ORGANISMAL COMMUNITIES
Cell communities in multicellular organisms are shaped by a fundamental organizing principle, instructing the relative sizes and reciprocal relationships to be enacted and maintained over time among different cell types. Such a seemingly simple fact represents the very essence of individual multicellular communities and mechanisms overlooking their correct implementation are central during development and throughout life[1-3]. For example, when part of liver tissue is lost (due to surgery or any injury), residual hepatocytes are immediately alerted and awakened from their quiescent state to enter cell cycle, divide and replenish the missing parenchyma. Conversely, following liver hyperplasia a wave of hepatocyte deletion ensues upon withdrawal of the inciting stimulus, until the original tissue mass is reinstated.
While the capacity to maintain stable tissue mass is remarkable, there is still more to it. In fact, another layer of complexity has been added to the above mechanism following the realization that quality control strategies are also at play to optimize cell fitness in tissue composition. Thus, not only the number of cells in a given tissue is constantly under control, but their functional efficiency is also monitored and actively selected for in order to limit any time-dependent decline. One such strategy is cell competition, consisting in the confrontation of homotypic cells with varying levels of fitness and resulting in the elimination of the weaker (losers) carried out by the stronger counterparts (winners). While primarily aimed at maintaining optimal functional proficiency in cell populations of normal tissues, it has been suggested that mechanisms driving cell competition can also be hijacked and exploited by pre-neoplastic and/or neoplastic cells to manifest their aggressive and dominant phenotype[4]. The aim of this review is to discuss the possible role of cell competition during hepato-carcinogenesis.
WHAT IS CELL COMPETITION
In its simplest definition, cell competition refers to a process whereby cells in a given tissue, which would be otherwise viable and functional, are instead outcompeted and cleared by the presence of a functionally more proficient population. The first report describing a similar scenario dates back to the mid the 1970s and refers to the elimination of Minute-mutant cells when confronted with wild type counterparts in the imaginal disk of Drosophila melanogaster[5]. The Minute mutation affects ribosomal protein genes and translates into a slower growth rate of heterozygous mutant cells. Several other mutations were subsequently identified to induce a loser phenotype in presence of wild type cells, including those involving basic cellular functions such as tissue patterning, protein translation and cell signaling[6].
In an attempt to outline the boundaries of cell competition, a series of biological features have been proposed to be associated with this phenomenon[7]. Firstly, cell competition is context-dependent, i.e., the fate of each cell type, the winner and the loser, results from their reciprocal interaction, be it direct or indirect; this is arguably the single most relevant attribute of cell competition, as well as the most complex to enact. Secondly, the winner cell proliferates following induction of cell death (apoptosis) of the loser cell, i.e., the two opposite events are temporally and mechanistically coordinated. As a corollary, biological forces driving cell competition tend to maintain appropriate tissue size and pattern. Furthermore, interactions conducive to cell competition occur within a relatively short range, being strongest at the interface between winner and loser cells. Last but not least, cell competition is restricted within defined developmental compartments[1], i.e., it occurs within discrete tissue boundaries that cannot be overridden by winner cells.
Within this definition, mechanism(s) governing cell competition are intertwined, at least in part, with those overlooking the fine balance between cell gain and cell loss, which in turn determine tissue size in any organ and organism[1,2]. Since alterations of the latter mechanisms represent a hallmark of neoplastic disease, it is all the more reasonable to propose that cancer cell populations may coopt strategies involving cell competition in order to selectively emerge vis a vis the surrounding counterparts[8,9].
COMPETITION FOR WHAT
Cell competition can only occur when a critical degree of phenotypic heterogeneity is present within a homotypic cell population; in addition, a limit must exist in the availability of whatever resource these cells are competing for. Molecular analysis carried out at the resolution of single cells has revealed that cell heterogeneity at genetic and epigenetic levels is indeed far more pervasive than previously thought even in normal adult tissues, raising the possibility that cell competition may not be a rare phenomenon[6]. Cells can compete for nutrients, growth factors and ultimately space, given the size constraints imposed on any tissue by homeostatic control mechanisms[3].
