Published online Jun 14, 2013. doi: 10.3748/wjg.v19.i22.3404
Revised: January 1, 2013
Accepted: February 2, 2013
Published online: June 14, 2013
Processing time: 202 Days and 3.3 Hours
AIM: To study the interplay between butyrate concentration and butyrate-producing bacteria in fecal samples of ulcerative colitis (UC) patients vs control individuals.
METHODS: Fecal samples were collected from 14 control individuals (hemorrhoid patients only) and 26 UC patients (severe: n = 12, moderate: n = 6, remission: n = 8), recruited by the gastroenterologist at the Department of Gastroenterology, All India Institute of Medical Sciences, New Delhi, India. Disease activity in UC patients was determined by clinical colitis activity index. We employed fluorescent in situ hybridization in combination with flow cytometry to enumerate the clostridium cluster population targeted by 16S rRNA gene probe. Major butyrate-producing species within this cluster were quantified to see if any change existed in control vs UC patients with different disease activity. This observed change was further validated by quantitative polymerase chain reaction. In addition to this, we carried out gas chromatography to evaluate the changes in concentration of major short chain fatty acids (SCFAs), namely acetate, n-butyrate, iso-butyrate, in the above samples. Student t test and Graph pad prism-6 were used to compare the data statistically.
RESULTS: There was a significant decrease of Clostridium coccoides (control, 25.69% ± 1.62% vs severe, 9.8% ± 2.4%, P = 0.0001) and Clostridium leptum clusters (control, 13.74% ± 1.05% vs severe, 6.2% ± 1.8%, P = 0.0001) in fecal samples of UC patients. Furthermore, we demonstrated that some butyrate-producing members of the clostridial cluster, like Fecalibacterium prausnitzii (control, 11.66% ± 1.55% vs severe, 6.01% ± 1.6%, P = 0.0001) and Roseburia intestinalis (control, 14.48% ± 1.52% vs severe, 9% ± 1.83%, P = 0.02) were differentially present in patients with different disease activity. In addition, we also demonstrated decreased concentrations of fecal SCFAs, especially of n-butyrate (control, 24.32 ± 1.86 mmol/μL vs severe, 12.74 ± 2.75 mmol/μL, P = 0.003), iso-butyrate (control, 1.70 ± 0.41 mmol/μL vs severe, 0.68 ± 0.24 mmol/μL, P = 0.0441) and acetate (control, 39.51 ± 1.76 mmol/μL vs severe, 32.12 ± 2.95 mmol/μL, P = 0.047), in the fecal samples of UC patients. The observed decrease of predominant butyrate producers of clostridial clusters correlated with the reduced SCFA levels in active UC patients. This was further confirmed by the restoration in the population of some butyrate producers with simultaneous increase in the level of SCFA in remission samples.
CONCLUSION: Our observations indicate that decreases in members of the clostridial cluster resulting in reduced butyrate levels contribute to the etiology of UC.
- Citation: Kumari R, Ahuja V, Paul J. Fluctuations in butyrate-producing bacteria in ulcerative colitis patients of North India. World J Gastroenterol 2013; 19(22): 3404-3414
- URL: https://www.wjgnet.com/1007-9327/full/v19/i22/3404.htm
- DOI: https://dx.doi.org/10.3748/wjg.v19.i22.3404
Inflammatory bowel disease (IBD), comprising of Crohn’s disease and ulcerative colitis (UC), is a class of chronic inflammatory disorders of the intestine. An increasing trend in the incidence and prevalence of IBD in the Asian population has been recognized for the past two decades[1]. The dynamic balance between commensal microbiota and host defensive responses at the mucosal frontier has a pivotal role in the initiation and pathogenesis of chronic IBD[2]. Whether the exaggerated immune response is exerted to all commensal bacteria, or to a subset or a single strain of bacteria, is not known[3]. Differences in fecal microbiota of healthy subjects and IBD patients have been enumerated using different techniques[4-6]. Impaired cellular metabolism, such as butyrate oxidation and short chain fatty acid (SCFA) fermentation, has shown strong association with altered gut microbiota in UC patients[7,8].
SCFAs, such as acetate, propionate and butyrate, are produced by intestinal microbial fermentation of mainly undigested dietary carbohydrates, specifically resistant starches and dietary fiber, but also in a minor part generated by dietary and endogenous proteins in the intestine[9]. SCFAs are important for normal intestinal biology[10]. They also stimulate colonic sodium and fluid absorption and exert proliferative effects on the colonocytes[11]. Therefore, monitoring the fluctuations in SCFA concentration may help in understanding the relation of dysbiosis with UC.
Members of Clostridium leptum (C. leptum) and Clostridium coccoides (C. coccoides) groups together constitute the majority of Firmicutes (low G + C content bacteria)[12], producing large amounts of butyrate that function as an energy source for colonic epithelial cells and inhibit mRNA expression of proinflammatory cytokines in the mucosa by inhibiting nuclear factor-κB (NF-κB) activation[13]. Butyrate has been reported to help in prevention of colorectal cancer[14]. Evidence indicates that bacteria related to Eubacterium hallii (E. hallii), Roseburia species and Eubacterium rectale (E. rectale) within cluster XIVa and Faecalibacterium prausnitzii (F. prausnitzii)-related bacteria within cluster IV are normally the two most abundant groups of human fecal bacteria that produce butyrate[15]. These species-level probes account for a mean of 7.7% of the population of the total human fecal microbiota[16]. However, both the clostridial clusters harbor a diverse collection of several species of butyrate producers and non-butyrate producers as well[17].
Dynamics of predominant butyrate producers at the species level during disease activity and their correlation with the fluctuation in SCFA have not been established clearly. Here we explored the alteration in population of dominant butyrate-producing bacterial species present in fecal samples of UC patients at different disease stages. Abundance of the butyrate-producing clostridial cluster group was estimated by fluorescent in situ hybridization (FISH) in combination with flow cytometry and real time polymerase chain reaction (PCR). Further enumeration was carried out for known predominant members of butyrate-producing bacteria such as (1) F. prausnitzii as a member of the C. leptum group; and (2) E. hallii and Roseburia intestinalis (R. intestinalis) as members of the C. coccoides group. We also checked the level of butyrate, iso-butyrate and acetate in fecal samples to see if any correlation exists with butyrate-producing bacteria during different disease conditions.
