote PIK3R1 expression through miR-198 | Cisplatin | [76] | |
| Gastric cancer | circFhashsa-circ-0058147 | ↑ | Regulation of cisplatin activity via miR-182-5p | Cisplatin | [77] | |
Hepatocellular carcinoma (HCC) has become one of the leading global cancers with an extremely high fatality rate. The prevalence of liver cancer is higher in men than in women, and it is related to risk factors such as hepatitis B and C, smoking, diabetes, and Aspergillus infections[78]. Studies have demonstrated that several circRNAs may either be potential biomarkers for the diagnosis of HCC or represent therapeutic targets for inhibiting tumor development. For example, hsa-circ-0004018 is highly sensitive to ,-fetoprotein and circ-0016788 may represent a personalized treatment for HCC patients[79]. Similarly, circZKSCAN1 and circPCNX may be used as diagnostic markers for liver cancer[80], while hsa-circRNA-100084 acts as a sponge for hsa-miR-23A-5P, promoting IGF2 expression[81]. circ-0051443 is transferred from normal cells to cancer cells via exosomes and exerts tumor-suppressing functions, inhibiting HCC[82]. circ-0091579 suppresses HCC proliferation and metastasis via targeting miRNA-490-3p. circABCB10 acts as a miR-670-3p sponge, upregulating HMG20A expression and thereby promoting HCC development. However, circABCB10 knockdown inhibits the migration and metastasis of tumor cells[83]. The mechanisms of action of circRNAs in liver cancer need to be further investigated to achieve better insights regarding their role and therapeutic potential.
Patients with liver cancer often face drug resistance. circRNAs have been found to influence drug resistance in several studies. For example, circRNA-101237 is associated with cisplatin sensitivity in HCC patients. Increased levels of circRNA101237 have been reported in the serum of cisplatin-resistant HCC patients and Huh7 cells, and patients with high expression typically have poor prognosis and low survival rates. circRNA_101237 can be used as an independent predictor of the prognosis of liver cancer patients[84]. hsa-circ-u0006294, hsa-circ-u0035944 and hsa-circ-u0084663 activate the Wnt/β-catenin pathway and inhibit HCC growth; of these, hsa-circ-u0006294 and hsa-circ-u0035944 have been found to be downregulated in HCC and influence sorafenib resistance[85]. Likewise, circ-0003418, which inhibits the Wnt/β-catenin pathway, is downregulated in HCC, but increases the sensitivity of HCC to cisplatin. Conversely, knocking down circ-0003418 promotes HCC growth and proliferation[86]. circFBXO11 is upregulated in HCC and promotes progression and oxaliplatin resistance via the miR-605/FOXO3/ABCB1 axis[87].
Liver resection and liver transplantation are common surgical treatments for patients with liver cancer[88]. Patients usually require radiotherapy and chemotherapy after surgical resection. However, with chemotherapy and targeted medication, drug resistance often occurs and the probability of liver cancer recurrence is high with poor prognosis. As presented in the results above, circRNAs have been shown to regulate HCC tumor growth, differentiation, apoptosis and migration. These and future studies may enable the application of circRNAs as potential prospective biomarkers in liver cancers, and the above-mentioned circRNAs may also have certain regulatory effects on drug resistance. Further in-depth research will likely provide new diagnostic indicators for liver cancer patients (Table 2).
| Cancer type | circRNA | Gene expression | Mechanism | Drug | Ref. | |
| Up | Down | |||||
| Liver cancer | circ100084 | ↑ | Promotes IGF2 expression by acting as a sponge for HSA-MIR-23A-5P | NR | [81] | |
| Liver cancer | circABCB10 | ↑ | Upregulates HMG20A expression by acting as a miR-670-3p sponge | NR | [83] | |
| Liver cancer | circRNA-101237 | ↑ | Increased levels in serum and Huh7 cells of cisplatin-resistant HCC patients | Cisplatin | [84] | |
| Liver cancer | Hsa-circ-u0006294, hsa-circ-u0035944, hsa-circ-u0084663 | ↓ | Plays a role by activating the Wnt/β-catenin pathway | Sorafenib | [85] | |
| Liver cancer | circ-0003418 | ↓ | Inhibits Wnt/β-catenin pathway | Cisplatin | [86] | |
| Liver cancer | circFBXO11 | ↑ | Uses miR-605 /FOXO3/ABCB1 axis adjustment | Oxaliplatin | [87] | |
Colorectal cancer (CRC) is the third most deadly and fourth most commonly diagnosed cancer in the world. Alcohol, tobacco, sedentary lifestyles and obesity have been reported to increase the risk of CRC[89]. It is estimated that by 2030, the global incidence of CRC will be as high as 60%, meaning that there will be nearly 1.4 million new patients[90]. Many circRNAs are dysregulated in CRC tissues compared with the normal mucosa. circNSUN2 is opposite to the 5p15 amplicon and interacts with YTHDC1, SRSF3, and NXF1. Upregulation of circNSUN2 in CRC promotes metastasis and is associated with poor prognosis; as a cancer-promoting circRNA, it may be used as a clinical diagnostic or prognostic marker in the future[91]. hsa-circ-101555 (circ101555) is also a cancer-promoting circRNA that sponges miR-597-5p, whose potential targets are CDK6 and RPA3. hsa-circ-101555 knockdown results in inhibition of CRC proliferation and metastasis, thus it may also be used as a future prognostic factor and therapeutic target for CRC[92]. High levels of circHIPK3 expression result in a low overall survival rate of CRC, and there is a negative correlation between circHIPK3 expression and CRC survival. circHIPK3 can inhibit the proliferation, migration and invasion of CRC cells[93]. circCCDC66 exerts its function by regulation of a subset of oncogenes, and studies have shown that circCCDC66 knockout inhibits CRC cell growth and invasion, while circCCDC66 upregulation results in poor CRC prognosis. Additionally, circCCDC66 knockdown increases the therapeutic effect of oxaliplatin on CRC[94,95]. hsa-circ-0000069 is highly expressed in CRC and its knockdown significantly inhibits the growth and proliferation of tumor cells[96].
In addition to their roles in both suppression and promotion of cancer, circRNAs also play a key role in drug resistance. Topoisomerase is widely distributed in eukaryotic cells and was identified as the target of the anticancer drug camptothecin in the 1970s. Irinotecan, a camptothecin derivative, is used as a treatment for CRC. It is an S-phase-specific, effective anticancer treatment for metastatic CRC. circRNAs may promote irinotecan resistance by regulating cancer-stem-cell-like characteristics[97]. For example, circ001680 modulates BMI1 via miR-340 and it can be used to induce irinotecan sensitivity in CRC, suppressing CRC proliferation and migration. circ001680 may thus represent a potential indicator of irinotecan chemotherapy resistance[98]. hsa-circ-0005963 inhibits glycolysis and reverses drug resistance by targeting pKM2 via miR-122 sponging: it increases oxaliplatin sensitivity in CRC via downregulation of PKM2[99]. circ-0000338, which is found in extracellular vesicles, has been found to improve 5-FU resistance in CRC cells. circ-0000338 may have dual regulatory effects, capable of both suppressing and causing cancer[100]. hsa-circ-32883 expression is upregulated in CRC, and the circRNA produced by the exon region exerts a cancer-promoting effect. Thus, hsa-circ-32883 may be a powerful biological target, but its mechanism of resistance relating to 5-FU and oxaliplatin is not yet clear[101]. The expression of circ0007031, circ0000504 and circ0007006 is elevated in CRC. Upregulation of hsa-circ-0000504 can reduce the inhibitory effect of hsa-miR485-5p on STAT3 and promotes chemical radiation resistance. Additionally, silencing STAT3 results in a significant reduction of 5-FU resistance and clone survival after irradiation[102]. hsa-circ-0079662 regulates HOXA9 and induces oxaliplatin resistance via hsa-miR-324-5p binding in CRC[103]. Studies have shown that circDDX17 can enhance CRC sensitivity to 5-Fu via upregulation of miR-31-5p/KANK1[104].
