Published online Aug 27, 2025. doi: 10.4254/wjh.v17.i8.107456
Revised: May 9, 2025
Accepted: July 16, 2025
Published online: August 27, 2025
Processing time: 149 Days and 20.3 Hours
Whether rtS106C+H126Y+D134E/rtS106C+H126Y+D134E+L269I (rtCYE/rtCYEI) mutations in the hepatitis B virus (HBV) reverse-transcriptase (RT) region are associated with tenofovir disoproxil fumarate (TDF) resistance is controversial.
To evaluate the presence of the rtCYE/rtCYEI mutations in a large cohort of Chinese patients with chronic HBV infection.
A total of 28236 patients who underwent drug resistance testing at the Fifth Medical Center of Chinese PLA General Hospital from 2007 to 2019 were enrolled. All patients received nucleoside/nucleotide analogues (NAs) therapy, and serum samples were collected for sequence analysis of the HBV RT domain with mutation analysis.
The detection rates of a single mutation of rtS106C, rtH126Y, rtD134E, and rtL269I were 8.21%, 3.20%, 2.55% and 61.49% in 23718 genotype C patients, and 1.31%, 1.76%, 0.21%, and 92.33% in 4266 genotype B patients, res
HBV rtCYE/rtCYEI mutations have a limited effect on TDF susceptibility and are not sufficient to cause TDF resistance.
Core Tip: Whether rtS106C+H126Y+D134E/rtS106C+H126Y+D134E+L269I (rtCYE/rtCYEI) mutations in the reverse-transcriptase region of hepatitis B virus (HBV) are associated with tenofovir resistance is controversial. The present study investigated many nucleoside/nucleotide analogues (NAs)-treated Chinese patients, focusing on the relationship between rtCYE/rtCYEI mutations and tenofovir resistance. We confirmed that rtCYE/rtCYEI mutations were rare (0.042%) in NAs-treated chronic HBV-infected patients and could appear in tenofovir treatment-naïve patients and remain sensitive to tenofovir. The effect of rtCYE/rtCYEI mutations on tenofovir susceptibility was limited. This study extends the current knowledge related to HBV resistance, which helps to explain the controversy and has clinical implications for resistance management.
- Citation: Si LL, Fan ZP, Liu WH, Chen RJ, Chen XY, Ji D, Li L, Chen C, Liao H, Wang J, Xu DP, Zhao J, Liu Y. Hepatitis B virus rtCYE/rtCYEI mutations may contribute limited tenofovir resistance: Analysis of a large sample of Chinese patients. World J Hepatol 2025; 17(8): 107456
- URL: https://www.wjgnet.com/1948-5182/full/v17/i8/107456.htm
- DOI: https://dx.doi.org/10.4254/wjh.v17.i8.107456
It is estimated that over 296 million people worldwide, including 86 million people in China are chronically infected with the hepatitis B virus (HBV) and these people are at risk for HBV-related liver diseases, such as liver cirrhosis (LC) and hepatocellular carcinoma (HCC)[1-3]. Currently, nucleoside/nucleotide (nt) analogues (NAs) are major anti-HBV drugs. Six NAs have been approved to treat chronic hepatitis B (CHB) in China, including lamivudine (LAM), adefovir dipivoxil (ADV), telbivudine (LdT), entecavir (ETV), tenofovir disoproxil fumarate (TDF, licensed in China in June 2014), and tenofovir alafenamide (TAF). ETV, TDF and TAF are currently recommended as first-line anti-HBV agents, whereas LAM, LdT and ADV were once popular anti-HBV agents used alone or in combination[4-6].
NAs have no direct inhibitory effect on HBV covalently closed circular DNA; therefore, long-term NAs therapy is required, which increases the risk of HBV resistance mutations[7,8]. The signature primary resistance mutations include rtM204I/V (LAMr) for LAM (rtM204I, also LdT resistance); rtA181V/rtN236T (ADVr) for ADV; and LAMr plus rtT184 substitutions (i.e., A/C/F/G/I/L/M/S), rtS202 (C/G/I), or rtM250 (I/L/V) (ETVr) for ETV. In addition, rtL80I and rtL180M are considered resistance-associated mutations as they can compensate for the replication capacity of rtM204I/V mutants[8,9]. Moreover, some unusual mutations have been found to confer primary resistance, such as rtN236V, rtA181S, rtA186T, and rtL180M+A181C+M204V[4,10,11].
It was recently reported that the rtS106C+H126Y+D134E/rtS106C+H126Y+D134E+L269I (rtCYE/rtCYEI) mutations conferred a 3.7-fold to 15.3-fold reduction in TDF sensitivity and were associated with TDF resistance[12,13], and the rtI (rtL269I) provided replication compensation for rtCYE backbone mutants[14]. However, the results were disputed by other investigators[15,16]. So far, data on the clinical significance and virological features of rtCYE/rtCYEI mutations remain very limited. To date, there is no solid evidence of a clear relationship between LAM/LdT/ADV/ETV exposure/resistance and the emergence of rtCYE/rtCYEI mutations, or for the introduction of TDF resistance by LAM-/LdT-/ADV-/ETV-resistance mutations in the clinic. This study aimed to further the knowledge of HBV rtCYE/rtCYEI mu
A total of 28236 patients whose blood samples were collected and underwent drug resistance testing (direct sequencing) at the Fifth Medical Center of Chinese PLA General Hospital (formerly named Beijing 302 Hospital) from July 2007 to June 2019 were enrolled in the study. In this cohort, all patients underwent testing due to clinical suspicion of drug resistance, not as part of general population screening. This cohort of patients has also been enrolled to assess HBV multidrug-resistance mutations[4]. These patients received LAM, LdT, ADV, ETV and/or TDF monotherapy or combination therapy and had no co-infection with other hepatitis viruses or human immunodeficiency virus infection. They were diagnosed using the Guidelines of Prevention and Treatment for Hepatitis B Disease, and the disease categories of chronic HBV infection were CHB, HBV-related LC, and HCC[2,17]. These patients were from the Beijing 302 Hospital database and they all provided their informed consent to use the samples for research before enrollment. The study was approved by the Ethics Committee of Beijing 302 Hospital (Reference Number: FRM-EC-018-02-V2.0).