Disease activity in UC patients was determined by simple clinical colitis activity index[18] and the patients with total index score of 7-10, > 10 and 0-2 were assigned moderate, severe and remission category, respectively[19]. The patients were recruited by the gastroenterologist at the Department of Gastroenterology of the All India Institute of Medical Sciences, New Delhi, India. Clinical and demographic features of UC patients and controls are shown in Table 1. Patients who had hemorrhoids only and showed no evidence of small and large intestinal disease were enrolled as control individuals in this study. No study patient had received any antibiotic treatment in the past three months before sample collection. Patients under any antibiotic or probiotic treatments were excluded from this study. The fecal samples were collected in sterile stool specimen containers and stored at -80 °C within 3 h of sample collection until further processing.
| Feature | UC (n = 26) | Control (n = 14) |
| Sex (F/M) | 11 (42.30)/15 (57.69) | 7 (50)/7(50) |
| Age at diagnosis (yr), mean ± SD | 38.35 ± 11.49 | 35.00 ± 14.14 |
| 15-40 | 18 (69.23) | 8 (57.14) |
| > 40 | 8 (30.76) | 6 (42.85) |
| Disease behavior | ||
| Severe | 12 (46.15) | - |
| Moderate | 6 (23.07) | - |
| Remission | 8 (30.76) | - |
| Disease extent | ||
| Proctitis | 4 (15.38) | - |
| Left sided colitis | 6 (23.00) | - |
| Pancolitis | 6 (23.00) | - |
| None of the above | 10 (38.46) | - |
| Smoking history | ||
| Yes | 3 (11.53) | 2 (14.28) |
| No | 23 (88.46) | 12 (85.70) |
| Treatment history | ||
| Immunosuppressant | 16 (61.53) | - |
| Steroids | 14 (53.84) | - |
| Appendectomy Y/N | 4 (15.38)/22 (84.61) | 0/14 |
| Family history Y/N | 2 (7.69)/24 (92.30) | 0/14 |
All probes were designed from the 16S rRNA gene. EUB 338 conserved within the bacterial domain was used as a positive control probe[20]; conversely NONEUB338 (reverse of EUB)[21] was used as a negative control probe. The positive control probe was double labeled with fluorescein isothiocyanate (FITC) at both the 5’ and 3’ end[22], whereas the negative control probe was labeled with FITC at the 5’ end and Cy5 at the 3’ end. We used two group-specific probes labeled with Cy5 at their 5’ end (Sigma, India) and three species-specific probes labeled with FITC at their 5’ end (Sigma, India). Two competitors (unlabeled) and 5 helper oligonucleotides (unlabeled) were used to increase the accessibility of the Clep 1156 probe[23] (Table 2).
| Primer or probe | Target (phylogenetic group) | Sequence (5’-3’) from 16S rRNA gene | Used in FISH-flow cytometry or qPCR | Ref. |
| NON338 | No bacteria | ACATCCTACGGGAGGC | Probe1 | [21] |
| EUB338 | Most bacteria | GCTGCCTCCCGTAGGAGT | Probe1 | [20] |
| Erec 482 | C. coccoides/E.rectale cluster | GCTTCTTAGTCARGTACCG | Probe1 | [34] |
| Clep 1156 | Clostridium leptum subgroup | GTTTTRTCAACGGCAGTC | Probe1 | [35] |
| Fpra-655 | F. prausnitzii | CGCCTACCTCTGCACTAC | Probe1 | [16] |
| Rint-623 | R. intestinalis subcluster | TTCCAATGCAGTACCGGG | Probe1 | [16] |
| Ehal-057 | E. halliii L2-7/E. halliii | TTGCACTGCCACCTACGC | Probe1 | [16] |
| Cp1 | Competitor 1 | GRTTTRTCAYCGGCAGTC | Competitor1 | [23] |
| Cp2 | Competitor 1 | GTVTTRTCBACGGCAGTC | Competitor1 | [23] |
| H1174 | Helper oligonucleotide | TTGACGTCRTCCCCACCTTCCTCC | Helper1 | [23] |
| H1129 | Helper oligonucleotide | TAGAGTGMTCTTGCGTA | Helper1 | [23] |
| H1090 | Helper oligonucleotide | GGTTGCGCTCGTTGCGGGACTTAA | Helper1 | [23] |
| H750 | Helper oligonucleotide | TCGHGCCTCAGCGTCAG | Helper1 | [23] |
| H122 | Helper oligonucleotide | GAAGGCAGGTTACTCACGC | Helper1 | [23] |
| Clep FP | C. leptum subgroup | CGTCAGCTCGTGTCGTGAGAT | Primer set2 | [36] |
| Clep RP | CGTCATCCCCACCTTCCTCC | |||
| C.cocci FP | C. coccoides subgroup | GCCACATTGGGACTGAGA | Primer set2 | [36] |
| C.cocci RP | GCTTCTTAGTCAGGTACCG | |||
| Fpraus FP | F. prausnitzii | GATGGCCTCGCGTCCGATTAG | Primer set2 | [37] |
| Fpraus RP | CCGAAGACCTTCTTCCTC | |||
| Rint FP | Roseburia/E. rectale cluster | CKGCAAGTCTGATGTGAAAG | Primer set2 | This study |
| Rint RP | GCGGGTCCCCGTCAATTCC | |||
| Ehal FP | E. hallii L2-7/E. hallii members | GCGTAGGTGGCAGTGCAA | Primer set2 | [38] |
| Ehal RP | GCACCGRAGCCTATACGG |
About 1 g of fecal sample was suspended in 9 mL of phosphate buffered saline (PBS) and vortexed with 10-15 glass beads for 5 min to homogenize the sample. This suspension was centrifuged at 1500 rpm for 1 min to pellet down the debris and the supernatant was collected. To fix the cells, 1 mL of this supernatant was incubated with 4% paraformaldehyde (1:3 ratio) at 4 °C, overnight. The fixed cells were washed twice with PBS and incubated in ethanol-PBS solution (1:1 ratio) at -20 °C for 2 h. For each hybridization reaction, 60 μL of fixed cells were used. The fixed cells were washed twice with PBS and resuspended in 50 μL hybridization buffer (900 mmol/L NaCl, 20 mmol/L Tris-HCl of pH 8, and 0.01% sodium dodecyl sulfate at pH 7.2). All hybridizations were performed in the dark at 50 °C for 16 h in the hybridization solution containing 4 ng/μL of the appropriate labeled probe. One hundred and fifty microliter of hybridization solution (without probe) was added to stop the reaction and cells were pelleted at 1610 g for 10 min. Hybridized cells were further resuspended in prewarmed washing buffer [65 mmol/L NaCl, 20 mmol/L Tris-HCl, 5 mmol/L diaminoethanetetraacetic acid (EDTA), 0.01% sodium dodecyl sulfate, pH 7.2] and incubated at 50 °C for 20 min to remove non-specific binding of the probe. Finally, cells were pelleted down at 6000 rpm for 10 min and suspended in 200 μL PBS[5,24,25]. An aliquot of 100 μL was added to 0.5 mL of flow sheath solution (Becton Dickinson) for flow cytometry analysis.