Taken together, circRNAs appear to play important roles in CRC which is summarized in Table 3.
| Cancer type | circRNA | Gene expression | Mechanism | Drug | Ref. | |
| Up | Down | |||||
| Colon cancer | circNSUN2 | ↑ | Interact with YTHDC1, SRSF3 and NXF1 | NR | [91] | |
| Colon cancer | hsa-circ-101555 | ↑ | Acts as miR-597-5p sponge | NR | [92] | |
| Colon cancer | circHIPK3 | ↑ | Acts as miR-7 sponge | NR | [93] | |
| Colon cancer | circCCDC66 | ↑ | DHX9 phosphorylation | Oxaliplatin | [94,95] | |
| Colon cancer | circ-001680 | ↑ | Adjust BMI1 via miR-340 | Irinotecan | [98] | |
| Colon cancer | hsa-circ-0005963 | ↑ | Acts as miR-122 sponge | Oxaliplatin | [99] | |
| Colon cancer | circ-0000338 | ↑ or ↓ | Knock out circ-0000338 improves drug resistance | 5-FU | [100] | |
| Colon cancer | hsa-circ-32883 | ↑ | uncertain | 5-FU, oxaliplatin | [101] | |
| Colon cancer | circ0007031, circ0000504, circ0007006 | ↑ | Use hsa-miR485-5p pathway | 5-FU | [102] | |
| Colon cancer | hsa-circ-0079662 | ↑ | Using the TNF-α pathway | oxaliplatin | [103] | |
| Colon cancer | circDDX17 | ↑ | Adjustable via miR-31-5p/KANK1 axis | 5-FU | [104] | |
Esophageal cancer (EC) is an extremely aggressive type of cancer with a 5-year survival rate between 15% and 20%. EC is the eighth most prevalent tumor in the world and the sixth leading cause of cancer-related deaths. Due to its poor prognosis and low survival rate, it is particularly important to identify the reliable biomarkers for its early diagnosis[105].
A study has reported 1045 upregulated and 1032 downregulated circRNAs in EC[106], and several studies have suggested that circRNAs play an important role in the regulation of gene transcription in EC. For example, hsa-cric-0004771 promotes esophageal squamous cell carcinoma via the miR-339-5P/CDC25A axis, and its knockdown inhibits proliferation and migration of development of esophageal squamous-cell carcinoma (ESCC). Nonetheless, its mechanisms of action have not been fully elucidated[107]. circUBAP2 is upregulated in ESCC and acts as a miR-422a sponge. circUBAP2 knockdown inhibits ESCC progression and prevents the spread of tumor cells[108]. hsa-circ-0006948 is likewise upregulated in ESCC and promotes the growth and development of tumor cells via the miR-490-3p/HMGA2 axis[109]. Similarly, circ-0000654 is upregulated in ESCC. It promotes proliferation and migration of tumor cells via the miR-149-5p/IL-6/STAT3 axis and tumor progression can be suppressed by circ-0000654 knockdown[110]. circGSK3 is another upregulated circRNA in ESCC. It promotes tumor cell proliferation via β-catenin signaling, which is positively correlated with poor prognosis[111]. circ-Foxo3 exhibits low expression in ESCC and modulates the miR-23a/PTEN pathway to inhibit ESCC development[112]. circPVT1 has been found to be significantly upregulated in EC and is suggested to play a role in the proliferation of TE-10 tumor cells via miR-4663[113].
ciRS-7 is abnormally increased in ESCC and promotes tumor cell growth. It may act as an oncogene, promoting promotes tumor cells proliferation by acting as a miR-876-5p sponge. Meng et al[114] have shown that ciRS-7 inhibits autophagy in ESCC induced by starvation or rapamycin immunosuppressive agents. Huang et al[115] confirmed that ciRS-7 promotes migration and invasion of ESCC cells via miR-7/KLF4 and NF-κB signaling pathways. Knockdown of KLF-4 in ESCC attenuates ciRS-7 invasion[114,115]. Drug resistance in EC is affected by cell proliferation, metastasis, glycolytic enzymes and other factors, among them circRNAs, lncRNAs, miRNAs, and acetyl-coenzyme, a synthetase short-chain family member 2 (ACSS2). Of the above-mentioned factors, lncRNAs have been particularly well studied. For example, ACSS2 affects the absorption of cisplatin in ESCC and protects cancer cells from cisplatin[116]. Conversely, research investigating the relationship between circRNAs and EC drug resistance is lacking. It has been demonstrated that circRNA-001275 upregulates Wnt family member 7A (Wnt7a), which triggers tumor cell growth via cisplatin, via miR-370-3p, inducing drug resistance[117]. Although there are few studies on circRNA in esophageal cancer drug resistance (Table 4), it can be expected that more circRNAs that play a role in the development and resistance of esophageal cancer will be discovered.
| Cancer type | circRNA | Gene expression | Mechanism | Drug | Ref. | |
| Up | Down | |||||
| Esophageal cancer | circUBAP2 | ↑ | Acts as miR-422a sponge | NR | [108] | |
| Esophageal cancer | hsa-circ-0006948 | ↑ | miR-490-3p to enhance HMGA2-induced EMT | NR | [109] | |
| Esophageal cancer | circ-0000654 | ↑ | miR-149-5p/IL-6/STAT3 pathway | NR | [110] | |
| Esophageal cancer | circGSK3 | ↑ | β-catenin signal passing | NR | [111] | |
| Esophageal cancer | circ-Foxo3 | ↓ | miR-23a/PTEN pathway | NR | [112] | |
| Esophageal cancer | circPVT1 | ↑ | Acts as miR-4663 sponge | NR | [113] | |
| Esophageal cancer | ciRS-7 | ↑ | Acts as a miR-876-5p sponge | Rapamycin | [114,115] | |
| Esophageal cancer | circRNA-001275 | ↑ | Acts as a miR-370-3p sponge | Cisplatin | [117] | |
Pancreatic cancer (PC) has the seventh highest mortality rate among all cancers. It is particularly common in North America, Australia and Europe, and the incidence of PC is higher in males than in females[118]. Several circRNAs have been found to be involved in PC and may be used as therapeutic targets. For example, circBFAR (hsa-circ-0009065) is upregulated in pancreatic ductal adenocarcinoma (PDAC), acting as a miR-34b-5p sponge, and modulates the MET/PI3K/Akt signaling pathway[119]. circFOXK2 is likewise upregulated in PDAC and promotes the expression of ANK1, GDNF and PAX6 by acting as a sponge for miR-942, ultimately promoting the proliferation of tumor cells. In line with this, knockdown of circFOXK2 was conducive to the inhibition of tumor progression[120]. circ-ADAM9 is highly expressed in PC. It activates the ERK/VEGF signaling pathway via miR-217[121]. circ-ASH2L is also highly expressed in PDAC and acts as a sponge for miR-34a, promoting tumor cell proliferation and tumor progression via regulation of Notch1 in PDAC[122]. circ-LDLRAD3 is also overexpressed in PC and regulates cell proliferation via the miR-137-3p/PTN axis. It affects prognosis of PC and its knockdown inhibits the invasion of tumor cells[123]. circ-0000977 is upregulated in PC and acts as an miR-153 sponge. Knockdown of circ-0000977 conveys the limiting effect of HIF1A and ADAM10 on PC cells under hypoxia[124]. hsa-circRNA-0007334 is upregulated in PDAC and competes for binding with hsa-miR-144-3p[125]. Similarly as in EC, high expression of ciRS-7 promotes PDAC development via acting as an miR-7 sponge and regulating the EGFR/STAT3 signaling pathway. Knockdown of ciRS-7 inhibits the proliferation and invasion of tumor cells[126]. circ-0030235 is upregulated in PDAC and acts as a sponge for miR-1253 and miR-1294. Its expression is a poor prognostic indicator in PDAC[127]. circZMYM2 is also upregulated in PC. It acts as a miR-335-5p sponge and regulates the downstream oncogene JMJD2C. Knockdown of circZMYM2 is conducive for the inhibition of tumor growth[128]. circ-0007534 is upregulated in PDAC and acts as a sponge for miR-625 and miR-892b, increasing the proliferation and invasion of tumor cells[129]. circ-IARS is upregulated in PDAC, absorbs miR-122 as a sponge, and increases RhoA activity and F-actin expression, reduces ZO-1 expression, and promotes tumors by enhancing endothelial cell monolayer permeability, invasion and metastasis[130]. Moreover, circ-PDE8A is upregulated in PDAC and acts as a sponge for miR-338, regulating MACC1 expression and promoting the growth of tumor cells via the MACC/MET/ERK or AKT signaling pathways[131].
Gemcitabine is an important drug for the treatment of advanced PC. A study showed 68 upregulated and 58 downregulated circRNAs in gemcitabine-treated PANC-1 cells of which two were significantly upregulated (chr14: 10140210 191014644448+, chr4: 52729603-52780244+)[132]. circHIPK3 is highly expressed in PC and promotes gemcitabine resistance in PC cells by acting as a miR-330-5p sponge and targeting RASSF1[133].