Biochemical and serological markers, along with HBV DNA levels, were routinely measured in the Central Clinical Laboratory of the Fifth Medical Center of Chinese PLA General Hospital. HBV DNA levels were determined using a real-time quantitative polymerase chain reaction (PCR) kit (Fosun Pharmaceutical Co., Ltd., Shanghai, China) with a lower limit of detection (LLOD) of 100 IU/mL for samples analyzed before April 2012 and 40 IU/mL for all other samples. Sequence analysis was performed as previously described[4]. Briefly, a 1225-bp-long viral gene fragment (nt 54-1278) covering the full-length of the reverse-transcriptase (RT) region (nt 130-1161) and overlapping S region (nt 155-835) was amplified using a one-tube nested PCR method with an LLOD of 10 IU/mL. Resistance mutations were analyzed using direct sequencing by the Sanger method. Co-existing nts at one position were determined if the sequencing peak of one nt reached 25% of the sequencing summit of another dominant nt at that position, given that direct sequencing can detect a particular mutant only if it is present at ≥ 20%[18]. Clone construction and sequencing (≥ 20 clones/sample) were per
The pTriEx-mod-1.1 vector (construct of the 1.1-mer genotype C HBV genome) was kindly provided by Professor Zoulim, University of Lyon. Information and applications of this vector have been described previously[19,20]. Patient-derived HBV RT sequences were included in the construction. Specifically, the sequence rtCYE+M204I was from patient 2, the sequences rtWT, rtCYE+L80I+L180M+M204I and rtCYEI+L80I+L180M+M204I were from patient 3, the sequence rtCYEI+A181V was from patient 4, the sequence rtCYE was from patient 7, the sequences rtCYEI and rtCYEI+N236T were from patient 9, and the sequence rtCYEI+M204I was from patient 11. In addition, the rtWT (laboratory) sequence was created by eliminating rtH126Y from patient 7. A QuikChange Lightning site-directed mutagenesis kit (Stratagene, La Jolla, CA, United States) was used with the primer (sense) 5’-ATCAACCACCAGCACGGGACCATGC-3’. The clones obtained were all validated by sequencing analysis.
The phenotypic analysis was performed as previously described[4]. Briefly, the constructs were transiently transfected into HepG2 cells before the addition of serially diluted TDF. HepG2 cells were first cultured in 24-well plates at 1 × 105 cells per well before transfection. Then, each well was transfected with 0.25 μg mutant or wild-type (WT) HBV genomic recombinant vectors using FuGene HD transfection reagent (Roche, Mannheim, Germany) and the transfection efficiency was normalized by β-galactosidase with a reporter plasmid (Promega, Madison, WI, United States). Four hours after transfection, TDF susceptibility analysis was performed using serially diluted TDF (0, 0.78, 3.125, 12.5, 50, and 200 mmol/L). After four days of culture, the cells were harvested, lysed, and viral core particles were immunoprecipitated using anti-HBc/protein A+G beads. HBV replicative intermediates in core particles were released and quantitated by real-time PCR (Chinese patent ZL 2013103921225). The real-time PCR method with a LOD was experimentally determined to be 40 IU/mL using a 10-fold dilution of DNA standards [from 102 to 108 IU/mL, with a linear correlation coefficient (R²) of 0.99973 over this range]. The relative replication capacity of mutants vs WT strains was assessed in NAs-free conditions. Drug susceptibility was determined by comparing the half maximal effective concentration (EC50) of the drug in mutant vs WT samples. The experiments were performed at least three times independently.
This method has been described in our previous studies[11]. Briefly, the modeling structure of HBV RT was constructed by SWISS-MODEL (https://www.swissmodel.expasy.org) based on the crystal structure of the human immunodeficiency virus RT (Protein Data Bank accession number; 1RTD). This HBV RT modeling structure was used as a WT to construct the structures of the HBV rtCYE and rtCYEI mutants. Autodock software (version 4.2.6, molecular graphics laboratory) was used to simulate the docking process and evaluate the binding energy of HBV RT and TDF. Autodock is a component of the MGLTools that is available for academic use. More detailed information on Autodock can be found at the following website: http://mgltools.scripps.edu/.
Data are presented as mean ± SD or median (inter-quartile range). Differences between variables and univariate analysis were examined by the Student’s t-test (two-tailed) or χ2 test. Results are presented as odds ratios (OR) and corresponding P values in univariate analysis. Multivariate analyses with logistic regression were used to determine independent factors. Statistical analysis was carried out using the Statistical Program for Social Sciences (SPSS 23.0 for Windows; SPSS Inc., Chicago, IL, United States). A P value < 0.05 was considered statistically significant.
The results of HBV genotyping by direct sequencing showed that of all 28236 patients, 9 (0.03%), 4266 (15.11%), 23718 (84.00%), and 243 (0.86%) were genotype A, B, C, and D HBV-infected patients, respectively. No other HBV genotypes were detected. The detection rates of single mutations rtC, rtY, rtE, and rtI in patients with genotype C were 8.21%, 3.20%, 2.55%, and 61.49%, respectively, and in patients with genotype B were 1.31%, 1.76%, 0.21%, and 92.33%, respectively. The rtCYE/rtCYEI mutations (with or without co-existing resistance mutations) were detected in 0.051% (12/23,718) of HBV genotype C infected patients, and in 0% of the other genotypes (Table 1). None of these 12 patients were receiving TDF therapy at the time of serum collection.