Data acquisition was performed with a FACS calibur flow cytometer (Becton Dickinson) which is equipped with an air-cooled argon-ion laser providing 15 mW at 488 nm light combined with a 635 nm red-diode laser. The 488 nm laser was used to measure the forward angle light scatter (FSC, in the 488/10 nm band pass filter), the side angle light scatter (SSC, in the 488/10 nm band pass) and the green fluorescence intensity conferred by FITC labeled probes (filter 1 in the 530/30 nm band pass filter). The red-diode laser was used to detect the red fluorescence conferred by Cy5 labeled probes (filter 4 in a 661/16 nm band pass filter). The acquisition threshold was set in the side scatter channel. All the parameters were collected as logarithmic signals. The rate of events in the flow was set at low (12 μL/s). A total of 25000 events were collected and subsequent analyses were conducted using the Cell Quest Software (Becton Dickinson).
Enumeration of bacterial groups was performed by a double staining method in the same reaction tube where the hybridization of the EUB338 probe labeled with FITC and the genus-specific probe labeled with Cy5 were combined. This led us to estimate the abundance of bacterial groups targeted by the respective Cy5-labeled probe as a proportion of total bacteria labeled with the EUB338 FITC probe. Next, the abundance of known butyrate producers was enumerated as cells hybridized with the FITC-labeled species-specific probe as a proportion of total cells hybridized with the respective genus-specific Cy5-labeled probe. Each time, the proportion of hybridized bacteria was corrected by subtracting the background fluorescence obtained with hybridization of the negative control probe NONEUB338.
Genomic DNA from human fecal samples (220 mg) was extracted using the Qiagen stool DNA kit and eluted in 50 μL of Tris-EDTA buffer. About 20 ng of DNA from each sample was used to analyze the bacterial population. All primer sets used in the study were designed from the 16S rRNA gene as shown in Table 2. Genus-specific primers were used to amplify respective genus and species from genomic DNA of the fecal samples of healthy individuals. The amplified product was cloned and sequenced and sequences were deposited in the EMBL database to obtain the accession numbers (Table 3). These 16S rRNA gene fragments containing plasmids were used as reference strains. The standard curves were constructed by serial dilutions of each reference clone prepared from 0.05 to 500000 pg/tube, corresponding to 1 × 10 to 1 × 107 copy numbers. The standard curve of the reference clones was used to extrapolate the numbers of bacteria present in the fecal samples. With the molecular mass of the plasmid and insert known, the copy number was calculated as follows: mass in Daltons (g/molecule) = [size of double-stranded (ds) product in base pairs (bp)] (330 Da × 2 nucleotides (nt)/bp]/Avogadro’s number.
| Bacteria | Source | Accession No. |
| C. leptum | Healthy human fecal sample | AM042697 |
| F. prausnitzii | Healthy human fecal sample | JX556686 |
| R. intestinalis | Healthy human fecal sample | JX556688 |
| E. hallii | Healthy human fecal sample | JX556687 |
Thus, the precise number of molecules (molecules/μL) = Conc./mass in Daltons[26].
Fecal SCFAs were analyzed using gas chromatography/flame ionization detection (GC-FID). An aliquot of fecal content (250 mg) was extracted with 1 mL of extraction buffer [0.1% (w/v) HgCl2 and 1% (v/v) H3PO4] supplemented with 0.045 mg/mL 2,2-dimethylbutyrate (as internal standard). The resulting slurry was centrifuged for 30 min at 5000 g at 4 °C, and the supernatant was filtered through a 0.2-μm filter. SCFAs in the supernatant collected were analyzed using a GC (Shimadzu-2010) equipped with FID and a stabilwax column (Restek, United States) of 30 m length, 530 μm diameter and 1 μm film thickness. The system was run with nitrogen as carrier gas at an inlet constant pressure of 18.1 kPa. Samples were run at an initial temperature of 120 °C for 0.5 min, and then with 8 °C/min change in temperature till it reached 220 °C and was held at 220 °C for 8 min for a total program time of 20.5 min[27]. SCFAs were identified using external standards consisting of acetate, iso-butyrate, n-butyrate (Sigma, India) and the concentration was calculated using the area percentage method.
Ethical clearance for the study was obtained from the Institute Ethics Committee, All India Institute of Medical Sciences, New Delhi. Written informed consent was obtained from all the participants.
The mean cell proportion and number of bacteria in fecal samples were estimated by FISH and qPCR in triplicate, and the results were expressed as a percentage of bacteria and number of bacteria, respectively. SCFA level was determined by GC, and the results were expressed in mmol/μL. Student’s t test was employed to check any significant changes in the SCFA concentrations with the changes in the disease activity. Graphpad prism-6 was used to analyze FISH and qPCR data.
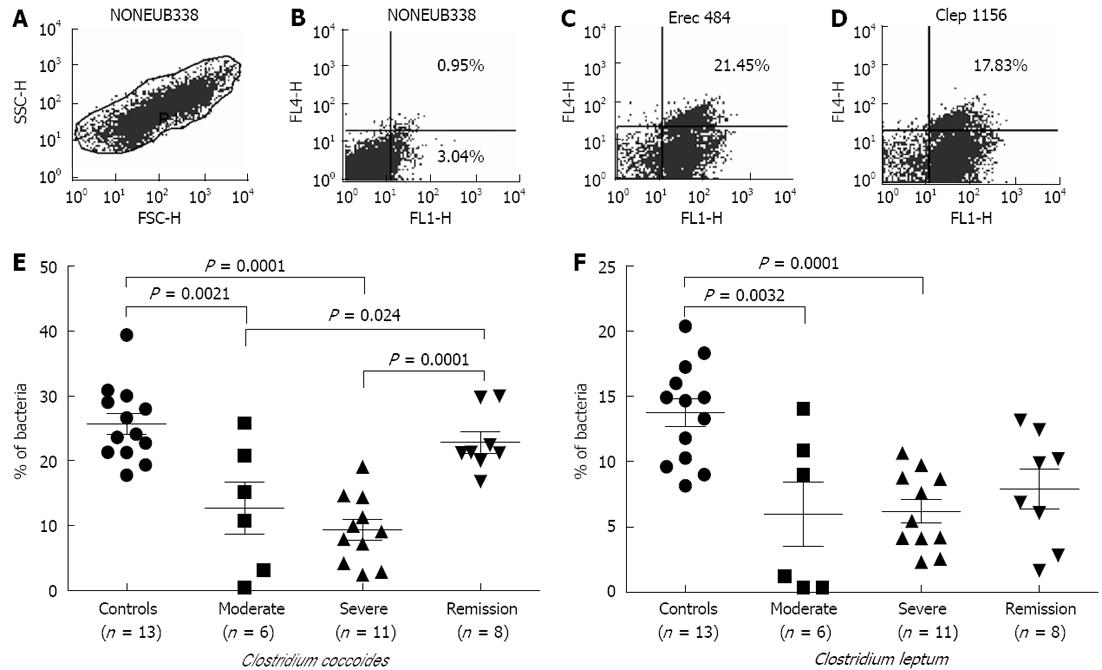
FISH-flow cytometry: In Figure 1A, the region R1 corresponds to relative size (FSC) with granularity (SSC) of the bacteria during flow cytometry with NONEUB338 hybridized cells. This region R1 was gated for further dot plots. Flow cytometric analysis of the hybridized samples gave a shift in signal of 1 log unit compared to the nonhybridized cells, enabling the specific detection and enumeration of the different bacterial groups (Figure 1B-D). Scoring of bacteria could not be achieved uniformly; therefore, sample size differed in each category.