A paradigmatic example in which the principle of cell competition is at play is the process of antibody affinity maturation in lymphoid germinal centers[8,9]. Heterogeneity is generated through somatic hypermutation in the gene coding for the B-cell receptor. The limited resource is represented by antigen availability: The lower the antigen concentration, the higher the affinity of the resulting antibodies. Lymphocytes are in fact positively selected through the binding of their mutated receptor to antigen, which in turn is dictated by the degree of affinity of the former to the latter. Lymphocytes that are unable to reach for antigen trough their receptor die by apoptosis. Thus, the competitive fitness of B lymphocyte clones rests on their ability to bind a rescuing or “trophic” factor, which is epitomized, in this case, by the incoming antigen.
The above sequence of events is similar, in essence, to the one described in the Minute-mutant of Drosophila wing imaginal disk referred to above, which is considered as a classical model of cell competition. It was in fact proposed that in this system, slow-growing Minute-mutant cells have a disadvantage in competing for a survival signal, and this leads to increased expression of a pro-apoptotic cascade and final clearance of mutants by wild type cells[10]. Parenthetically, it is important to point out that the latter interpretation has been questioned by subsequent studies suggesting that differences in growth rate per se between mutant and wild type cells are sufficient to account for their unbalanced contribution to wing development, while cell competition per se would not appear to play a major role in the process[5]. These findings also indicate that a slower growth rate does not necessarily entail a loser phenotype compared to faster homotypic counterparts, i.e., for cell competition to be enforced other critical differences must be present.
WHAT IS CELL FITNESS
The latter consideration brings us to the core issues pertaining cell competition: What parameters are measured between winner and loser cells to assess relative fitness and how is this accomplished? We must acknowledge that only scattered information is available so far to answer these questions. Given that evolutionary processes (including cell competition) have selected over time for cells with better and better functional proficiency, it follows that normal tissues are populated with cells with near-optimal performing capacity. This in turn implies that any damage to any cell will likely result in a decreased fitness, laying the basis for its clearance by neighboring normal cells through cell competition[11-13]. Thus, any damage above a given threshold can potentially trigger cell deletion. However, this is not a cell-autonomous process, as postulated for the classical p53-dependend apoptosis elicited in response to DNA damage[14]. Rather, it relies on the presence of surrounding cells and results from the confrontation of different levels of cellular fitness, within the biological boundaries of cell competition[15,16]. There is obviously a fundamental difference between these two strategies leading to clearance of damaged cells. While cell-autonomous mechanisms operate at single-cell level and do not take into account overall tissue function, deletion of damaged cells via cell competition is only triggered when fitter cells are available and can possibly replace the ones that are lost. The latter consideration supports the contention that cell competition is an integral part of regulatory networks overlooking tissue maintenance and homeostasis[17,18]. Specific strategies to pursue this goal can vary even in the same tissue during different developmental phases. In mouse skin, the early embryonic epithelium is single-layered and loser cell disposal is carried out through their direct phagocytosis by surrounding winner cells; however, as the epidermis becomes stratified, loser cells are extruded from the basal layer along the differentiation conveyor and are eventually shed out[19].
Sensing relative cellular fitness is therefore an essential step in the process of cell competition[6]. One of the parameters that has emerged as relevant in this regard is the expression of Myc protein[20-23]. Higher cellular levels of this transcription factor confer a winner phenotype both during Drosophila development[20], in early mammalian embryo[21,24-26] and in adult, post-mitotic tissues such heart[27]. Furthermore, overexpression of Myc is associated with a super-competitor phenotype, which is able to outcompete wild type cells[20]. Similar to Myc, other genes important for cell anabolism have been implicated as triggers of cell competition, including those involved in the Hippo, Wnt/Wingless, Ras/mitogen-activated protein kinases and Janus kinase/signal transducers and activators of transcription (JAK/STAT) pathways, among others[21]. Conversely, defects in genes implicated in the determination of cell polarity and tissue patterning impose a loser phenotype on the affected cells in presence of wild type counterparts[28,29].
So far, the best characterized direct sensor of cell fitness is the Flower system[30,31]. It consists of three splice isoform proteins, Flowerubi, FlowerLoseA and FlowerLoseB and only the former is expressed under steady state conditions. However, when cell fitness decreases, expression of Flowerubi is down-regulated, while levels of both FlowerLoseA and FlowerLoseB increase, generating what has been referred to as the “flower code”, which targets cells for survival (Flowerubi) or apoptosis (FlowerLoseA and FlowerLoseB)[23,24]. Human Flower isoforms have recently been reported and evidence was presented that a similar winner/loser code is also operative in human cells[32].