| Mutation(s) | Genotype A (n = 9) | Genotype B (n = 4266) | Genotype C (n = 23718) | Genotype D (n = 243) |
| rtS106C | 0 (0.00) | 56 (1.31) | 1947 (8.21) | 1 (0.41) |
| rtH126Y | 7 (77.78) | 75 (1.76) | 759 (3.20) | 3 (1.23) |
| rtD134E | 0 (0.00) | 9 (0.21) | 605 (2.55) | 0 (0.00) |
| rtL269I | 2 (22.22) | 3939 (92.33) | 14584 (61.49) | 231 (95.06) |
| rtS106C+H126Y | 0 (0.00) | 0 (0.00) | 114 (0.48) | 0 (0.00) |
| rtS106C+H126Y+D134E | 0 (0.00) | 0 (0.00) | 6 (0.03) | 0 (0.00) |
| rtS106C+H126Y+D134E+L269I | 0 (0.00) | 0 (0.00) | 6 (0.03) | 0 (0.00) |
We constructed the HBV 1.1-mers where the original RT gene was replaced by the patient-derived gene and obtained multiple clonal constructs. By sequencing these constructs, we verified that all 12 patients harbored rtCYE/rtCYEI mu
| Patient | Date1 | GT | HBV DNA (log10 IU/mL) | ALT (U/L) | HBeAg | NAs exposure (month) | Direct sequencing | Clonal sequencing (positive clone number/total tested clone number) |
| P1 | 2009.02 | C | 8.16 | 55 | + | LAM (19) | rtS106C+H126Y+D134E | rtS106C+H126Y+D134E (18/20), rtS106C+H126Y+D134E+L80I+M204I (1/20), rtS106C+H126Y+D134E+L269I (1/20) |
| P2 | 2009.05 | C | 8.66 | 53 | + | LdT (18) | rtS106C+H126Y+D134E+M204I | rtS106C+H126Y+D134E+M204I (19/20), rtS106C+H126Y+D134E+L80I+M204I (1/20) |
| P3 | 2012.04 | C | 8.10 | 589 | + | LAM (33) | rtS106C+H126Y+D134E+L80I+L180M+M204I | rtS106C+H126Y+D134E+L80I+L180M+M204I (17/20), rtS106C+H126Y+D134E+L269I+L80I+L180M+M204I (3/20), rtS106C+H126Y+L80I+L180M+M204I (1/20) |
| P4 | 2013.06 | C | 6.77 | 375 | + | ADV (55) | rtS106C+H126Y+D134E+L269I+A181V | rtS106C+H126Y+D134E+L269I+A181V (17/20), rtH126Y+L269I+A181V (3/20) |
| P5 | 2013.08 | C | 1.60 | 20 | + | ADV (17)→ETV (15) | rtS106C+H126Y+D134E+L269I | rtS106C+H126Y+D134E+L269I (20/20) |
| P6 | 2013.08 | C | 5.52 | 20 | - | LdT (16) | rtS106C+H126Y+D134E+L269I+M204I | rtS106C+H126Y+D134E+L269I (18/20), rtS106C+H126Y+D134E+L269I+M204I (2/20) |
| P7 | 2008.06 | C | 5.21 | 37 | + | LAM (24)→ADV (18) | rtS106C+H126Y+D134E | rtH126Y (6/20), rtS106C+H126Y+D134E (14/20) |
| P8 | 2008.08 | C | 6.61 | 109 | + | LdT (16) | rtS106C+H126Y+D134E+L80I+M204I | rtS106C+H126Y+D134E+L80I+M204I (19/20), rtS106C+H126Y+D134E+L269I+L80I+M204I (1/20) |
| P9 | 2009.04 | C | 3.77 | 87 | - | ADV (32) | rtS106C+H126Y+D134E+L269I+N236T | rtS106C+H126Y+D134E+L269I (16/20), rtS106C+H126Y+D134E+L269I+N236T (4/20) |
| P10 | 2010.08 | C | 6.88 | 165 | - | LAM (28)→ADV (12) | rtS106C+H126Y+D134E | rtS106C+H126Y+D134E (20/20) |
| P11 | 2012.09 | C | 4.50 | 35 | - | LAM (38) | rtS106C+H126Y+D134E+L269I+M204I | rtS106C+H126Y+D134E+L269I+M204I (20/20) |
| P12 | 2014.06 | C | 4.18 | 75 | - | LAM (36)→ADV (25) | rtS106C+H126Y+D134E+L269I | rtS106C+H126Y+D134E+L269I (16/20), rtS106C+H126Y+D134E (4/20) |
Clinical characteristics of rtCYE/rtCYEI mutation-positive and -negative patients are compared in Table 3. Clinical data analysis showed that no association was observed between the occurrence of rtCYE/rtCYEI mutations and LAM, LdT, ADV, and/or ETV exposure, and none of the rtCYE/rtCYEI-positive patients received TDF treatment. Univariate analysis showed that there were no significant differences in the clinical parameters between the two patient groups. In the subsequent multivariate analysis, there were no significant differences in the clinical parameters between the two patient groups.
| rtCYE/rtCYEI (+) (n = 12) | rtCYE/rtCYEI (-) (n = 28224) | Univariate OR (95%CI) | P value | |