In UC patients belonging to the moderate and severe disease categories, we observed significant decreases in members of the C. coccoides and C. leptum groups compared to control individuals (Figure 1E and F). Among the C. coccoides cluster, decreases in moderate disease samples attained the P value of 0.0021, while there was a P value of 0.0001 in the case of severe patient samples in comparison to controls. However, in the case of C. leptum, the P values were 0.0032 in moderate and 0.0001 in severe categories of samples, respectively, as compared to controls. Samples in the remission stage showed significant restoration in the population of the C. coccoides group (P = 0.0001); although an increasing trend was observed in the members of the C. leptum group, this did not attain a significant value.
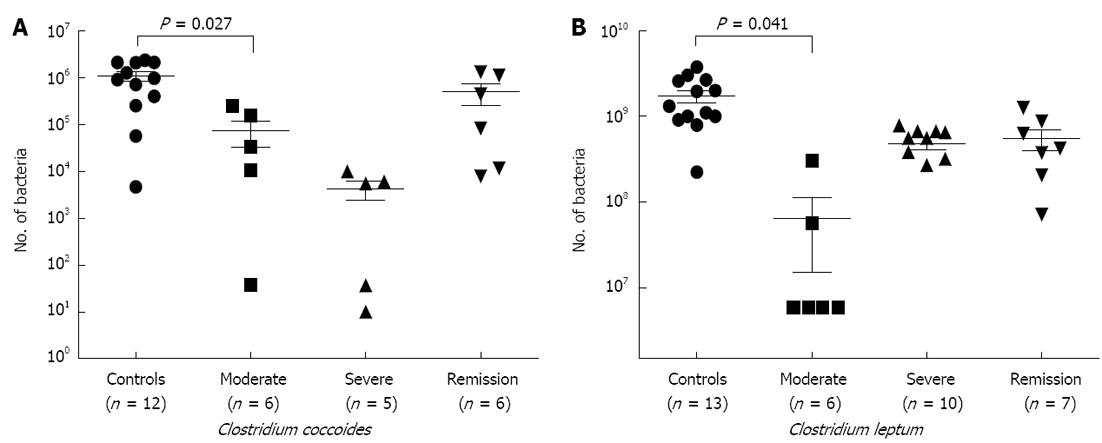
Quantitative polymerase chain reaction: In order to validate our FISH-flow cytometry data, we carried out a qPCR study (Figure 2A and B), where we observed significant decreases in both the members of C. coccoides group (P = 0.027) and C. leptum group (P = 0.041) in samples of moderate and severe disease stages. Members of both the clusters showed restoration of bacteria in samples of remission category; however, this did not attain a significant P value.
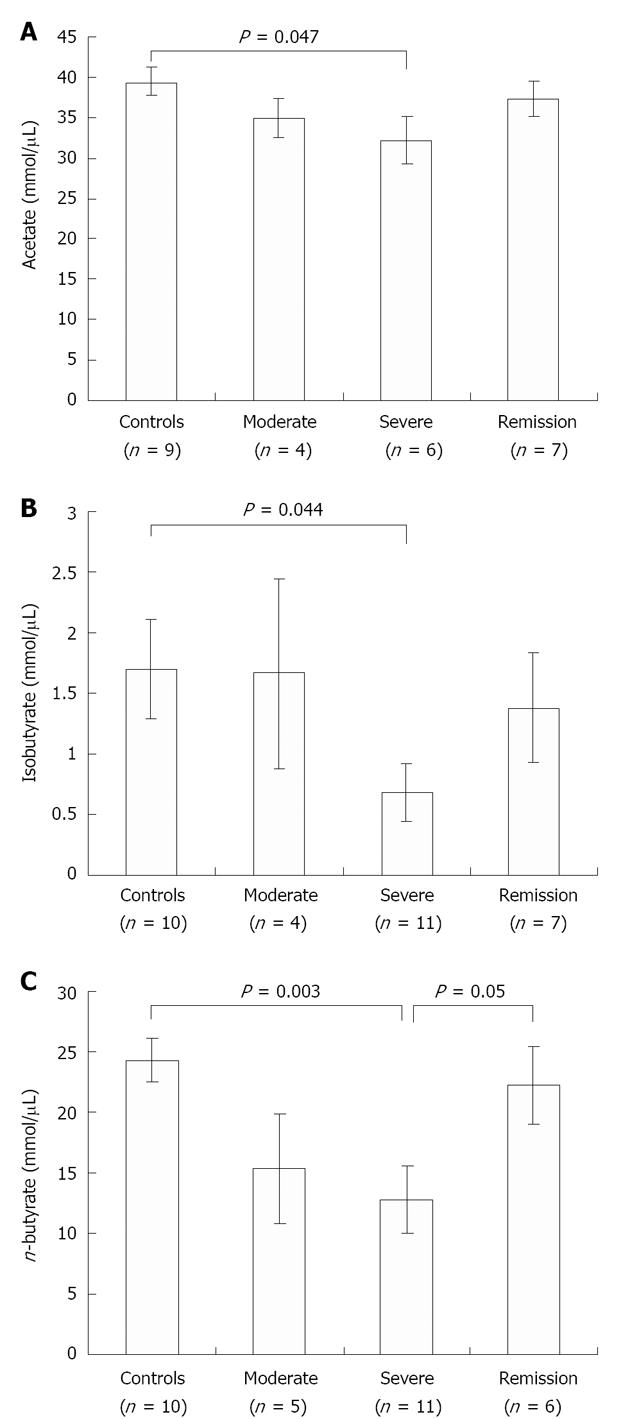
SCFA quantification by GC: We further quantified the change in concentration of fecal SCFAs, namely acetate, n-butyrate and iso-butyrate, in control vs UC patient samples by GC. The concentrations of butyrate (P = 0.003), iso-butyrate (P = 0.044) and acetate (P = 0.047), were significantly reduced in severe UC samples when compared with control samples (Figure 3). As expected, during the remission stage n-butyrate level significantly restored back to normal level (P = 0.05) as seen in control individuals, confirming that the decrease observed during disease conditions reflects the loss of butyrate-producing bacteria (Figure 3C).