Overall, circRNAs carry out important roles in tumors. Following further studies, they are expected to serve as new therapeutic targets and biomarkers for tumor diagnosis in the future[134] (Table 5).
| Cancer type | circRNA | Gene expression | Mechanism | Drug | Ref. | |
| Up | Down | |||||
| Pancreatic cancer | circBFAR | ↑ | Acts as a miR-34b-5p sponge | NR | [119] | |
| Pancreatic cancer | circFOXK2 | ↑ | Acts as miR-942 sponge | NR | [120] | |
| Pancreatic cancer | circ-ADAM9 | ↑ | Activate ERK/VEGF signaling pathway through miR-217 | NR | [121] | |
| Pancreatic cancer | circ-ASH2L | ↑ | Acts as a sponge for miR-34a | NR | [122] | |
| Pancreatic cancer | circ-LDLRAD3 | ↑ | Adjustable via miR-137-3p/PTN axis | NR | [123] | |
| Pancreatic cancer | circ-0000977 | ↑ | Acts as a sponge for miR-153 | NR | [124] | |
| Pancreatic cancer | hsa-circRNA-0007334 | ↑ | Competitive binding through hsa-miR-144-3p | NR | [125] | |
| pancreatic cancer | ciRS-7 | ↑ | Acts as miR-7 sponge | NR | [126] | |
| Pancreatic cancer | circ-0030235 | ↑ | Acts as miR-1253 and miR-1294 sponge | NR | [127] | |
| Pancreatic cancer | circZMYM2 | ↑ | Acts as a miR-335-5p sponge | NR | [128] | |
| Pancreatic cancer | circ-0007534 | ↑ | Acts as miR-625, miR-892b sponge | NR | [129] | |
| Pancreatic cancer | circ-IARS | ↑ | Acts as miR-122 sponge | NR | [130] | |
| Pancreatic cancer | circ-PDE8A | ↑ | As a sponge of miR-338 to regulate MACC1 | NR | [131] | |
| Pancreatic cancer | circHIPK3 | ↑ | Acts as a miR-330-5p sponge | Gemcitabine | [133] | |
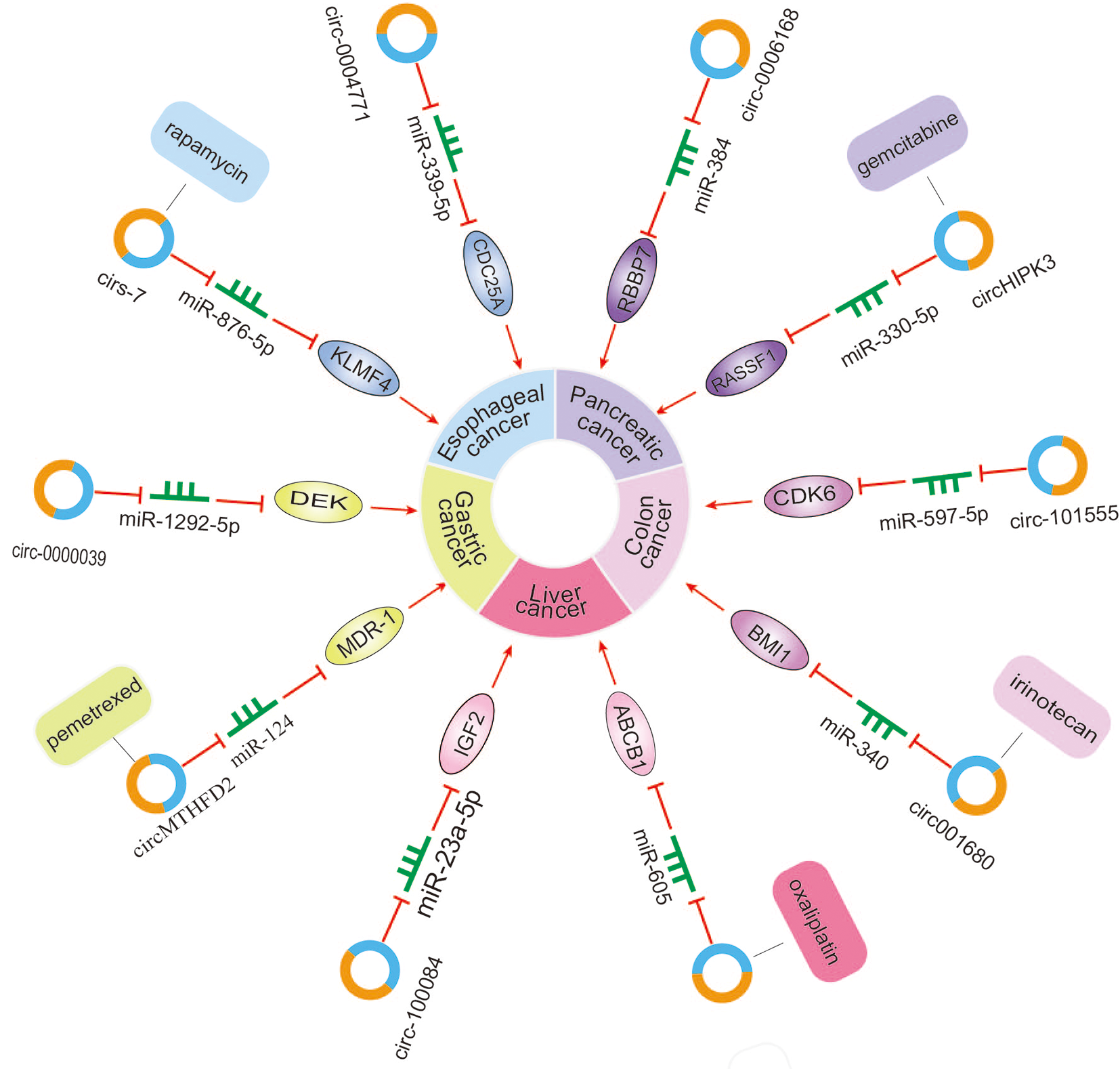
Following chemotherapy, cancers may relapse, and many patients gradually develop resistance to several anticancer drugs. In the present study, we have summarized findings that report that circRNAs may inhibit or promote tumor growth and migration by acting as miR sponges or other pathways in vitro and in vivo. circRNAs may play the role of vectors in drug resistance due to their unique structure, promoting an inhibitory effect of drugs on tumor cells. Drug resistance of tumor cells is associated with many factors and the underlying mechanisms are complex. Resistance mechanisms conveyed by circRNAs may be related to glycolysis, ATP levels, or other factors. With regard to the preliminary results of studies on drug resistance, several mechanisms and pathways have been reported to be involved in the regulation of drug resistance by circRNAs in esophageal, pancreatic, colon, liver and gastric cancer. Although further exploring the mechanisms may enable the development of preventive mechanisms against drug resistance among cancer cells, there is still a long way to go for understanding circRNAs and tumor drug resistance (Figure 3).
There is currently a lack of monitoring methods and treatment tools for the development of gastrointestinal tumors and the occurrence of drug resistance. The pathogenesis of gastrointestinal tumors is diverse, including many unstable factors such as DNA damage repair, genome instability and mutations, gene transcription regulation, chromosome modifications, inflammation, tobacco and alcohol, obesity, and autoimmunity. circRNAs exert rich biological functions and play an important role in gastrointestinal tumors. This article discusses the mechanisms of tumor development and drug resistance of gastrointestinal tumors related to circRNAs. While the exact mechanisms of involvement of circRNAs have not been fully elucidated, their important roles in various tumors as observed in initial studies have attracted a lot of attention and have huge potential for future clinical approaches. With advances in biotechnology, new applications such as genetic testing, polymerase chain reaction, high-throughput sequencing, and in-depth research in gastrointestinal tumors, it is expected that circRNAs could be developed into diagnostics or therapeutic targets for the treatment of gastrointestinal tumors. In case circRNAs can become a mass-produced product due to technological innovation, they could be used for clinical treatment. Noninvasive detection distinguishing malignant and benign tumors may also be enabled via biotechnological advances. Other options could include the development of new prognosis indicators for patients and monitoring of drug sensitivity, all aimed to ultimately extend patient life and disease-free survival. Based on the presented literature, there is hope that circRNAs may become an effective therapeutic target for cancer treatment in the future.
Provenance and peer review: Unsolicited article; Externally peer reviewed.