CELL COMPETITION IN THE LIVER
Virtually no information is available on the possible role of cell competition in the liver during normal development and throughout post-natal life. However, a few studies, mostly using cell transplantation systems, have indicated that hepatic tissue appears to be susceptible to undergo this process. About 20 years ago, we proposed that a mechanism consistent with cell competition was possibly involved in a newly developed model of massive liver repopulation[33,34]. In this experimental system, animals (rats) are treated with retrorsine, a naturally occurring pyrrolizidine alkaloid that causes persistent DNA damage associated with a chronic mitotic block in targeted hepatocytes. It is noteworthy that damaged hepatocytes are able to sustain normal liver function and in fact retrorsine treated animals survive for up to 2 years[35]. However, the most intriguing finding was observed following cell transplantation. When normal syngeneic hepatocytes were orthotopically delivered after treatment with the alkaloid, they are able to massively replace endogenous parenchymal cells, with greater that 90% repopulation at the end of the process[33]. It was suggested that “the presence of normal transplanted cells may trigger selective deletion of RS-damaged resident hepatocytes, possibly through apoptosis”[33], a process that is fully consistent with cell competition (Figure 1). A few years later, a study by Oertel et al[36] proposed cell competition as the basis for the selective expansion of transplanted normal embryonic hepatic cells in the liver of syngeneic adult rats[36], similar to results obtained following transplantation of young adult hepatocytes in the liver of aged recipients[37].
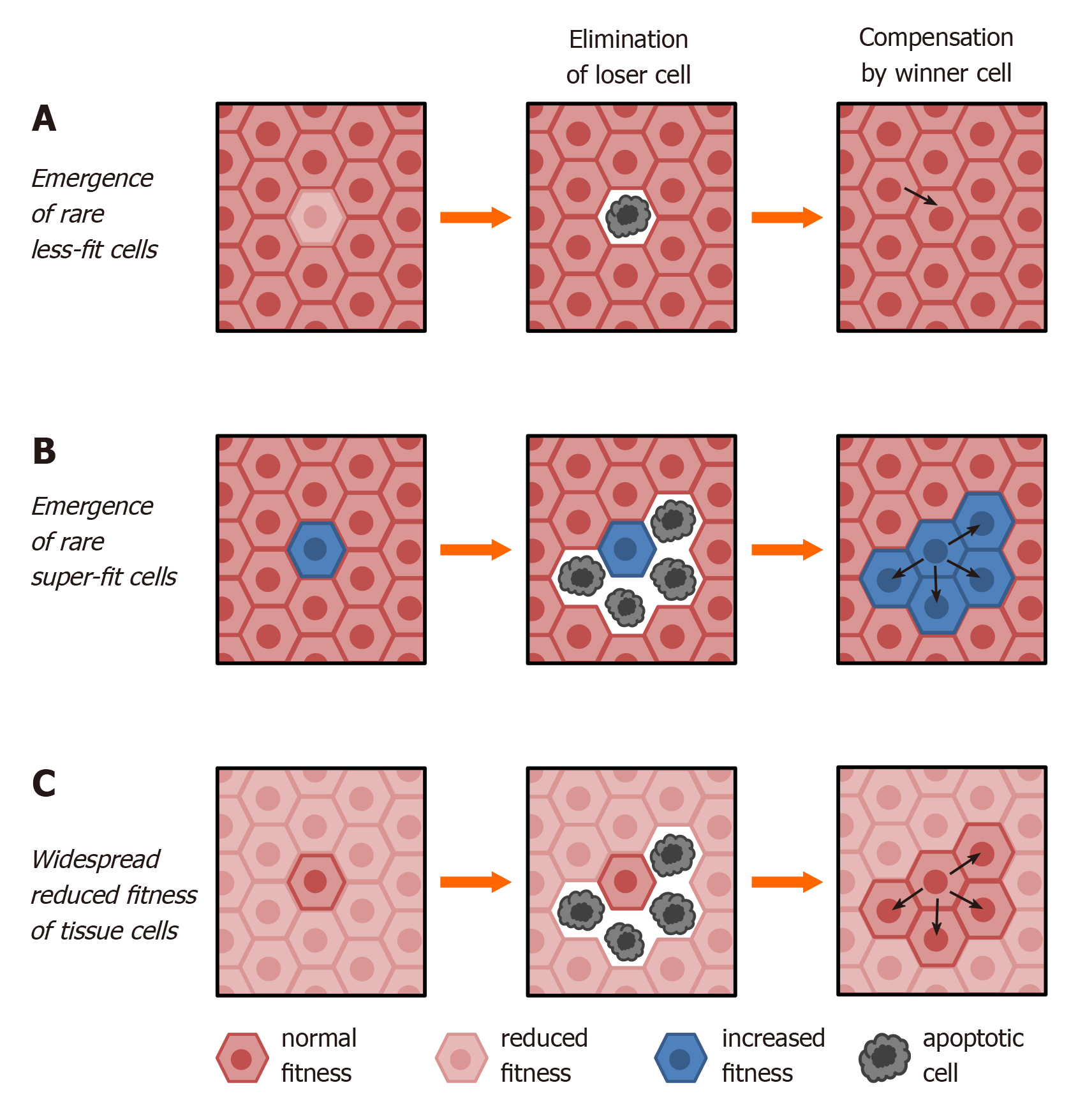
Figure 1 Modes of canonical cell competition.
A: Wild type cells (dark brown) are endowed with higher fitness (winner phenotype) compared to rare altered cells (loser phenotype
While common molecular pathways involved in cell competition were not investigated under the above experimental conditions, biological coordinates of the described phenomena support the hypothesis that liver repopulation by transplanted hepatocytes is, at least in part, the outcome of a differential fitness comparison between resident and donor-derived cells. Accordingly, a cell-autonomous decrease in proliferative competitiveness was reported in aged vs young hepatocytes upon transplantation in the same microenvironment in vivo[38].