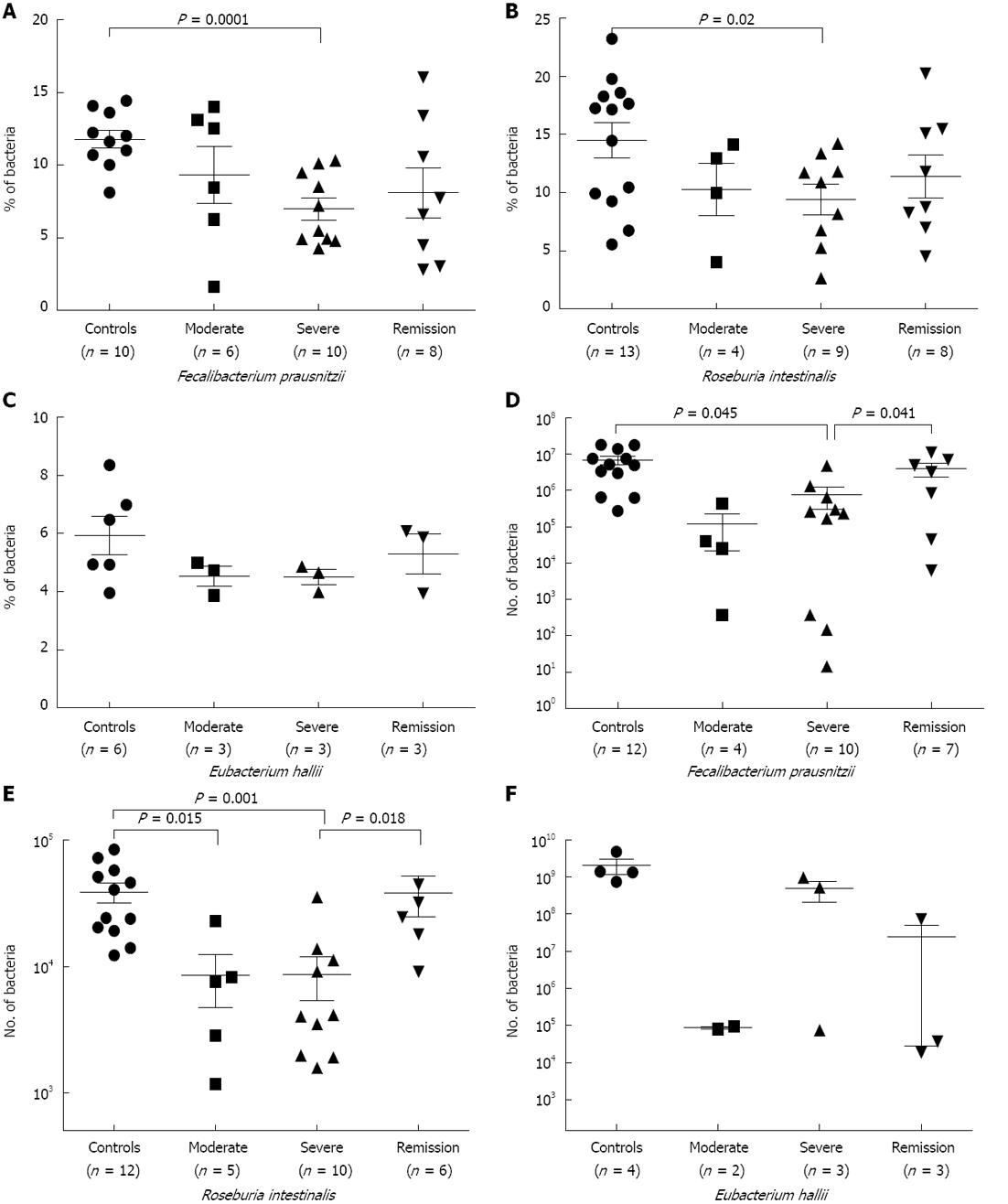
FISH-flow cytometry: Next, we evaluated the concentration of predominant members of both clostridial clusters (XIVa and IV). The population of F. prausnitzii, a member of the C. leptum group, was significantly low in UC samples of severe (P = 0.0001) category of disease in comparison to control samples (Figure 4A). Samples from the remission stage did not show significant restoration of F. prausnitzii in our FISH-flow cytometry experiment, as was observed for the C. leptum cluster during remission (Figures 1F and 2B).
R. intestinalis, a member of the C. coccoides group, was also significantly low (P = 0.02) in patient samples of severe category disease (Figure 4B); however, this did not show significant restoration in the samples of remission stage. The population of E. halii, another member of the C. coccoides group, decreased in disease conditions, but not significantly (Figure 4C). We failed to detect E. hallii in the majority of samples either by flow cytometry or qPCR, indicating low abundance of these bacteria in our study population. An increase in the population of R. intestinalis was recorded during remission stage in comparison to severe stage of the disease (Figure 4B and C). This was in agreement with our FISH-flow cytometry data of C. coccoides group. Abundance of R. intestinalis, E. hallii and F. prausnitzii was calculated out of total microbiota, as shown in Table 4. We detected a higher representation of R. intestinalis compared to E. hallii and F. prausnitzii. Therefore, we can infer that at the species level, abundance of the members of the C. coccoides cluster was higher in comparison to the members of the C. leptum cluster.
| Cluster or species | Out of total microbiota | Species | Out of respective cluster |
| Control fecal samples | |||
| C. coccoides cluster | 25.69% ± 1.62% | R. intestinalis | 14.48% ± 1.52% |
| E. hallii | 5.93% ± 0.54% | ||
| C. leptum cluster | 13.74% ± 1.05% | F. prausnitzii | 11.66% ± 1.55% |
| R .intestinalis1 | 3.71% ± 0.39% | ||
| E. hallii1 | 1.52% ± 0.13% | ||
| F. prausnitzii1 | 1.60% ± 0.21% | ||
| UC fecal samples (moderate) | |||
| C. coccoides cluster | 12.70% ± 4.60% | R. intestinalis | 10.00% ± 2.25% |
| E. hallii | 4.03% ± 0.3% | ||
| C. leptum cluster | 6.40% ± 3.40% | F. prausnitzii | 7.5% ± 2.3% |
| R .intestinalis1 | 1.27% ± 0.10% | ||
| E. hallii1 | 0.51% ± 0.01% | ||
| F. prausnitzii1 | 0.48% ± 0.14% | ||
| UC fecal samples (severe) | |||
| C. coccoides cluster | 9.80% ± 2.40% | R. intestinalis | 9% ± 1.83% |
| E. hallii | 4.2% ± 0.2% | ||
| C. leptum cluster | 6.20% ± 1.80% | F. prausnitzii | 6.01% ± 1.6% |
| R .intestinalis1 | 0.82% ± 0.04% | ||
| E. hallii1 | 0.41% ± 0.01% | ||
| F. prausnitzii1 | 0.37% ± 0.02% | ||
| UC fecal samples (remission) | |||
| C. coccoides cluster | 12.99% ± 2.65% | R. intestinalis | 11.40% ± 1.83% |
| E. hallii | 5.07% ± 0.6% | ||
| C. leptum cluster | 7.89% ± 1.50% | F. prausnitzii | 7.40% ± 2.32% |
| R .intestinalis1 | 1.48% ± 0.04% | ||
| E. hallii1 | 0.65% ± 0.01% | ||
| F. prausnitzii1 | 0.58% ± 0.03% | ||
As expected, the population of F. prausnitzii was significantly reduced in the samples of severe category of UC disease (P = 0.045) (Figure 4D) as compared to control. In addition, abundance of the same species was restored at remission stage when compared to severe stage (P = 0.041) (Figure 4D). Similarly, the R. intestinalis population showed significant reduction in samples of moderate (P = 0.015) and severe (P = 0.001) stage of the disease in comparison to control (Figure 4E). The recovery in the population of R. intestinalis was also seen in the remission category of samples when compared to severe category (P = 0.018) (Figure 4E). However, qPCR analysis in the case of E. hallii did not show any significant reduction; in addition, E. hallii was undetected in the majority of samples observed by FISH-flow cytometry (Figure 4F).