Specialty type: Gastroenterology and hepatology
Country/Territory of origin: China
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Deshwal H S-Editor: Gong ZM L-Editor: Kerr C P-Editor: Guo X
| 1. | Maia FCP, Silva TA, Generoso SV, Correia MITD. Malnutrition is associated with poor health-related quality of life in surgical patients with gastrointestinal cancer. Nutrition. 2020;75-76:110769. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 27] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 2. | Varughese LA, Lau-Min KS, Cambareri C, Damjanov N, Massa R, Reddy N, Oyer R, Teitelbaum U, Tuteja S. DPYD and UGT1A1 Pharmacogenetic Testing in Patients with Gastrointestinal Malignancies: An Overview of the Evidence and Considerations for Clinical Implementation. Pharmacotherapy. 2020;40:1108-1129. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 20] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 3. | Nie H, Wang Y, Liao Z, Zhou J, Ou C. The function and mechanism of circular RNAs in gastrointestinal tumours. Cell Prolif. 2020;53:e12815. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 44] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 4. | Sanger HL, Klotz G, Riesner D, Gross HJ, Kleinschmidt AK. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci U S A. 1976;73:3852-3856. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1253] [Cited by in RCA: 1690] [Article Influence: 34.5] [Reference Citation Analysis (0)] |
| 5. | Cocquerelle C, Mascrez B, Hétuin D, Bailleul B. Mis-splicing yields circular RNA molecules. FASEB J. 1993;7:155-160. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 758] [Cited by in RCA: 926] [Article Influence: 28.9] [Reference Citation Analysis (0)] |
| 6. | Capel B, Swain A, Nicolis S, Hacker A, Walter M, Koopman P, Goodfellow P, Lovell-Badge R. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell. 1993;73:1019-1030. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 781] [Cited by in RCA: 882] [Article Influence: 27.6] [Reference Citation Analysis (0)] |
| 7. | Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One. 2012;7:e30733. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1589] [Cited by in RCA: 1952] [Article Influence: 150.2] [Reference Citation Analysis (0)] |
| 8. | Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, Loewer A, Ziebold U, Landthaler M, Kocks C, le Noble F, Rajewsky N. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333-338. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6253] [Cited by in RCA: 6009] [Article Influence: 500.8] [Reference Citation Analysis (0)] |
| 9. | Conn SJ, Pillman KA, Toubia J, Conn VM, Salmanidis M, Phillips CA, Roslan S, Schreiber AW, Gregory PA, Goodall GJ. The RNA binding protein quaking regulates formation of circRNAs. Cell. 2015;160:1125-1134. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1218] [Cited by in RCA: 1611] [Article Influence: 161.1] [Reference Citation Analysis (0)] |
| 10. | Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L, Hanan M, Wyler E, Perez-Hernandez D, Ramberger E, Shenzis S, Samson M, Dittmar G, Landthaler M, Chekulaeva M, Rajewsky N, Kadener S. Translation of CircRNAs. Mol Cell. 2017;66:9-21.e7. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 988] [Cited by in RCA: 1348] [Article Influence: 168.5] [Reference Citation Analysis (0)] |
| 11. | Wu X, Xiao Y, Ma J, Wang A. Circular RNA: A novel potential biomarker for skin diseases. Pharmacol Res. 2020;158:104841. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 38] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 12. | Li B, Li Y, Hu L, Liu Y, Zhou Q, Wang M, An Y, Li P. Role of Circular RNAs in the Pathogenesis of Cardiovascular Disease. J Cardiovasc Transl Res. 2020;13:572-583. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 22] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 13. | Yu G, Yang Z, Peng T, Lv Y. Circular RNAs: Rising stars in lipid metabolism and lipid disorders. J Cell Physiol. 2021;236:4797-4806. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 37] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 14. | Yang Y, Gao X, Zhang M, Yan S, Sun C, Xiao F, Huang N, Yang X, Zhao K, Zhou H, Huang S, Xie B, Zhang N. Novel Role of FBXW7 Circular RNA in Repressing Glioma Tumorigenesis. J Natl Cancer Inst. 2018;110. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 568] [Cited by in RCA: 823] [Article Influence: 102.9] [Reference Citation Analysis (0)] |
| 15. | Prats AC, David F, Diallo LH, Roussel E, Tatin F, Garmy-Susini B, Lacazette E. Circular RNA, the Key for Translation. Int J Mol Sci. 2020;21. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 81] [Cited by in RCA: 90] [Article Influence: 18.0] [Reference Citation Analysis (0)] |
| 16. | Zhang L, Hou C, Chen C, Guo Y, Yuan W, Yin D, Liu J, Sun Z. The role of N6-methyladenosine (m6A) modification in the regulation of circRNAs. Mol Cancer. 2020;19:105. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 95] [Cited by in RCA: 223] [Article Influence: 44.6] [Reference Citation Analysis (0)] |
| 17. | Nakamoto K, Abe N, Tsuji G, Kimura Y, Tomoike F, Shimizu Y, Abe H. Chemically synthesized circular RNAs with phosphoramidate linkages enable rolling circle translation. Chem Commun (Camb). 2020;56:6217-6220. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 8] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 18. | Shan C, Zhang Y, Hao X, Gao J, Chen X, Wang K. Biogenesis, functions and clinical significance of circRNAs in gastric cancer. Mol Cancer. 2019;18:136. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 99] [Cited by in RCA: 172] [Article Influence: 28.7] [Reference Citation Analysis (0)] |
| 19. | Huang C, Shan G. What happens at or after transcription: Insights into circRNA biogenesis and function. Transcription. 2015;6:61-64. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 97] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 20. | Di Timoteo G, Dattilo D, Centrón-Broco A, Colantoni A, Guarnacci M, Rossi F, Incarnato D, Oliviero S, Fatica A, Morlando M, Bozzoni I. Modulation of circRNA Metabolism by m6A Modification. Cell Rep. 2020;31:107641. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 244] [Article Influence: 61.0] [Reference Citation Analysis (0)] |
| 21. | Zhou WY, Cai ZR, Liu J, Wang DS, Ju HQ, Xu RH. Circular RNA: metabolism, functions and interactions with proteins. Mol Cancer. 2020;19:172. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 389] [Cited by in RCA: 780] [Article Influence: 156.0] [Reference Citation Analysis (0)] |
| 22. | Chen LL. The expanding regulatory mechanisms and cellular functions of circular RNAs. Nat Rev Mol Cell Biol. 2020;21:475-490. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 417] [Cited by in RCA: 999] [Article Influence: 199.8] [Reference Citation Analysis (0)] |
| 23. | Qu S, Zhong Y, Shang R, Zhang X, Song W, Kjems J, Li H. The emerging landscape of circular RNA in life processes. RNA Biol. 2017;14:992-999. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 233] [Cited by in RCA: 325] [Article Influence: 36.1] [Reference Citation Analysis (0)] |
| 24. | Bai Y, Zhang Y, Han B, Yang L, Chen X, Huang R, Wu F, Chao J, Liu P, Hu G, Zhang JH, Yao H. Circular RNA DLGAP4 Ameliorates Ischemic Stroke Outcomes by Targeting miR-143 to Regulate Endothelial-Mesenchymal Transition Associated with Blood-Brain Barrier Integrity. J Neurosci. 2018;38:32-50. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 259] [Cited by in RCA: 316] [Article Influence: 45.1] [Reference Citation Analysis (0)] |
| 25. | Wang S, Tang D, Wang W, Yang Y, Wu X, Wang L, Wang D. circLMTK2 acts as a sponge of miR-150-5p and promotes proliferation and metastasis in gastric cancer. Mol Cancer. 2019;18:162. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 62] [Cited by in RCA: 64] [Article Influence: 10.7] [Reference Citation Analysis (0)] |
| 26. | Cervera-Carles L, Dols-Icardo O, Molina-Porcel L, Alcolea D, Cervantes-Gonzalez A, Muñoz-Llahuna L, Clarimon J. Assessing circular RNAs in Alzheimer's disease and frontotemporal lobar degeneration. Neurobiol Aging. 2020;92:7-11. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 33] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 27. | Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495:384-388. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4631] [Cited by in RCA: 6034] [Article Influence: 502.8] [Reference Citation Analysis (0)] |
| 28. | Qu X, Li Z, Chen J, Hou L. The emerging roles of circular RNAs in CNS injuries. J Neurosci Res. 2020;98:1485-1497. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 18] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 29. | He J, Xie Q, Xu H, Li J, Li Y. Circular RNAs and cancer. Cancer Lett. 2017;396:138-144. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 145] [Cited by in RCA: 176] [Article Influence: 22.0] [Reference Citation Analysis (0)] |