| Age (years) | 43 (33-49) | 42 (32-51) | 3.78 (-7.88-6.95) | 0.903 |
| Gender (male) | 10 (83.33) | 22419 (79.43) | 0.77 (0.17-3.53) | 1.00 |
| Genotype (A/B/C/D%) | 0/0/100/0 | 0.03/15.11/84.00/0.86 | 0.515 | |
| HBV DNA (log10 IU/mL) | 5.37 (3.40-6.69) | 4.03 (2.72-6.00) | 0.64 (-1.96-0.56) | 0.275 |
| ALT (U/L) | 65.0 (35.5-151.0) | 38.00 (23.0-72.0) | 57.48 (-153.95-71.37) | 0.473 |
| AST (U/L) | 33.0 (29.0-114.5) | 35.0 (25.0-66.0) | 73.71 (-179.46-109.51) | 0.635 |
| TBIL (µmol/L) | 13.85 (12.5-21.1) | 14.2 (10.2-22.0) | 20.32 (-29.46-50.21) | 0.610 |
| HBeAg (+) | 7 (58.33) | 17438 (61.78) | 0.87 (0.28-2.73) | 0.806 |
| LAM/LdT exposure1 | 9 (75.00) | 15419 (54.63) | 0.40 (0.11-1.48) | 0.260 |
| ADV exposure1 | 6 (50.00) | 16971 (60.13) | 1.51 (0.49-4.68) | 0.474 |
| ETV exposure1 | 1 (8.33) | 8698 (30.82) | 4.90 (0.63-37.96) | 0.169 |
| TDF exposure1 | 0 (0.00) | 390 (1.38) | 0.986 (0.98-0.99) | 1.000 |
| TAF exposure1 | 0 | 0 |
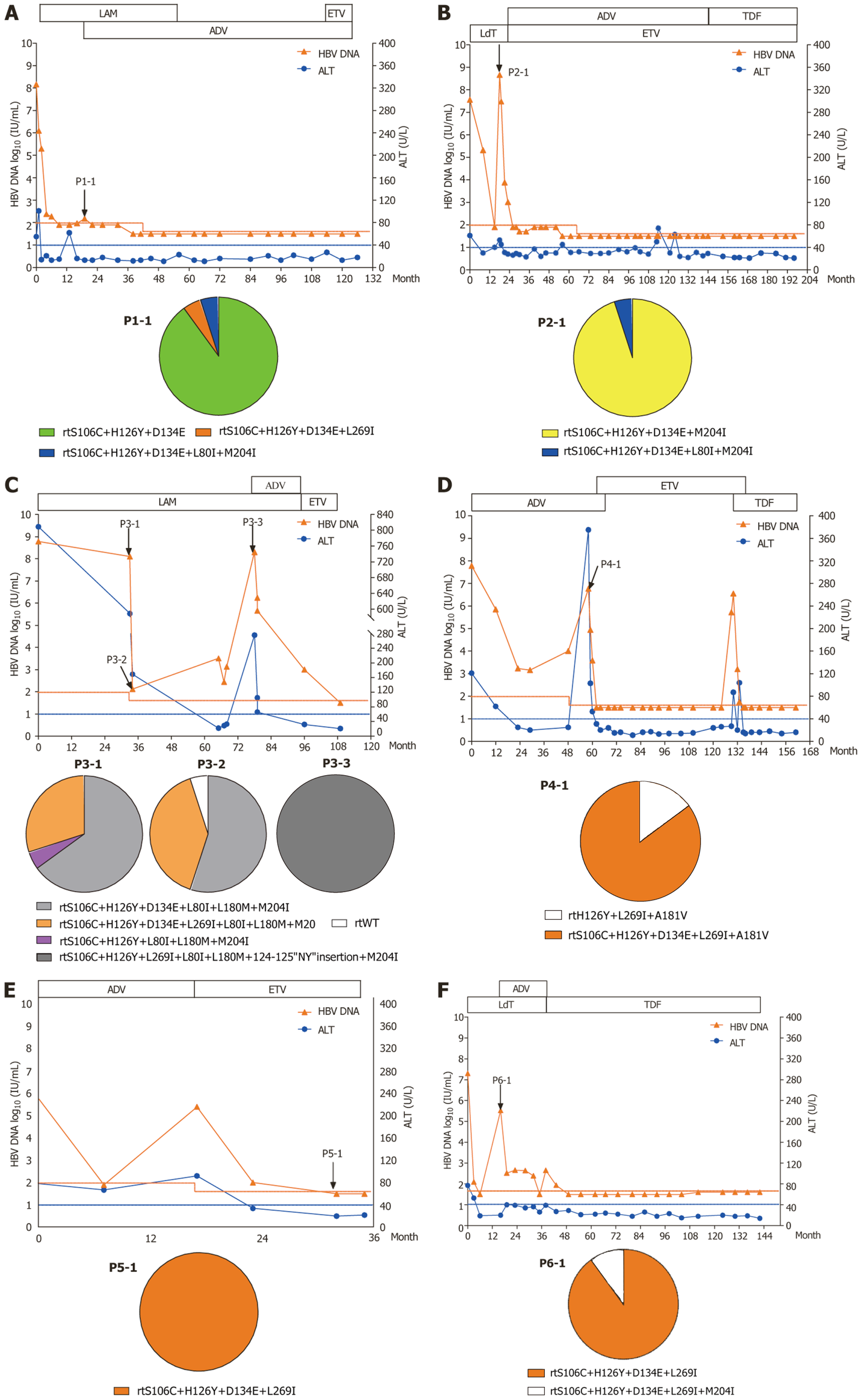
We retrospectively studied the 12 patients with rtCYE/rtCYEI mutations. Among them, six patients had a very long follow-up and provided available clinical data for analysis. The clinical course of these patients is shown in Figure 1.
Patient 1 received 19 months of LAM monotherapy from February 2009. In September 2010, ADV was added to the regimen until October 2013. Subsequently, ADV monotherapy was maintained for 60 months. In October 2018, antiviral therapy was adjusted to ADV combined with ETV for 12 months. Serum was sampled and tested at month 19 during LAM monotherapy. Using clone sequencing, mutants rtCYE (90%), rtCYEI (5%), and rtCYEI+L80I+M204I (5%) were detected (Figure 1A).
Patient 2 received 18 months of LdT therapy starting in November 2007 and had a clinical breakthrough before admission to Beijing 302 Hospital. Antiviral therapy was subsequently changed to ADV combined with ETV for 123 months, followed by ETV combined with TDF for 19 months. Sustained viral suppression was observed in the subsequent 36 months of TDF monotherapy. Serum (P2-1) was sampled in May 2009, and mutants rtCYE+M204I (95%) and rtCYE+L80I+M204I (5%) were detected simultaneously (Figure 1B).
Patient 3 received 78 months of LAM, 18 months of LAM+ADV, and 12 months of ETV therapy. Serum samples (P3-1, P3-2, and P3-3) were collected at months 33, 34, and 78 of LAM therapy. Mutants rtCYE+L80I+L180M+M204I (85%) and rtCYEI+L80I+L180M+M204I (15%) were dominantly detected at months 33 and 34. At month 78, an unreported mutant containing the rtCYI+L80I+L180M+M204I mutation plus rt124-125NY insertion was exclusively detected in tested viral clones (Figure 1C).
Patient 4 successively received 58 months of ADV, 4 months of ADV+ETV, 68 months of ETV, 6 months of ETV+TDF, and 25 months of TDF therapy. At the end of ADV therapy, a clinical breakthrough occurred. Serum at this time-point (P4-1) was sampled and mutant rtCYEI+A181V (85%) was dominantly detected (Figure 1D).
Patient 5 successively received 17 months of ADV and 18 months of ETV therapy. Serum at month 15 of ETV therapy (P5-1) was sampled and the rtCYEI mutant was exclusively detected in tested viral clones (Figure 1E).
Patient 6 successively received 16 months of LdT, 22 months of LdT+ADV, and 96 months of TDF therapy. Serum at the end of LdT monotherapy (P6-1) was sampled. Mutants rtCYEI (90%) and rtCYEI+M204I (10%) were detected in tested viral clones (Figure 1F).