Our study revealed significant reduction in the members of both C. coccoides and C. leptum groups in fecal samples obtained from the severe disease category of UC patients in comparison to controls. FISH-flow cytometry and qPCR analysis of fecal samples belonging to the above groups supported the observations made by Takaishi et al[4] and Sokol et al[5].
We quantified the abundance of predominant butyrate-producing species of clostridial clusters to see their association with acetate and butyrate. Since in most cases butyryl-CoA:acetate CoA-transferase rather than butyrate kinase appears to perform the final step in butyrate synthesis, we targeted bacteria possessing butyryl CoA:acetate CoA-transferase for butyrate synthesis, e.g., F. prausnitzii and Roseburia spp/E. rectale which apparently lack butyrate kinase activity[15,28]. It is known that Roseburia spp. and F. prausnitzii strains contribute in butyrate production and many strains are associated with the net consumption of both acetate and carbohydrate[15]. Our study revealed that both the above species exhibited low abundance in the samples of severe category of UC and tended to restore their population during remission.
F. prausnitzii has been shown to exhibit anti-inflammatory effects on cellular and trinitrobenzenesulphonic acid-induced colitis models, partly due to secreted metabolites that are able to block NF-κB activation and interleukin 8 production[29]. In our observations (Table 4), F. prausnitzii accounted for 11.66% ± 1.55% of the C. leptum group, and R. intestinalis and E. hallii accounted for 14.48% ± 1.52% and 5.93% ± 0.54% of the C. coccoides group, respectively, in control individuals. In total these three species accounted for 6.83% of the total fecal flora. Previous studies have already reported low counts of F. prausnitzii in UC patients[30].
We observed a significant decrease in concentrations of butyrate, iso-butyrate and acetate in UC fecal samples compared to controls through SCFA quantification by GC analysis, supporting earlier observations[4,31].
An intriguing link between the level of SCFA and an intracellular energy sensor for the maintenance of intestinal barrier function has been suggested[10]. Our study further substantiates a link between the fluctuations of butyrate production and the changes in the numbers of butyrate producers during UC. Lack of butyrate availability may lead to compromised intestinal barrier function resulting in increased exposure of luminal content to the host immune system, thus exaggerating the immune response[10].
The reduced population of dominant butyrate producers and decreases in concentrations of butyrate and acetate in diseased samples indicate impaired butyrate supply in the colon, which may lead to energy deficiency for colonocytes. The resurgence of butyrate-producing bacteria at the remission stage and simultaneous increases in butyrate concentration, as observed here, show an association with UC and support the energy deficiency hypothesis of IBD[32]. Earlier studies have shown a requirement of acetate by F. prausnitzii and R. intestinalis for survival and other activities like production of butyrate[15,33]. Thus, major butyrogenic species depend on other bacteria, including net producers of acetate (e.g., amylolytic bifidobacteria) and other bacterial species capable of degrading a variety of complex carbohydrates[15]. These bacteria play an important role in net butyrate production and may be associated with dysbiosis during UC.
Acetate and butyrate are major SCFAs and decreases in their concentration may affect the overall SCFA concentration. SCFAs in the colon maintain mild acidic conditions which boost the formation of butyrate by favoring growth of butyrate-producing bacteria and allowing them to compete against gram-negative bacteria such as Bacteroides spp to maintain the homeostasis[9]. Thus, a reduced butyrate level may lead to an increased Gram-negative bacteria population due to reduced competition. This was validated by our observation of increased representation of Bacteroides in UC samples (data not shown).
Our data showing reduced abundance of three predominant butyrate-producing species indicate that during UC, butyrate deficiency may not be solely due to reduced uptake of butyrate by the inflamed mucosa as reported by Canani et al[9], but also due to reduced abundance of predominant butyrate producers and conversely lower production of butyrate.
The limitation of our findings is that most of the microbial data are not presented in absolute values but in percentages; therefore, it is difficult to critically assess the real changes in the microbial composition. The study is performed in a very restricted number of subjects due to the exclusion criteria followed during sample collection.
In conclusion, our study shows a decreased abundance of predominant butyrate producers like R. intestinalis and F. prausnitzii belonging to clostridial clusters in the UC disease condition. This decrease was found to correlate with reduced SCFA concentration in UC patients. We thus provide evidence that reduced butyrate levels during the diseased state are due to less abundance of these species in UC. It is evident from our data that decreased levels of these species resulting in reduced butyrate levels may be associated with the etiology of UC. This was further demonstrated in our analysis of the samples from remission patients, where the targeted organisms tended to revert back to normal level. It would be of interest in the future to extend this approach to studying the diversity of each member of the clostridial cluster, including changes in non-butyrate producers.
Inflammatory bowel disease (IBD), comprising of Crohn’s disease and ulcerative colitis (UC), is a class of chronic inflammatory disorders of the intestine and involves impaired barrier function. A rising trend in the incidence and prevalence of inflammatory bowel IBD in the Asian population has been recognized for the past two decades. IBD is considered as an exaggerated immune response, exerted either to all commensal bacteria or to a subset or to a single strain of bacteria.
Differences in fecal microbiota of healthy subjects and IBD patients have been identified. Commensal gut microbiota have been reported to exert various multiple beneficial effects on host gut epithelium. short chain fatty acids (SCFAs), like butyrate, produced by microbial fermentation of undigested carbohydrates have been depicted to regulate transepithelial transport, colonocyte proliferation and differentiation, mucosal inflammation, intestinal motility and barrier function. The homeostasis between commensal micobiota and host defensive responses at the mucosal level has a pivotal role in the initiation and pathophysiology of chronic IBD. Therefore, monitoring the fluctuations in butyrogenic bacteria and SCFA level may help in understanding the relation of dysbiosis with UC.