| 30. | Holdt LM, Stahringer A, Sass K, Pichler G, Kulak NA, Wilfert W, Kohlmaier A, Herbst A, Northoff BH, Nicolaou A, Gäbel G, Beutner F, Scholz M, Thiery J, Musunuru K, Krohn K, Mann M, Teupser D. Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat Commun. 2016;7:12429. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 697] [Cited by in RCA: 858] [Article Influence: 95.3] [Reference Citation Analysis (0)] |
| 31. | Chen J, Li Y, Zheng Q, Bao C, He J, Chen B, Lyu D, Zheng B, Xu Y, Long Z, Zhou Y, Zhu H, Wang Y, He X, Shi Y, Huang S. Circular RNA profile identifies circPVT1 as a proliferative factor and prognostic marker in gastric cancer. Cancer Lett. 2017;388:208-219. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 446] [Cited by in RCA: 570] [Article Influence: 63.3] [Reference Citation Analysis (0)] |
| 32. | Li R, Jiang J, Shi H, Qian H, Zhang X, Xu W. CircRNA: a rising star in gastric cancer. Cell Mol Life Sci. 2020;77:1661-1680. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 249] [Cited by in RCA: 276] [Article Influence: 55.2] [Reference Citation Analysis (0)] |
| 33. | Petkovic S, Müller S. RNA circularization strategies in vivo and in vitro. Nucleic Acids Res. 2015;43:2454-2465. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 236] [Cited by in RCA: 267] [Article Influence: 26.7] [Reference Citation Analysis (0)] |
| 34. | Villaseñor R, Baubec T. Regulatory mechanisms governing chromatin organization and function. Curr Opin Cell Biol. 2021;70:10-17. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 33] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 35. | Kim Y, Yu H. Shaping of the 3D genome by the ATPase machine cohesin. Exp Mol Med. 2020;52:1891-1897. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 20] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 36. | Tang Z, Li X, Zhao J, Qian F, Feng C, Li Y, Zhang J, Jiang Y, Yang Y, Wang Q, Li C. TRCirc: a resource for transcriptional regulation information of circRNAs. Brief Bioinform. 2019;20:2327-2333. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 39] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 37. | Tan S, Kang Y, Li H, He HQ, Zheng L, Wu SQ, Ai K, Zhang L, Xu R, Zhang XZ, Zhao XK, Zhu X. circST6GALNAC6 suppresses bladder cancer metastasis by sponging miR-200a-3p to modulate the STMN1/EMT axis. Cell Death Dis. 2021;12:168. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 37] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 38. | Lacerda R, Menezes J, Romão L. More than just scanning: the importance of cap-independent mRNA translation initiation for cellular stress response and cancer. Cell Mol Life Sci. 2017;74:1659-1680. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 97] [Article Influence: 12.1] [Reference Citation Analysis (0)] |
| 39. | Dong R, Ma XK, Chen LL, Yang L. Increased complexity of circRNA expression during species evolution. RNA Biol. 2017;14:1064-1074. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 125] [Cited by in RCA: 162] [Article Influence: 18.0] [Reference Citation Analysis (0)] |
| 40. | Hu X, Zhu M, Zhang X, Liu B, Liang Z, Huang L, Xu J, Yu L, Li K, Zar MS, Xue R, Cao G, Gong C. Identification and characterization of circular RNAs in the silkworm midgut following Bombyx mori cytoplasmic polyhedrosis virus infection. RNA Biol. 2018;15:292-301. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 43] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 41. | Granados-Riveron JT, Aquino-Jarquin G. The complexity of the translation ability of circRNAs. Biochim Biophys Acta. 2016;1859:1245-1251. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 113] [Cited by in RCA: 158] [Article Influence: 17.6] [Reference Citation Analysis (0)] |
| 42. | Starke S, Jost I, Rossbach O, Schneider T, Schreiner S, Hung LH, Bindereif A. Exon circularization requires canonical splice signals. Cell Rep. 2015;10:103-111. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 482] [Cited by in RCA: 615] [Article Influence: 55.9] [Reference Citation Analysis (0)] |
| 43. | Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, Zhong G, Yu B, Hu W, Dai L, Zhu P, Chang Z, Wu Q, Zhao Y, Jia Y, Xu P, Liu H, Shan G. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22:256-264. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1619] [Cited by in RCA: 2213] [Article Influence: 221.3] [Reference Citation Analysis (0)] |
| 44. | Lei M, Zheng G, Ning Q, Zheng J, Dong D. Translation and functional roles of circular RNAs in human cancer. Mol Cancer. 2020;19:30. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 202] [Cited by in RCA: 498] [Article Influence: 99.6] [Reference Citation Analysis (0)] |
| 45. | Tang Q, Hann SS. Biological Roles and Mechanisms of Circular RNA in Human Cancers. Onco Targets Ther. 2020;13:2067-2092. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 88] [Cited by in RCA: 129] [Article Influence: 25.8] [Reference Citation Analysis (0)] |
| 46. | Li X, Wang J, Zhang C, Lin C, Zhang J, Zhang W, Lu Y, Zheng L, Li X. Circular RNA circITGA7 inhibits colorectal cancer growth and metastasis by modulating the Ras pathway and upregulating transcription of its host gene ITGA7. J Pathol. 2018;246:166-179. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 135] [Cited by in RCA: 206] [Article Influence: 29.4] [Reference Citation Analysis (0)] |
| 47. | Xie M, Yu T, Jing X, Ma L, Fan Y, Yang F, Ma P, Jiang H, Wu X, Shu Y, Xu T. Exosomal circSHKBP1 promotes gastric cancer progression via regulating the miR-582-3p/HUR/VEGF axis and suppressing HSP90 degradation. Mol Cancer. 2020;19:112. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 136] [Cited by in RCA: 290] [Article Influence: 58.0] [Reference Citation Analysis (0)] |
| 48. | Bannister AJ, Kouzarides T. Regulation of chromatin by histone modifications. Cell Res. 2011;21:381-395. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3359] [Cited by in RCA: 3991] [Article Influence: 285.1] [Reference Citation Analysis (0)] |
| 49. | Tang L, Xiong W, Zhang L, Wang D, Wang Y, Wu Y, Wei F, Mo Y, Hou X, Shi L, Xiong F, Zhang S, Gong Z, Liao Q, Xiang B, Zhang W, Zhou M, Li X, Li G, Guo C, Zeng Z. circSETD3 regulates MAPRE1 through miR-615-5p and miR-1538 sponges to promote migration and invasion in nasopharyngeal carcinoma. Oncogene. 2021;40:307-321. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 55] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 50. | Jie M, Wu Y, Gao M, Li X, Liu C, Ouyang Q, Tang Q, Shan C, Lv Y, Zhang K, Dai Q, Chen Y, Zeng S, Li C, Wang L, He F, Hu C, Yang S. CircMRPS35 suppresses gastric cancer progression via recruiting KAT7 to govern histone modification. Mol Cancer. 2020;19:56. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 71] [Cited by in RCA: 122] [Article Influence: 24.4] [Reference Citation Analysis (0)] |
| 51. | He Y, Zhang Q, Zheng Q, Yu X, Guo W. Distinct 5-methylcytosine profiles of circular RNA in human hepatocellular carcinoma. Am J Transl Res. 2020;12:5719-5729. [PubMed] |
| 52. | Zhao Z, Song J, Tang B, Fang S, Zhang D, Zheng L, Wu F, Gao Y, Chen C, Hu X, Weng Q, Yang Y, Tu J, Ji J. CircSOD2 induced epigenetic alteration drives hepatocellular carcinoma progression through activating JAK2/STAT3 signaling pathway. J Exp Clin Cancer Res. 2020;39:259. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 66] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 53. | Wang Y, Shi S, Zhang Q, Dong H, Zhang J. MicroRNA-206 upregulation relieves circTCF25-induced osteosarcoma cell proliferation and migration. J Cell Physiol. 2020;. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 8] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 54. | Wang Y, Xu R, Zhang D, Lu T, Yu W, Wo Y, Liu A, Sui T, Cui J, Qin Y, Dong Y, Leng X, Kong D, Du W, Huang Z, Su W, Yuan T, Sun X, Wang J, Jiao W. Circ-ZKSCAN1 regulates FAM83A expression and inactivates MAPK signaling by targeting miR-330-5p to promote non-small cell lung cancer progression. Transl Lung Cancer Res. 2019;8:862-875. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 77] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 55. | Pan G, Mao A, Liu J, Lu J, Ding J, Liu W. Circular RNA hsa_circ_0061825 (circ-TFF1) contributes to breast cancer progression through targeting miR-326/TFF1 signalling. Cell Prolif. 2020;53:e12720. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 92] [Cited by in RCA: 97] [Article Influence: 19.4] [Reference Citation Analysis (0)] |