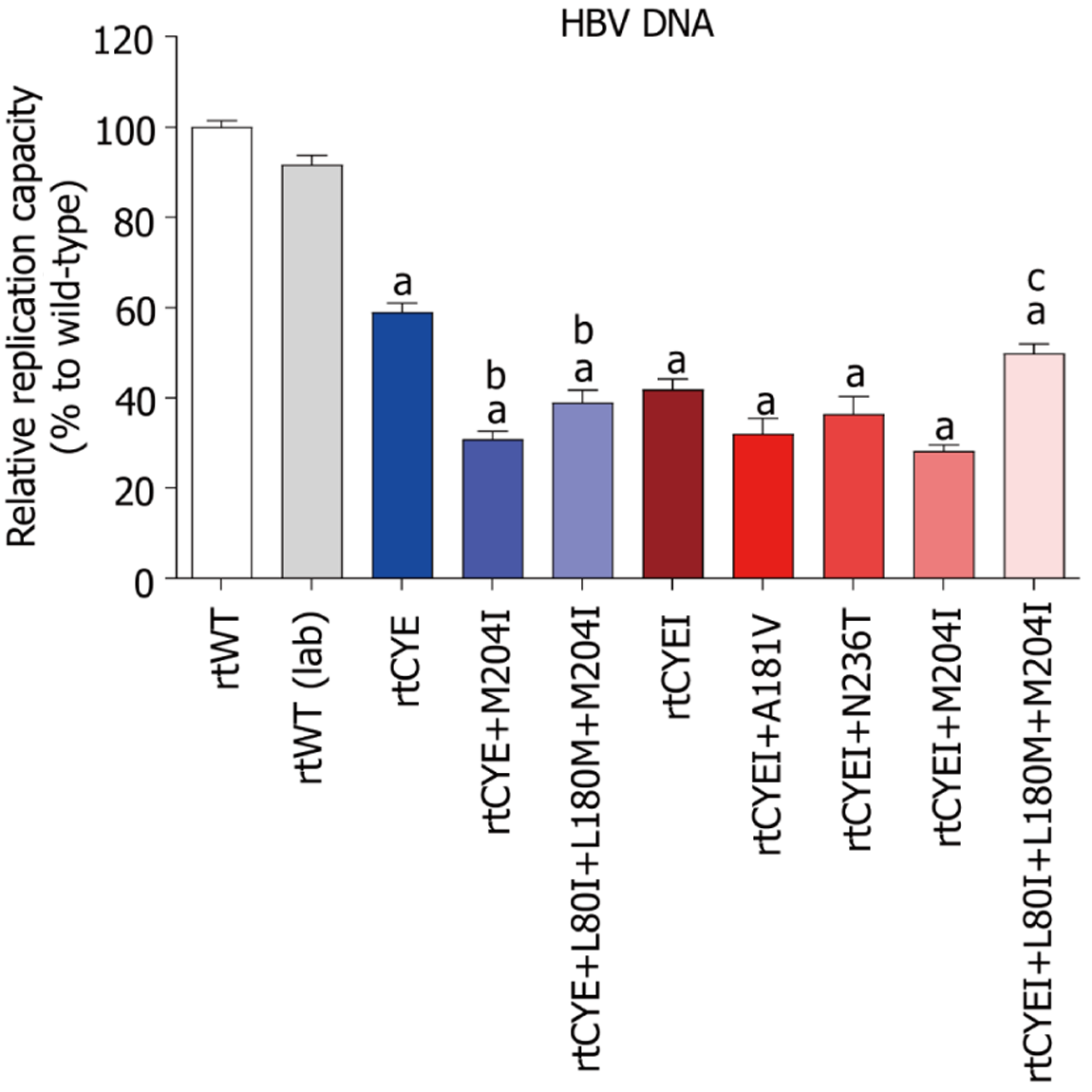
Phenotypic analysis was performed on representative viral strains with typical mutational patterns that emerged in rtCYE/rtCYEI-harboring patients. These included viral strains with rtCYE/rtCYEI mutations alone, combined with LAM-/LdT-/ADV-resistance mutations, as well as the WT. As shown in Figure 2, all rtCYE- and rtCYEI-containing mutants had significantly reduced HBV replication capacity compared with the rtWT strain in the absence of drug pressure. Specifically, the replication capacities of rtCYE and rtCYEI mutants were 58.9% and 41.8% of the rtWT, respectively; the replication capacities of rtCYE+M204I and rtCYE+L80I+L180M+M204I mutants were 30.8% and 38.9% of the rtWT, respectively; and the replication capacities of rtCYEI+A181V, rtCYEI+N236T, rtCYEI+M204I, and rtCYEI+L80I+L180M+M204I mutants were 32.0%, 36.4%, 28.2%, and 49.8% of the rtWT, respectively. The original data are shown in Supplementary Table 1.
The EC50 of rtCYE, rtCYEI, rtCYE+LAMr, and rtCYEI+LAMr mutants decreased by less than 2-fold, suggesting similar TDF susceptibility of the mutants compared with the rtWT strain. The mutants rtCYEI+ADVr (rtA181V/N236T) showed reduced TDF susceptibility of 6.2- to 9.9-fold compared with the rtWT strain (Table 4). The original data are shown in Supplementary Table 2.
| Viral strain | Tenofovir disoproxil fumarate | ||
| EC50 (μmol/L)2 | Fold3 | P value1 | |
| rtWT (wild-type) | 5.79 ± 1.59 | 1.0 | - |
| rtWT (laboratory) | 5.90 ± 1.95 | 1.0 | 0.93 |
| rtCYE | 7.41 ± 0.97 | 1.3 | 0.09 |
| rtCYEI | 8.51 ± 1.29 | 1.5 | < 0.05 |
| rtCYE+M204I | 6.95 ± 0.64 | 1.2 | 0.17 |
| rtCYE+L80I+L180M+M204I | 6.92 ± 1.69 | 1.2 | 0.31 |
| rtCYEI+M204I | 8.92 ± 0.87 | 1.5 | < 0.05 |
| rtCYEI+L80I+L180M+M204I | 10.23 ± 1.01 | 1.8 | < 0.05 |
| rtCYEI+A181V | 36.15 ± 5.99 | 6.2 | < 0.05 |
| rtCYEI+N236T | 57.33 ± 5.30 | 9.9 | < 0.05 |
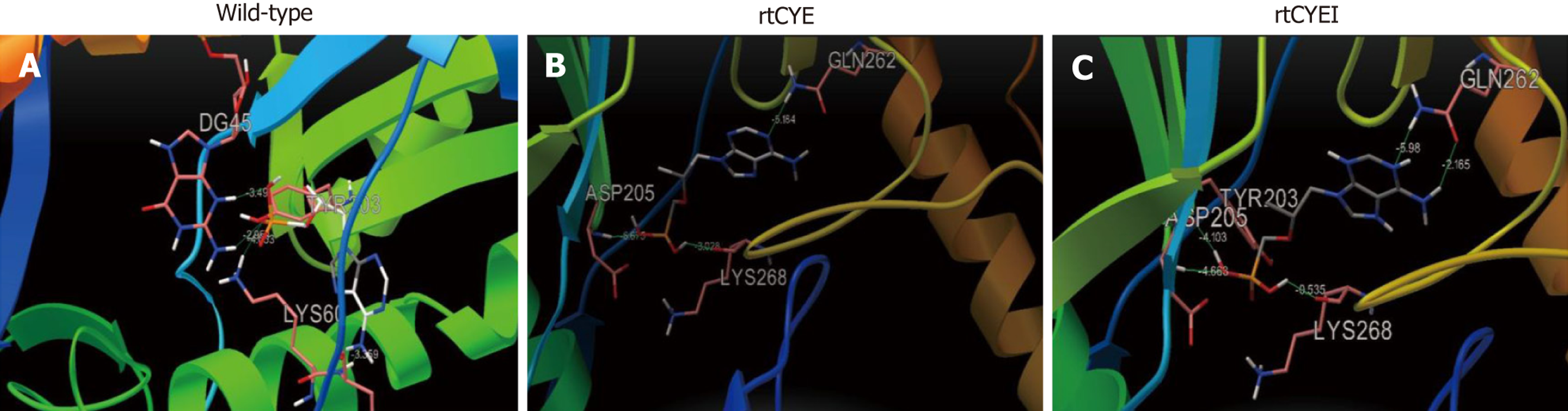
The effect of rtCYE and rtCYEI mutations on the binding ability of HBV RT to TDF was evaluated using Autodock software. The modeled structures of WT and mutant RT containing rtCYE, or rtCYEI mutations were used as receptors to analyze their binding energy to TDF as a ligand (Figure 3). The lower binding energy signifies the more stable conformation. The binding energy of the rtWT to TDF was -6.18 kcal/mol, while the binding energies of the rtCYE and rtCYEI mutants were -6.36 kcal/mol and -6.83 kcal/mol, respectively, with no significant difference observed (P = 0.335).