This work was carried out to study the fluctuations in butyrate-producing bacteria and simultaneous measurement of butyrate concentration in fecal samples of UC vs control individuals belonging to the northern part of India. The data were also correlated with patients differing in disease activity. Decreases in the level of butyrate producers in UC patients coincided with the changes in butyrate concentration during active stage disease. However, the levels reverted back to normal during remission stage showing the role of these bacteria in the disease etiology. Such comparisons have not been made so far to show the above relationship with disease activity.
It is speculated that by measuring the level of these species and concentration of SCFA in stool samples (a non-invasive method), authors can indicate the status of UC patients.
Dysbiosis (also called dysbacteriosis) refers to a condition with microbial imbalances on or within the body. Dysbiosis is most prominent in the digestive tract or on the skin, but can also occur on any exposed surface or mucous membrane such as the vagina, lungs, mouth, nose, sinuses, ears, nails or eyes. It has been associated with different illnesses, such as IBD, as imbalances in the intestinal microbiome may be associated with bowel inflammation and chronic fatigue syndrome.
This is a methodologically well performed, interesting report, addressing the potential role of gut commensal microbiota in the pathophysiology of UC.
P- Reviewer Martinez V S- Editor Jiang L L- Editor Logan S E- Editor Zhang DN
| 1. | Thia KT, Loftus EV, Sandborn WJ, Yang SK. An update on the epidemiology of inflammatory bowel disease in Asia. Am J Gastroenterol. 2008;103:3167-3182. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 371] [Cited by in RCA: 408] [Article Influence: 24.0] [Reference Citation Analysis (1)] |
| 2. | Xavier RJ, Podolsky DK. Unravelling the pathogenesis of inflammatory bowel disease. Nature. 2007;448:427-434. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2894] [Cited by in RCA: 3351] [Article Influence: 186.2] [Reference Citation Analysis (11)] |
| 3. | Stenson WF, Snapper SB. Challenges in IBD research: Assessing progress and rethinking the research agenda. Inflamm Bowel Dis. 2008;14:687-708. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 4. | Takaishi H, Matsuki T, Nakazawa A, Takada T, Kado S, Asahara T, Kamada N, Sakuraba A, Yajima T, Higuchi H. Imbalance in intestinal microflora constitution could be involved in the pathogenesis of inflammatory bowel disease. Int J Med Microbiol. 2008;298:463-472. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 223] [Cited by in RCA: 248] [Article Influence: 13.8] [Reference Citation Analysis (0)] |
| 5. | Sokol H, Seksik P, Rigottier-Gois L, Lay C, Lepage P, Podglajen I, Marteau P, Doré J. Specificities of the fecal microbiota in inflammatory bowel disease. Inflamm Bowel Dis. 2006;12:106-111. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 334] [Cited by in RCA: 323] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 6. | Manichanh C, Rigottier-Gois L, Bonnaud E, Gloux K, Pelletier E, Frangeul L, Nalin R, Jarrin C, Chardon P, Marteau P. Reduced diversity of faecal microbiota in Crohn’s disease revealed by a metagenomic approach. Gut. 2006;55:205-211. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1633] [Cited by in RCA: 1676] [Article Influence: 88.2] [Reference Citation Analysis (0)] |
| 7. | Vernia P, Marcheggiano A, Caprilli R, Frieri G, Corrao G, Valpiani D, Di Paolo MC, Paoluzi P, Torsoli A. Short-chain fatty acid topical treatment in distal ulcerative colitis. Aliment Pharmacol Ther. 1995;9:309-313. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 150] [Cited by in RCA: 159] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 8. | Gibson GR. Physiology and ecology of the sulphate-reducing bacteria. J Appl Bacteriol. 1990;69:769-797. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 149] [Cited by in RCA: 115] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 9. | Canani RB, Costanzo MD, Leone L, Pedata M, Meli R, Calignano A. Potential beneficial effects of butyrate in intestinal and extraintestinal diseases. World J Gastroenterol. 2011;17:1519-1528. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 802] [Cited by in RCA: 909] [Article Influence: 64.9] [Reference Citation Analysis (11)] |
| 10. | Peng L, Li ZR, Green RS, Holzman IR, Lin J. Butyrate enhances the intestinal barrier by facilitating tight junction assembly via activation of AMP-activated protein kinase in Caco-2 cell monolayers. J Nutr. 2009;139:1619-1625. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1443] [Cited by in RCA: 1339] [Article Influence: 83.7] [Reference Citation Analysis (0)] |
| 11. | Scheppach W. Effects of short chain fatty acids on gut morphology and function. Gut. 1994;35:S35-S38. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 517] [Cited by in RCA: 506] [Article Influence: 16.3] [Reference Citation Analysis (0)] |
| 12. | Lay C, Rigottier-Gois L, Holmstrøm K, Rajilic M, Vaughan EE, de Vos WM, Collins MD, Thiel R, Namsolleck P, Blaut M. Colonic microbiota signatures across five northern European countries. Appl Environ Microbiol. 2005;71:4153-4155. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 196] [Cited by in RCA: 196] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 13. | Segain JP, Raingeard de la Blétière D, Bourreille A, Leray V, Gervois N, Rosales C, Ferrier L, Bonnet C, Blottière HM, Galmiche JP. Butyrate inhibits inflammatory responses through NFkappaB inhibition: implications for Crohn’s disease. Gut. 2000;47:397-403. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 886] [Cited by in RCA: 973] [Article Influence: 38.9] [Reference Citation Analysis (0)] |
| 14. | Scheppach W, Bartram HP, Richter F. Role of short-chain fatty acids in the prevention of colorectal cancer. Eur J Cancer. 1995;31A:1077-1080. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 236] [Cited by in RCA: 228] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 15. | Duncan SH, Holtrop G, Lobley GE, Calder AG, Stewart CS, Flint HJ. Contribution of acetate to butyrate formation by human faecal bacteria. Br J Nutr. 2004;91:915-923. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 276] [Cited by in RCA: 334] [Article Influence: 15.9] [Reference Citation Analysis (0)] |
| 16. | Hold GL, Schwiertz A, Aminov RI, Blaut M, Flint HJ. Oligonucleotide probes that detect quantitatively significant groups of butyrate-producing bacteria in human feces. Appl Environ Microbiol. 2003;69:4320-4324. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 215] [Cited by in RCA: 252] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 17. | Louis P, Young P, Holtrop G, Flint HJ. Diversity of human colonic butyrate-producing bacteria revealed by analysis of the butyryl-CoA: acetate CoA-transferase gene. Environ Microbiol. 2010;12:304-314. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 449] [Cited by in RCA: 564] [Article Influence: 35.3] [Reference Citation Analysis (0)] |