| 56. | Li Y, Li C, Xu R, Wang Y, Li D, Zhang B. A novel circFMN2 promotes tumor proliferation in CRC by regulating the miR-1182/hTERT signaling pathways. Clin Sci (Lond). 2019;133:2463-2479. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 66] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 57. | Wang GJ, Yu TY, Li YR, Liu YJ, Deng BB. Circ_0000190 suppresses gastric cancer progression potentially via inhibiting miR-1252/PAK3 pathway. Cancer Cell Int. 2020;20:351. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 16] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 58. | Kumala S, Fujarewicz K, Jayaraju D, Rzeszowska-Wolny J, Hancock R. Repair of DNA strand breaks in a minichromosome in vivo: kinetics, modeling, and effects of inhibitors. PLoS One. 2013;8:e52966. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 7] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 59. | Gamper AM, Rofougaran R, Watkins SC, Greenberger JS, Beumer JH, Bakkenist CJ. ATR kinase activation in G1 phase facilitates the repair of ionizing radiation-induced DNA damage. Nucleic Acids Res. 2013;41:10334-10344. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 57] [Cited by in RCA: 58] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 60. | Podralska M, Ciesielska S, Kluiver J, van den Berg A, Dzikiewicz-Krawczyk A, Slezak-Prochazka I. Non-Coding RNAs in Cancer Radiosensitivity: MicroRNAs and lncRNAs as Regulators of Radiation-Induced Signaling Pathways. Cancers (Basel). 2020;12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 41] [Cited by in RCA: 47] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 61. | Xu X, Zhang J, Tian Y, Gao Y, Dong X, Chen W, Yuan X, Yin W, Xu J, Chen K, He C, Wei L. CircRNA inhibits DNA damage repair by interacting with host gene. Mol Cancer. 2020;19:128. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 89] [Cited by in RCA: 255] [Article Influence: 51.0] [Reference Citation Analysis (0)] |
| 62. | Jonaitis P, Kiudelis V, Streleckiene G, Gedgaudas R, Skieceviciene J, Kupcinskas J. Novel Biomarkers in the Diagnosis of Benign and Malignant GI Diseases. Dig Dis. 2021;. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 7] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 63. | Xu T, Wang M, Jiang L, Ma L, Wan L, Chen Q, Wei C, Wang Z. CircRNAs in anticancer drug resistance: recent advances and future potential. Mol Cancer. 2020;19:127. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 64] [Cited by in RCA: 80] [Article Influence: 16.0] [Reference Citation Analysis (0)] |
| 64. | Wei L, Sun J, Zhang N, Zheng Y, Wang X, Lv L, Liu J, Xu Y, Shen Y, Yang M. Noncoding RNAs in gastric cancer: implications for drug resistance. Mol Cancer. 2020;19:62. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 279] [Cited by in RCA: 334] [Article Influence: 66.8] [Reference Citation Analysis (0)] |
| 65. | Yuan X, Yuan Y, He Z, Li D, Zeng B, Ni Q, Yang M, Yang D. The Regulatory Functions of Circular RNAs in Digestive System Cancers. Cancers (Basel). 2020;12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 21] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 66. | Lin X, Huang C, Chen Z, Wang H, Zeng Y. CircRNA_100876 Is Upregulated in Gastric Cancer (GC) and Promotes the GC Cells' Growth, Migration and Invasion via miR-665/YAP1 Signaling. Front Genet. 2020;11:546275. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 18] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 67. | Pu Z, Xu M, Yuan X, Xie H, Zhao J. Circular RNA circCUL3 Accelerates the Warburg Effect Progression of Gastric Cancer through Regulating the STAT3/HK2 Axis. Mol Ther Nucleic Acids. 2020;22:310-318. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 46] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 68. | Wang H, Sun G, Xu P, Lv J, Zhang X, Zhang L, Wang S, Cao J, Xia Y, Xuan Z, Li B, Huang X, Jiang T, Fang L, Xu Z. Circular RNA TMEM87A promotes cell proliferation and metastasis of gastric cancer by elevating ULK1 via sponging miR-142-5p. J Gastroenterol. 2021;56:125-138. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 30] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 69. | Fang J, Chen W, Meng X. Downregulating circRNA_0044516 Inhibits Cell Proliferation in Gastric Cancer Through miR-149/Wnt1/β-catenin Pathway. J Gastrointest Surg. 2021;25:1696-1705. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 13] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 70. | Fan D, Wang C, Wang D, Zhang N, Yi T. Circular RNA circ_0000039 enhances gastric cancer progression through miR-1292-5p/DEK axis. Cancer Biomark. 2021;30:167-177. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 12] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 71. | Ma C, Wang X, Yang F, Zang Y, Liu J, Xu X, Li W, Jia J, Liu Z. Circular RNA hsa_circ_0004872 inhibits gastric cancer progression via the miR-224/Smad4/ADAR1 successive regulatory circuit. Mol Cancer. 2020;19:157. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 49] [Cited by in RCA: 137] [Article Influence: 27.4] [Reference Citation Analysis (0)] |
| 72. | Peng Y, Wang HH. Cir-ITCH inhibits gastric cancer migration, invasion and proliferation by regulating the Wnt/β-catenin pathway. Sci Rep. 2020;10:17443. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 31] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 73. | Xia T, Pan Z, Zhang J. CircPDZD8 promotes gastric cancer progression by regulating CHD9 via sponging miR-197-5p. Aging (Albany NY). 2020;12:19352-19364. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 18] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 74. | Xu QY, Xie MJ, Huang J, Wang ZW. Effect of circ MTHFD2 on resistance to pemetrexed in gastric cancer through regulating expression of miR-124. Eur Rev Med Pharmacol Sci. 2019;23:10290-10299. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 15] [Reference Citation Analysis (0)] |
| 75. | Liu YY, Zhang LY, Du WZ. Circular RNA circ-PVT1 contributes to paclitaxel resistance of gastric cancer cells through the regulation of ZEB1 expression by sponging miR-124-3p. Biosci Rep. 2019;39. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 69] [Cited by in RCA: 104] [Article Influence: 20.8] [Reference Citation Analysis (0)] |
| 76. | Huang X, Li Z, Zhang Q, Wang W, Li B, Wang L, Xu Z, Zeng A, Zhang X, He Z, Li Q, Sun G, Wang S, Zhang L, Xu H. Circular RNA AKT3 upregulates PIK3R1 to enhance cisplatin resistance in gastric cancer via miR-198 suppression. Mol Cancer. 2019;18:71. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 291] [Cited by in RCA: 310] [Article Influence: 51.7] [Reference Citation Analysis (0)] |
| 77. | Huang XX, Zhang Q, Hu H, Jin Y, Zeng AL, Xia YB, Xu L. A novel circular RNA circFN1 enhances cisplatin resistance in gastric cancer via sponging miR-182-5p. J Cell Biochem. 2020;. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 46] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 78. | Hu J, Li P, Song Y, Ge YX, Meng XM, Huang C, Li J, Xu T. Progress and prospects of circular RNAs in Hepatocellular carcinoma: Novel insights into their function. J Cell Physiol. 2018;233:4408-4422. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 46] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 79. | Cheng F, Wang L, Zhang J. Circular RNA 0016788 displays as a biomarker for tumor progression and poor prognosis in surgical hepatocellular carcinoma patients. J Clin Lab Anal. 2020;34:e23300. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 15] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 80. | Yao Z, Luo J, Hu K, Lin J, Huang H, Wang Q, Zhang P, Xiong Z, He C, Huang Z, Liu B, Yang Y. ZKSCAN1 gene and its related circular RNA (circZKSCAN1) both inhibit hepatocellular carcinoma cell growth, migration, and invasion but through different signaling pathways. Mol Oncol. 2017;11:422-437. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 219] [Cited by in RCA: 263] [Article Influence: 32.9] [Reference Citation Analysis (0)] |
| 81. | Yang J, Li Y, Yu Z, Zhou Y, Tu J, Lou J, Wang Y. Circular RNA Circ100084 functions as sponge of miR23a5p to regulate IGF2 expression in hepatocellular carcinoma. Mol Med Rep. 2020;21:2395-2404. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 9] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 82. | Chen W, Quan Y, Fan S, Wang H, Liang J, Huang L, Chen L, Liu Q, He P, Ye Y. Exosome-transmitted circular RNA hsa_circ_0051443 suppresses hepatocellular carcinoma progression. Cancer Lett. 2020;475:119-128. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 121] [Cited by in RCA: 212] [Article Influence: 42.4] [Reference Citation Analysis (0)] |