Typical viral resistance mutations have several distinguishing characteristics, including an association with drug therapy, resulting in viral rebound or inadequate virological response, emergence in multiple patients exposed to the drug, the ability to confer viral resistance in in vitro phenotypic assays, and reversion to WT sequences in the absence of selective antiviral pressure[21]. As few studies have questioned whether HBV rtCYEI mutations have these distinguishing characteristics, more convincing clinical and laboratory evidence is needed for classification. The present study provides the first data on the clinical prevalence of HBV rtCYE/rtCYEI mutations in a large cohort of Chinese patients who underwent resistance testing. The reasons for studying rtCYE and rtCYEI mutations simultaneously were as follows: (1) rtCYE mutations have been reported to be directly associated with TDF resistance in two patients[12]; (2) The rtCYE mutations conferred decreased TDF sensitivity and served as the backbone of rtCYEI mutations[14]; and (3) Among the 12 patients harboring rtCYE/rtCYEI mutations in our study, the co-existence of rtCYE and rtCYEI mutants was identified in 4 patients as shown in Table 2, suggesting that the emergence of the two mutational patterns is closely related.
Consistent with previous studies[12,16], our results showed that rtCYE/rtCYEI mutations were rarely detected (0.042% of all patients) and were only observed in genotype C HBV. The prevalence of the mutations was mainly due to the low incidence and genotype-bias of rtD134E mutation (Table 1). In China, genotypes B and C HBV dominate the epidemiological landscape, exhibiting distinct clinical and virological profiles. Comparative analyses reveal that genotype C HBV demonstrates higher propensity for multidrug resistance mutations and accelerated disease progression compared to genotype B HBV[22,23]. Notably, genotype C HBV exhibits significantly elevated LAM resistance rates and a characteristic ADV resistance profile marked by increased rtA181V mutations but reduced rtN236T mutations[22]. Importantly, higher baseline resistance rates were observed in treatment-naïve genotype C HBV-infected patients, suggesting potential innate resistance mechanisms[23]. Our findings emphasized the necessity for enhanced resistance surveillance and genotype-guided therapeutic strategies in genotype C HBV-infected patients. Additionally, the rtD134E mutation ("E") exhibits a lower prevalence in genotype B (0.21%, Table 1) compared to genotype C (2.55%, Table 1), suggesting a geno
Regarding phenotypic resistance, a small decrease (2- to 9-fold increase in EC50) in vitro ADV susceptibility may confer clinical resistance, and an increase in the EC50 of > 10-fold (usually > 50-fold) for ETV and TDF, and > 500-fold for LAM are required to confer clinical resistance[8,26]. In our study, the rtCYE and rtCYEI mutants exhibited less than a 2-fold change in TDF susceptibility with or without concomitant LAMr. In contrast, the rtCYEI+A181V/N236T mutant showed a 6.2- to 9.9-fold decrease in TDF susceptibility. This decrease was attributed primarily to the rtA181V/N236T mutations, as both ADV and TDF are NAs and ADVr has been documented to slightly reduce TDF susceptibility[16,27].
Our results indicate that rtCYE/rtCYEI mutations do not directly cause TDF resistance. First, none of the 12 rtCYE/rtCYEI mutation-positive patients had a history of TDF exposure, let alone the association of viral rebound with TDF treatment. This observation suggests that rtCYE/rtCYEI mutations may occur in TDF treatment naïve patients, as reported by Blackard et al[15]. Second, the pre-existence of the rtCYE/rtCYEI mutations did not have a negative impact on subsequent TDF therapy in the observation of 3 representative patients (P2, P4, and P6). Third, phenotypic analysis showed that the rtCYE/rtCYEI mutants had little or no effect on TDF sensitivity, as recently reported by Liu et al[28]. In addition, molecular modeling showed that the binding energy changes in the RT regions of rtCYE and rtCYEI mutants were limited compared with the WT RT region. This provides circumstantial evidence that the rtCYE and rtCYEI mutations have no robust influence on TDF activity.
Compared with ADV, TDF has a similar molecular structure and anti-HBV effects, but far fewer side effects, which greatly increases its dosage (300 mg/day for TDF vs 10 mg/day for ADV), meaning that the genetic barrier threshold for TDF resistance is much higher than that for ADV resistance. It was estimated that a 60-fold decrease in drug susceptibility is required to satisfy the definition of a primary TDF-resistant mutation/mutation pattern. In addition to resistance mutations, certain mutations may negatively affect the efficacy of NAs therapy. One example is the rtA181T/sW172stop mutation, which forms a truncated S protein, leading to intracellular retention of viral particles and atypical extent of virological breakthrough[29]. Another example is the rtS78T/sC69stop mutation, which has been reported to emerge in ETV+TDF-refractory patients. It has limited effects on drug susceptibility but has a potential influence on anti-HBV response[30]. Moreover, the efficacy of NAs therapy could be weakened by HBV mutations that change specific T-cell epitopes and viral replication capacity[28,31].