| 18. | Walmsley RS, Ayres RC, Pounder RE, Allan RN. A simple clinical colitis activity index. Gut. 1998;43:29-32. [PubMed] |
| 19. | Tursi A, Brandimarte G, Papa A, Giglio A, Elisei W, Giorgetti GM, Forti G, Morini S, Hassan C, Pistoia MA. Treatment of relapsing mild-to-moderate ulcerative colitis with the probiotic VSL#3 as adjunctive to a standard pharmaceutical treatment: a double-blind, randomized, placebo-controlled study. Am J Gastroenterol. 2010;105:2218-2227. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 391] [Cited by in RCA: 353] [Article Influence: 23.5] [Reference Citation Analysis (0)] |
| 20. | Amann RI, Krumholz L, Stahl DA. Fluorescent-oligonucleotide probing of whole cells for determinative, phylogenetic, and environmental studies in microbiology. J Bacteriol. 1990;172:762-770. [PubMed] |
| 21. | Wallner G, Amann R, Beisker W. Optimizing fluorescent in situ hybridization with rRNA-targeted oligonucleotide probes for flow cytometric identification of microorganisms. Cytometry. 1993;14:136-143. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 913] [Cited by in RCA: 818] [Article Influence: 25.6] [Reference Citation Analysis (0)] |
| 22. | Stoecker K, Dorninger C, Daims H, Wagner M. Double labeling of oligonucleotide probes for fluorescence in situ hybridization (DOPE-FISH) improves signal intensity and increases rRNA accessibility. Appl Environ Microbiol. 2010;76:922-926. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 129] [Cited by in RCA: 117] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 23. | Saunier K, Rougé C, Lay C, Rigottier-Gois L, Doré J. Enumeration of bacteria from the Clostridium leptum subgroup in human faecal microbiota using Clep1156 16S rRNA probe in combination with helper and competitor oligonucleotides. Syst Appl Microbiol. 2005;28:454-464. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 10] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 24. | Zoetendal EG, Ben-Amor K, Harmsen HJ, Schut F, Akkermans AD, de Vos WM. Quantification of uncultured Ruminococcus obeum-like bacteria in human fecal samples by fluorescent in situ hybridization and flow cytometry using 16S rRNA-targeted probes. Appl Environ Microbiol. 2002;68:4225-4232. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 91] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 25. | Langendijk PS, Schut F, Jansen GJ, Raangs GC, Kamphuis GR, Wilkinson MH, Welling GW. Quantitative fluorescence in situ hybridization of Bifidobacterium spp. with genus-specific 16S rRNA-targeted probes and its application in fecal samples. Appl Environ Microbiol. 1995;61:3069-3075. [PubMed] |
| 26. | Whelan JA, Russell NB, Whelan MA. A method for the absolute quantification of cDNA using real-time PCR. J Immunol Methods. 2003;278:261-269. [PubMed] |
| 27. | Martin FP, Dumas ME, Wang Y, Legido-Quigley C, Yap IK, Tang H, Zirah S, Murphy GM, Cloarec O, Lindon JC. A top-down systems biology view of microbiome-mammalian metabolic interactions in a mouse model. Mol Syst Biol. 2007;3:112. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 380] [Cited by in RCA: 343] [Article Influence: 19.1] [Reference Citation Analysis (0)] |
| 28. | Louis P, Flint HJ. Diversity, metabolism and microbial ecology of butyrate-producing bacteria from the human large intestine. FEMS Microbiol Lett. 2009;294:1-8. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1125] [Cited by in RCA: 1466] [Article Influence: 91.6] [Reference Citation Analysis (0)] |
| 29. | Sokol H, Pigneur B, Watterlot L, Lakhdari O, Bermúdez-Humarán LG, Gratadoux JJ, Blugeon S, Bridonneau C, Furet JP, Corthier G. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc Natl Acad Sci USA. 2008;105:16731-16736. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2747] [Cited by in RCA: 3199] [Article Influence: 188.2] [Reference Citation Analysis (0)] |
| 30. | Sokol H, Seksik P, Furet JP, Firmesse O, Nion-Larmurier I, Beaugerie L, Cosnes J, Corthier G, Marteau P, Doré J. Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm Bowel Dis. 2009;15:1183-1189. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 857] [Cited by in RCA: 912] [Article Influence: 57.0] [Reference Citation Analysis (0)] |
| 31. | Huda-Faujan N, Abdulamir AS, Fatimah AB, Anas OM, Shuhaimi M, Yazid AM, Loong YY. The impact of the level of the intestinal short chain Fatty acids in inflammatory bowel disease patients versus healthy subjects. Open Biochem J. 2010;4:53-58. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 210] [Cited by in RCA: 261] [Article Influence: 17.4] [Reference Citation Analysis (0)] |
| 32. | Roediger WE. The colonic epithelium in ulcerative colitis: an energy-deficiency disease? Lancet. 1980;2:712-715. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 469] [Cited by in RCA: 472] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 33. | Duncan SH, Barcenilla A, Stewart CS, Pryde SE, Flint HJ. Acetate utilization and butyryl coenzyme A (CoA):acetate-CoA transferase in butyrate-producing bacteria from the human large intestine. Appl Environ Microbiol. 2002;68:5186-5190. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 500] [Cited by in RCA: 510] [Article Influence: 22.2] [Reference Citation Analysis (0)] |
| 34. | Franks AH, Harmsen HJ, Raangs GC, Jansen GJ, Schut F, Welling GW. Variations of bacterial populations in human feces measured by fluorescent in situ hybridization with group-specific 16S rRNA-targeted oligonucleotide probes. Appl Environ Microbiol. 1998;64:3336-3345. [PubMed] |
| 35. | Sghir A, Gramet G, Suau A, Rochet V, Pochart P, Dore J. Quantification of bacterial groups within human fecal flora by oligonucleotide probe hybridization. Appl Environ Microbiol. 2000;66:2263-2266. [PubMed] |
| 36. | Verma AK, Verma R, Ahuja V, Paul J. Real-time analysis of gut flora in Entamoeba histolytica infected patients of Northern India. BMC Microbiol. 2012;12:183. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 70] [Cited by in RCA: 79] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 37. | Wang RF, Cao WW, Cerniglia CE. PCR detection and quantitation of predominant anaerobic bacteria in human and animal fecal samples. Appl Environ Microbiol. 1996;62:1242-1247. [PubMed] |
| 38. | Ramirez-Farias C, Slezak K, Fuller Z, Duncan A, Holtrop G, Louis P. Effect of inulin on the human gut microbiota: stimulation of Bifidobacterium adolescentis and Faecalibacterium prausnitzii. Br J Nutr. 2009;101:541-550. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 530] [Cited by in RCA: 595] [Article Influence: 35.0] [Reference Citation Analysis (0)] |