| 83. | Fu Y, Cai L, Lei X, Wang D. Circular RNA ABCB10 promotes hepatocellular carcinoma progression by increasing HMG20A expression by sponging miR-670-3p. Cancer Cell Int. 2019;19:338. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 25] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 84. | Zhou S, Wei J, Wang Y, Liu X. Cisplatin resistance-associated circRNA_101237 serves as a prognostic biomarker in hepatocellular carcinoma. Exp Ther Med. 2020;19:2733-2740. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 21] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 85. | Wu MY, Tang YP, Liu JJ, Liang R, Luo XL. Global transcriptomic study of circRNAs expression profile in sorafenib resistant hepatocellular carcinoma cells. J Cancer. 2020;11:2993-3001. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 18] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 86. | Chen H, Liu S, Li M, Huang P, Li X. circ_0003418 Inhibits Tumorigenesis And Cisplatin Chemoresistance Through Wnt/β-Catenin Pathway In Hepatocellular Carcinoma. Onco Targets Ther. 2019;12:9539-9549. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 63] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 87. | Li J, Qin X, Wu R, Wan L, Zhang L, Liu R. Circular RNA circFBXO11 modulates hepatocellular carcinoma progress and oxaliplatin resistance through miR-605/FOXO3/ABCB1 axis. J Cell Mol Med. 2020;24:5152-5161. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 52] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 88. | Sun P, Fan X, Hu X, Fu X, Wei Q, Zang Y. circPCNX and Pecanex Promote Hepatocellular Carcinoma Cell Viability by Inhibiting miR-506. Cancer Manag Res. 2019;11:10957-10967. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 89. | Rawla P, Sunkara T, Barsouk A. Epidemiology of colorectal cancer: incidence, mortality, survival, and risk factors. Prz Gastroenterol. 2019;14:89-103. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 444] [Cited by in RCA: 1059] [Article Influence: 176.5] [Reference Citation Analysis (1)] |
| 90. | Karimzadeh MR, Zarin M, Ehtesham N, Khosravi S, Soosanabadi M, Mosallaei M, Pourdavoud P. MicroRNA binding site polymorphism in inflammatory genes associated with colorectal cancer: literature review and bioinformatics analysis. Cancer Gene Ther. 2020;27:739-753. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 91. | Chen RX, Chen X, Xia LP, Zhang JX, Pan ZZ, Ma XD, Han K, Chen JW, Judde JG, Deas O, Wang F, Ma NF, Guan X, Yun JP, Wang FW, Xu RH, Dan Xie. N6-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis. Nat Commun. 2019;10:4695. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 239] [Cited by in RCA: 515] [Article Influence: 85.8] [Reference Citation Analysis (0)] |
| 92. | Chen Z, Ren R, Wan D, Wang Y, Xue X, Jiang M, Shen J, Han Y, Liu F, Shi J, Kuang Y, Li W, Zhi Q. Hsa_circ_101555 functions as a competing endogenous RNA of miR-597-5p to promote colorectal cancer progression. Oncogene. 2019;38:6017-6034. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 72] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 93. | Zeng K, Chen X, Xu M, Liu X, Hu X, Xu T, Sun H, Pan Y, He B, Wang S. CircHIPK3 promotes colorectal cancer growth and metastasis by sponging miR-7. Cell Death Dis. 2018;9:417. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 447] [Cited by in RCA: 488] [Article Influence: 69.7] [Reference Citation Analysis (0)] |
| 94. | Hsiao KY, Lin YC, Gupta SK, Chang N, Yen L, Sun HS, Tsai SJ. Noncoding Effects of Circular RNA CCDC66 Promote Colon Cancer Growth and Metastasis. Cancer Res. 2017;77:2339-2350. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 416] [Cited by in RCA: 519] [Article Influence: 64.9] [Reference Citation Analysis (0)] |
| 95. | Lin YC, Yu YS, Lin HH, Hsiao KY. Oxaliplatin-Induced DHX9 Phosphorylation Promotes Oncogenic Circular RNA CCDC66 Expression and Development of Chemoresistance. Cancers (Basel). 2020;12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 32] [Cited by in RCA: 49] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 96. | Guo JN, Li J, Zhu CL, Feng WT, Shao JX, Wan L, Huang MD, He JD. Comprehensive profile of differentially expressed circular RNAs reveals that hsa_circ_0000069 is upregulated and promotes cell proliferation, migration, and invasion in colorectal cancer. Onco Targets Ther. 2016;9:7451-7458. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 92] [Cited by in RCA: 129] [Article Influence: 14.3] [Reference Citation Analysis (0)] |
| 97. | Fujita K, Kubota Y, Ishida H, Sasaki Y. Irinotecan, a key chemotherapeutic drug for metastatic colorectal cancer. World J Gastroenterol. 2015;21:12234-12248. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 185] [Cited by in RCA: 225] [Article Influence: 22.5] [Reference Citation Analysis (2)] |
| 98. | Jian X, He H, Zhu J, Zhang Q, Zheng Z, Liang X, Chen L, Yang M, Peng K, Zhang Z, Liu T, Ye Y, Jiao H, Wang S, Zhou W, Ding Y, Li T. Hsa_circ_001680 affects the proliferation and migration of CRC and mediates its chemoresistance by regulating BMI1 through miR-340. Mol Cancer. 2020;19:20. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 124] [Cited by in RCA: 148] [Article Influence: 29.6] [Reference Citation Analysis (0)] |
| 99. | Wang X, Zhang H, Yang H, Bai M, Ning T, Deng T, Liu R, Fan Q, Zhu K, Li J, Zhan Y, Ying G, Ba Y. Exosome-delivered circRNA promotes glycolysis to induce chemoresistance through the miR-122-PKM2 axis in colorectal cancer. Mol Oncol. 2020;14:539-555. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 294] [Cited by in RCA: 397] [Article Influence: 79.4] [Reference Citation Analysis (0)] |
| 100. | Hon KW, Ab-Mutalib NS, Abdullah NMA, Jamal R, Abu N. Extracellular Vesicle-derived circular RNAs confers chemoresistance in Colorectal cancer. Sci Rep. 2019;9:16497. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 91] [Cited by in RCA: 107] [Article Influence: 17.8] [Reference Citation Analysis (0)] |
| 101. | Abu N, Hon KW, Jeyaraman S, Yahaya A, Abdullah NM, Mustangin M, Sulaiman SA, Jamal R, Ab-Mutalib NS. Identification of differentially expressed circular RNAs in chemoresistant colorectal cancer. Epigenomics. 2019;11:875-884. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 43] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 102. | Ehrhart IC, Parker PE, Weidner WJ, Dabney JM, Scott JB, Haddy FJ. Coronary vascular and myocardial responses to carotid body stimulation in the dog. Am J Physiol. 1975;229:754-760. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 27] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 103. | Lai M, Liu G, Li R, Bai H, Zhao J, Xiao P, Mei J. Hsa_circ_0079662 induces the resistance mechanism of the chemotherapy drug oxaliplatin through the TNF-α pathway in human colon cancer. J Cell Mol Med. 2020;24:5021-5027. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 44] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 104. | Ren TJ, Liu C, Hou JF, Shan FX. CircDDX17 reduces 5-fluorouracil resistance and hinders tumorigenesis in colorectal cancer by regulating miR-31-5p/KANK1 axis. Eur Rev Med Pharmacol Sci. 2020;24:1743-1754. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 25] [Reference Citation Analysis (0)] |
| 105. | Naeli P, Pourhanifeh MH, Karimzadeh MR, Shabaninejad Z, Movahedpour A, Tarrahimofrad H, Mirzaei HR, Bafrani HH, Savardashtaki A, Mirzaei H, Hamblin MR. Circular RNAs and gastrointestinal cancers: Epigenetic regulators with a prognostic and therapeutic role. Crit Rev Oncol Hematol. 2020;145:102854. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 127] [Cited by in RCA: 120] [Article Influence: 24.0] [Reference Citation Analysis (0)] |
| 106. | Fan L, Cao Q, Liu J, Zhang J, Li B. Circular RNA profiling and its potential for esophageal squamous cell cancer diagnosis and prognosis. Mol Cancer. 2019;18:16. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 72] [Cited by in RCA: 130] [Article Influence: 21.7] [Reference Citation Analysis (0)] |
| 107. | Huang E, Fu J, Yu Q, Xie P, Yang Z, Ji H, Wang L, Luo G, Zhang Y, Li K. CircRNA hsa_circ_0004771 promotes esophageal squamous cell cancer progression via miR-339-5p/CDC25A axis. Epigenomics. 2020;12:587-603. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 53] [Article Influence: 10.6] [Reference Citation Analysis (0)] |