Differences in methodology may cause deviations in results. There are two popular methods for phenotypic analysis of HBV drug resistance. One is a Southern blotting-based method used by the South Korean investigation team to detect intracellular HBV replicative intermediates. This method has the advantages of good specificity and strong visualization, but it has disadvantages such as low detection sensitivity and time-consuming operation. The other is the real-time PCR-based method used by a United States investigation team to detect extracellular productive HBV DNA in culture. This method has the advantages of high detection sensitivity and time-saving operation but its specificity is relatively low. We have developed a real-time PCR-based method that uses anti-HBc/protein A+G to capture core particles and determine the intracellular replicative intermediates of HBV in these core particles. Comparable results were obtained in deter
This study was restricted by serial serum sample collection and longitudinal clinical information acquisition, which prevented us from monitoring the dynamic changes of rtCYE/rtCYEI mutants in most patients of interest. As the enrolled patients were the population for clinically required drug resistance testing, the number of patients treated with TDF in our study was small.
Taken together, our findings suggest that HBV rtCYE/rtCYEI mutations were not specifically associated with TDF treatment and the mutations alone were not sufficient to confer TDF resistance. This conclusion is consistent with previous reports from other investigators[15,28]. Our study provides new insights into clinical HBV drug resistance management.
In summary, this study highlights several key findings regarding rtCYE/rtCYEI mutations in chronic HBV-infected patients. First, these mutations were rarely observed (0.042%) in NAs-treated cohorts. Notably, they could emerge spontaneously in treatment-naïve patients without prior TDF exposure while retaining susceptibility to TDF. Functional analysis revealed that rtCYE/rtCYEI mutants exhibited significantly impaired viral replication capacity, with reductions of 41.1% to 71.8% compared to rtWT HBV. Although TDF susceptibility remained largely preserved for single rtCYE/rtCYEI mutants, combination with classical resistance mutations (rtA181V/rtN236T) led to moderate reductions in TDF sensitivity (6.2- and 9.9-fold increases in EC50, respectively). Collectively, these findings suggest that rtCYE/rtCYEI mutations alone have limited clinical impact on TDF efficacy but warrant vigilance when co-occurring with established resistance-associated substitutions. The natural replication deficiency of these mutants may further constrain their transmission potential and clinical significance.
We are grateful to Jiu-Zeng Dai and Zeng-Tao Yao for their contributions to resistance testing in clinical practice.
| 1. | Polaris Observatory Collaborators. Global prevalence, treatment, and prevention of hepatitis B virus infection in 2016: a modelling study. Lancet Gastroenterol Hepatol. 2018;3:383-403. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1260] [Cited by in RCA: 1216] [Article Influence: 173.7] [Reference Citation Analysis (2)] |
| 2. | Hou J, Wang G, Wang F, Cheng J, Ren H, Zhuang H, Sun J, Li L, Li J, Meng Q, Zhao J, Duan Z, Jia J, Tang H, Sheng J, Peng J, Lu F, Xie Q, Wei L; Chinese Society of Hepatology, Chinese Medical Association; Chinese Society of Infectious Diseases, Chinese Medical Association. Guideline of Prevention and Treatment for Chronic Hepatitis B (2015 Update). J Clin Transl Hepatol. 2017;5:297-318. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 154] [Cited by in RCA: 186] [Article Influence: 23.3] [Reference Citation Analysis (0)] |
| 3. | GBD 2019 Hepatitis B Collaborators. Global, regional, and national burden of hepatitis B, 1990-2019: a systematic analysis for the Global Burden of Disease Study 2019. Lancet Gastroenterol Hepatol. 2022;7:796-829. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 416] [Cited by in RCA: 416] [Article Influence: 138.7] [Reference Citation Analysis (0)] |
| 4. | Liu Y, Chen R, Liu W, Si L, Li L, Li X, Yao Z, Liao H, Wang J, Li Y, Zhao J, Xu D. Investigation of multidrug-resistance mutations of hepatitis B virus (HBV) in a large cohort of chronic HBV-infected patients with treatment of nucleoside/nucleotide analogs. Antiviral Res. 2021;189:105058. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 9] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 5. | Chinese Society of Infectious Diseases; Chinese Medical Association; Chinese Society of Hepatology, Chinese Medical Association. [The guidelines of prevention and treatment for chronic hepatitis B (2019 version)]. Zhonghua Gan Zang Bing Za Zhi. 2019;27:938-961. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 112] [Reference Citation Analysis (0)] |
| 6. | Jiang C, Zhang ZH, Li JX. Current status of drug therapy for chronic hepatitis B. World J Gastroenterol. 2025;31:99443. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 7. | Terrault NA, Lok ASF, McMahon BJ, Chang KM, Hwang JP, Jonas MM, Brown RS Jr, Bzowej NH, Wong JB. Update on Prevention, Diagnosis, and Treatment of Chronic Hepatitis B: AASLD 2018 Hepatitis B Guidance. Clin Liver Dis (Hoboken). 2018;12:33-34. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 129] [Cited by in RCA: 223] [Article Influence: 31.9] [Reference Citation Analysis (0)] |
| 8. | Lok AS, Zoulim F, Locarnini S, Bartholomeusz A, Ghany MG, Pawlotsky JM, Liaw YF, Mizokami M, Kuiken C; Hepatitis B Virus Drug Resistance Working Group. Antiviral drug-resistant HBV: standardization of nomenclature and assays and recommendations for management. Hepatology. 2007;46:254-265. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 359] [Cited by in RCA: 364] [Article Influence: 20.2] [Reference Citation Analysis (0)] |
| 9. | Zoulim F, Locarnini S. Optimal management of chronic hepatitis B patients with treatment failure and antiviral drug resistance. Liver Int. 2013;33 Suppl 1:116-124. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 56] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 10. | Hayashi S, Murakami S, Omagari K, Matsui T, Iio E, Isogawa M, Watanabe T, Karino Y, Tanaka Y. Characterization of novel entecavir resistance mutations. J Hepatol. 2015;63:546-553. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 43] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 11. | Liu Y, Zhou Y, Li X, Niu M, Chen R, Shao J, Si L, Luo D, Lin Y, Li L, Zhang K, Xiao X, Xu Z, Liu M, Lu M, Zoulim F, Xu D. Hepatitis B virus mutation pattern rtL180M+A181C+M204V may contribute to entecavir resistance in clinical practice. Emerg Microbes Infect. 2019;8:354-365. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 19] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 12. | Park ES, Lee AR, Kim DH, Lee JH, Yoo JJ, Ahn SH, Sim H, Park S, Kang HS, Won J, Ha YN, Shin GC, Kwon SY, Park YK, Choi BS, Lee YB, Jeong N, An Y, Ju YS, Yu SJ, Chae HB, Yu KS, Kim YJ, Yoon JH, Zoulim F, Kim KH. Identification of a quadruple mutation that confers tenofovir resistance in chronic hepatitis B patients. J Hepatol. 2019;70:1093-1102. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 94] [Cited by in RCA: 99] [Article Influence: 16.5] [Reference Citation Analysis (0)] |