| 108. | Wu Y, Zhi L, Zhao Y, Yang L, Cai F. Knockdown of circular RNA UBAP2 inhibits the malignant behaviours of esophageal squamous cell carcinoma by microRNA-422a/Rab10 axis. Clin Exp Pharmacol Physiol. 2020;47:1283-1290. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 22] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 109. | Pan Z, Lin J, Wu D, He X, Wang W, Hu X, Zhang L, Wang M. Hsa_circ_0006948 enhances cancer progression and epithelial-mesenchymal transition through the miR-490-3p/HMGA2 axis in esophageal squamous cell carcinoma. Aging (Albany NY). 2019;11:11937-11954. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 71] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 110. | Xu Z, Tie X, Li N, Yi Z, Shen F, Zhang Y. Circular RNA hsa_circ_0000654 promotes esophageal squamous cell carcinoma progression by regulating the miR-149-5p/IL-6/STAT3 pathway. IUBMB Life. 2020;72:426-439. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 52] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 111. | Hu X, Wu D, He X, Zhao H, He Z, Lin J, Wang K, Wang W, Pan Z, Lin H, Wang M. circGSK3β promotes metastasis in esophageal squamous cell carcinoma by augmenting β-catenin signaling. Mol Cancer. 2019;18:160. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 63] [Cited by in RCA: 135] [Article Influence: 22.5] [Reference Citation Analysis (0)] |
| 112. | Xing Y, Zha WJ, Li XM, Li H, Gao F, Ye T, Du WQ, Liu YC. Circular RNA circ-Foxo3 inhibits esophageal squamous cell cancer progression via the miR-23a/PTEN axis. J Cell Biochem. 2020;121:2595-2605. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 70] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 113. | Zhong R, Chen Z, Mo T, Li Z, Zhang P. Potential Role of circPVT1 as a proliferative factor and treatment target in esophageal carcinoma. Cancer Cell Int. 2019;19:267. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 44] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 114. | Meng L, Liu S, Ding P, Chang S, Sang M. Circular RNA ciRS-7 inhibits autophagy of ESCC cells by functioning as miR-1299 sponge to target EGFR signaling. J Cell Biochem. 2020;121:1039-1049. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 49] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 115. | Huang H, Wei L, Qin T, Yang N, Li Z, Xu Z. Circular RNA ciRS-7 triggers the migration and invasion of esophageal squamous cell carcinoma via miR-7/KLF4 and NF-κB signals. Cancer Biol Ther. 2019;20:73-80. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 80] [Cited by in RCA: 109] [Article Influence: 15.6] [Reference Citation Analysis (0)] |
| 116. | Mi L, Zhou Y, Wu D, Tao Q, Wang X, Zhu H, Gao X, Wang J, Ling R, Deng J, Mao C, Chen D. ACSS2/AMPK/PCNA pathwaydriven proliferation and chemoresistance of esophageal squamous carcinoma cells under nutrient stress. Mol Med Rep. 2019;20:5286-5296. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 8] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 117. | Zou FW, Yang SZ, Li WY, Liu CY, Liu XH, Hu CH, Liu ZH, Xu S. circRNA_001275 upregulates Wnt7a expression by competitively sponging miR3703p to promote cisplatin resistance in esophageal cancer. Int J Oncol. 2020;57:151-160. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 21] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 118. | Arnold M, Abnet CC, Neale RE, Vignat J, Giovannucci EL, McGlynn KA, Bray F. Global Burden of 5 Major Types of Gastrointestinal Cancer. Gastroenterology. 2020;159:335-349.e15. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 857] [Cited by in RCA: 1211] [Article Influence: 242.2] [Reference Citation Analysis (0)] |
| 119. | Guo X, Zhou Q, Su D, Luo Y, Fu Z, Huang L, Li Z, Jiang D, Kong Y, Chen R, Chen C. Circular RNA circBFAR promotes the progression of pancreatic ductal adenocarcinoma via the miR-34b-5p/MET/Akt axis. Mol Cancer. 2020;19:83. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 56] [Cited by in RCA: 90] [Article Influence: 18.0] [Reference Citation Analysis (0)] |
| 120. | Wong CH, Lou UK, Li Y, Chan SL, Tong JH, To KF, Chen Y. CircFOXK2 Promotes Growth and Metastasis of Pancreatic Ductal Adenocarcinoma by Complexing with RNA-Binding Proteins and Sponging MiR-942. Cancer Res. 2020;80:2138-2149. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 58] [Cited by in RCA: 108] [Article Influence: 21.6] [Reference Citation Analysis (0)] |
| 121. | Xing C, Ye H, Wang W, Sun M, Zhang J, Zhao Z, Jiang G. Circular RNA ADAM9 facilitates the malignant behaviours of pancreatic cancer by sponging miR-217 and upregulating PRSS3 expression. Artif Cells Nanomed Biotechnol. 2019;47:3920-3928. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 42] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 122. | Chen Y, Li Z, Zhang M, Wang B, Ye J, Zhang Y, Tang D, Ma D, Jin W, Li X, Wang S. Circ-ASH2L promotes tumor progression by sponging miR-34a to regulate Notch1 in pancreatic ductal adenocarcinoma. J Exp Clin Cancer Res. 2019;38:466. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 49] [Cited by in RCA: 72] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 123. | Yao J, Zhang C, Chen Y, Gao S. Downregulation of circular RNA circ-LDLRAD3 suppresses pancreatic cancer progression through miR-137-3p/PTN axis. Life Sci. 2019;239:116871. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 58] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 124. | Ou ZL, Luo Z, Wei W, Liang S, Gao TL, Lu YB. Hypoxia-induced shedding of MICA and HIF1A-mediated immune escape of pancreatic cancer cells from NK cells: role of circ_0000977/miR-153 axis. RNA Biol. 2019;16:1592-1603. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 75] [Cited by in RCA: 131] [Article Influence: 21.8] [Reference Citation Analysis (0)] |
| 125. | Yang J, Cong X, Ren M, Sun H, Liu T, Chen G, Wang Q, Li Z, Yu S, Yang Q. Circular RNA hsa_circRNA_0007334 is Predicted to Promote MMP7 and COL1A1 Expression by Functioning as a miRNA Sponge in Pancreatic Ductal Adenocarcinoma. J Oncol. 2019;2019:7630894. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 56] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 126. | Liu L, Liu FB, Huang M, Xie K, Xie QS, Liu CH, Shen MJ, Huang Q. Circular RNA ciRS-7 promotes the proliferation and metastasis of pancreatic cancer by regulating miR-7-mediated EGFR/STAT3 signaling pathway. Hepatobiliary Pancreat Dis Int. 2019;18:580-586. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 78] [Cited by in RCA: 110] [Article Influence: 18.3] [Reference Citation Analysis (0)] |
| 127. | Xu Y, Yao Y, Gao P, Cui Y. Upregulated circular RNA circ_0030235 predicts unfavorable prognosis in pancreatic ductal adenocarcinoma and facilitates cell progression by sponging miR-1253 and miR-1294. Biochem Biophys Res Commun. 2019;509:138-142. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 73] [Article Influence: 12.2] [Reference Citation Analysis (0)] |
| 128. | An Y, Cai H, Zhang Y, Liu S, Duan Y, Sun D, Chen X, He X. circZMYM2 Competed Endogenously with miR-335-5p to Regulate JMJD2C in Pancreatic Cancer. Cell Physiol Biochem. 2018;51:2224-2236. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 61] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 129. | Hao L, Rong W, Bai L, Cui H, Zhang S, Li Y, Chen D, Meng X. Upregulated circular RNA circ_0007534 indicates an unfavorable prognosis in pancreatic ductal adenocarcinoma and regulates cell proliferation, apoptosis, and invasion by sponging miR-625 and miR-892b. J Cell Biochem. 2019;120:3780-3789. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 84] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 130. | Li J, Li Z, Jiang P, Peng M, Zhang X, Chen K, Liu H, Bi H, Liu X, Li X. Circular RNA IARS (circ-IARS) secreted by pancreatic cancer cells and located within exosomes regulates endothelial monolayer permeability to promote tumor metastasis. J Exp Clin Cancer Res. 2018;37:177. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 179] [Cited by in RCA: 292] [Article Influence: 41.7] [Reference Citation Analysis (0)] |
| 131. | Li Z, Yanfang W, Li J, Jiang P, Peng T, Chen K, Zhao X, Zhang Y, Zhen P, Zhu J, Li X. Tumor-released exosomal circular RNA PDE8A promotes invasive growth via the miR-338/MACC1/MET pathway in pancreatic cancer. Cancer Lett. 2018;432:237-250. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 187] [Cited by in RCA: 269] [Article Influence: 38.4] [Reference Citation Analysis (0)] |
| 132. | Shao F, Huang M, Meng F, Huang Q. Circular RNA Signature Predicts Gemcitabine Resistance of Pancreatic Ductal Adenocarcinoma. Front Pharmacol. 2018;9:584. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 56] [Cited by in RCA: 73] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 133. | Liu Y, Xia L, Dong L, Wang J, Xiao Q, Yu X, Zhu H. CircHIPK3 Promotes Gemcitabine (GEM) Resistance in Pancreatic Cancer Cells by Sponging miR-330-5p and Targets RASSF1. Cancer Manag Res. 2020;12:921-929. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 73] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 134. | Zhang HD, Jiang LH, Sun DW, Hou JC, Ji ZL. CircRNA: a novel type of biomarker for cancer. Breast Cancer. 2018;25:1-7. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 422] [Cited by in RCA: 593] [Article Influence: 74.1] [Reference Citation Analysis (0)] |