| 13. | Revill PA, Tu T, Netter HJ, Yuen LKW, Locarnini SA, Littlejohn M. The evolution and clinical impact of hepatitis B virus genome diversity. Nat Rev Gastroenterol Hepatol. 2020;17:618-634. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 107] [Article Influence: 21.4] [Reference Citation Analysis (2)] |
| 14. | Lee JH, Kim KH. Reply to: "In response to identification of a quadruple mutation that confers tenofovir resistance in chronic hepatitis B patients.". J Hepatol. 2019;71:1260-1261. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 15. | Blackard JT, Kwara A, Sherman KE. In response to identification of a quadruple mutation that confers tenofovir resistance in chronic hepatitis B patients. J Hepatol. 2019;71:1259-1260. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 4] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 16. | Liu Y, Chang S, Martin R, Flaherty J, Mo H, Feierbach B. Characterization of Hepatitis B virus polymerase mutations A194T and CYEI and tenofovir disoproxil fumarate or tenofovir alafenamide resistance. J Viral Hepat. 2021;28:30-39. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 11] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 17. | Chinese Society of Hepatology; Chinese Medical Association; Chinese Society of Infectious Diseases, Chinese Medical Association. Guideline on prevention and treatment of chronic hepatitis B in China (2005). Chin Med J (Engl). 2007;120:2159-2173. [PubMed] |
| 18. | Zoulim F, Locarnini S. Hepatitis B virus resistance to nucleos(t)ide analogues. Gastroenterology. 2009;137:1593-608.e1. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 517] [Cited by in RCA: 540] [Article Influence: 33.8] [Reference Citation Analysis (0)] |
| 19. | Durantel D, Carrouée-Durantel S, Werle-Lapostolle B, Brunelle MN, Pichoud C, Trépo C, Zoulim F. A new strategy for studying in vitro the drug susceptibility of clinical isolates of human hepatitis B virus. Hepatology. 2004;40:855-864. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 83] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 20. | Zoulim F. In vitro models for studying hepatitis B virus drug resistance. Semin Liver Dis. 2006;26:171-180. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 40] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 21. | Yang H, Westland CE, Delaney WE 4th, Heathcote EJ, Ho V, Fry J, Brosgart C, Gibbs CS, Miller MD, Xiong S. Resistance surveillance in chronic hepatitis B patients treated with adefovir dipivoxil for up to 60 weeks. Hepatology. 2002;36:464-473. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 141] [Cited by in RCA: 130] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 22. | Li X, Liu Y, Xin S, Ji D, You S, Hu J, Zhao J, Wu J, Liao H, Zhang XX, Xu D. Comparison of Detection Rate and Mutational Pattern of Drug-Resistant Mutations Between a Large Cohort of Genotype B and Genotype C Hepatitis B Virus-Infected Patients in North China. Microb Drug Resist. 2017;23:516-522. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 15] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 23. | Wang L, Han F, Duan H, Ji F, Yan X, Fan Y, Wang K. Hepatitis B virus pre-existing drug resistant mutation is related to the genotype and disease progression. J Infect Dev Ctries. 2017;11:727-732. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 5] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 24. | Villet S, Pichoud C, Villeneuve JP, Trépo C, Zoulim F. Selection of a multiple drug-resistant hepatitis B virus strain in a liver-transplanted patient. Gastroenterology. 2006;131:1253-1261. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 135] [Cited by in RCA: 132] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 25. | Villet S, Billioud G, Pichoud C, Lucifora J, Hantz O, Sureau C, Dény P, Zoulim F. In vitro characterization of viral fitness of therapy-resistant hepatitis B variants. Gastroenterology. 2009;136:168-176.e2. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 52] [Cited by in RCA: 58] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 26. | Svarovskaia ES, Curtis M, Zhu Y, Borroto-Esoda K, Miller MD, Berg T, Lavocat F, Zoulim F, Kitrinos KM. Hepatitis B virus wild-type and rtN236T populations show similar early HBV DNA decline in adefovir refractory patients on a tenofovir-based regimen. J Viral Hepat. 2013;20:131-140. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 21] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 27. | Liu Y, Wu C, Chen R, Li X, Xu Z, Li Q, Li L, Wang FS, Yang D, Lu M, Xu D. Molecular cloning and phenotypic analysis of drug-resistance mutants with relevant S-region variants of HBV for a patient during 189-month anti-HBV treatment. Antivir Ther. 2019;24:237-246. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 5] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 28. | Liu T, Sun Q, Gu J, Cen S, Zhang Q. Characterization of the tenofovir resistance-associated mutations in the hepatitis B virus isolates across genotypes A to D. Antiviral Res. 2022;203:105348. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 12] [Reference Citation Analysis (0)] |
| 29. | Warner N, Locarnini S. The antiviral drug selected hepatitis B virus rtA181T/sW172* mutant has a dominant negative secretion defect and alters the typical profile of viral rebound. Hepatology. 2008;48:88-98. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 179] [Cited by in RCA: 187] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 30. | Shirvani-Dastgerdi E, Winer BY, Celià-Terrassa T, Kang Y, Tabernero D, Yagmur E, Rodríguez-Frías F, Gregori J, Luedde T, Trautwein C, Ploss A, Tacke F. Selection of the highly replicative and partially multidrug resistant rtS78T HBV polymerase mutation during TDF-ETV combination therapy. J Hepatol. 2017;67:246-254. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 49] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 31. | Zhang Y, Ren Y, Wu Y, Zhao B, Qiu L, Li X, Xu D, Liu J, Gao GF, Meng S. The L60V variation in hepatitis B virus core protein elicits new epitope-specific cytotoxic T lymphocytes and enhances viral replication. J Virol. 2013;87:8075-8084. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 21] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 32. | Phillips S, Chokshi S, Chatterji U, Riva A, Bobardt M, Williams R, Gallay P, Naoumov NV. Alisporivir inhibition of hepatocyte cyclophilins reduces HBV replication and hepatitis B surface antigen production. Gastroenterology. 2015;148:403-14.e7. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 34] [Cited by in RCA: 42] [Article Influence: 4.2] [Reference Citation Analysis (0)] |