If cell transplantation is not performed, a slow process of repopulation sustained by endogenous hepatocytes occurs in rat liver exposed to RS, giving rise to regenerative nodules[39,40]. A similar scenario has also been reported in the liver of patients affected by type 1 tyrosinemia[41], the human counterpart of a well characterized mouse model of liver repopulation[42]. Whether cell competition is driving and/or contributing to either or both processes is an intriguing possibility that remains to be explored.
CELL COMPETITION IN THE PATHOGENESIS OF CANCER
We are now back to the central question of this review: Is there a role for cell completion in cancer and, more specifically, in liver carcinogenesis, From a molecular standpoint, numerous pathways that have been proposed to have a role in cell competition (summarized above) are also implicated in the pathogenesis of neoplastic disease. It would appear therefore quite reasonable to expect that alterations in these pathways might cause a defective control in the mechanisms of cell competition and, almost inevitably, a parallel increase in the risk of cancer. As an example, the Myc protein is often overexpressed in human cancer[43] and is also a known driver of cell competition[7]. Accordingly, cells with up-regulated Myc can express a super-competitor phenotype[23] and this could contribute, at least theoretically, towards a neoplastic behavior[22].
However, the potential role of cell competition in the multistep process of neoplastic development is far from being confined to situations of altered molecular control. In fact, the emerging picture is that biological mechanisms underlying cell competition under normal conditions are directly relevant to the pathogenesis of cancer from early stages to advanced disease[4]. A compelling case is the extrusion of altered, potentially pre-neoplastic cells from epithelia orchestrated by normal surrounding counterparts and referred to as epithelial defense against cancer[12]. For example, normal cells were able to induce Warburg-like metabolic changes in RasV12-transformed cells, leading to their removal from mouse intestinal epithelium[11]. Similarly, cell competition by normal cells is able to eliminate scrib mutant cells from Drosophila imaginal disk, while in the absence of wild type counterparts mutant clones do not die and progress to form tumors[29].
As one can predict, this protective strategy is not foolproof and, more to the point, it can be altered in its efficiency by environmental influences. A recent study indicated that removal of RasV12-transformed cells from mouse intestine was decreased following feeding a high fat diet (HFD), due to altered lipid metabolism and HFD-induced inflammatory changes; treatment with aspirin was able to mitigate HFD negative effects on transformed cell clearance[44]. Similarly, hematopoietic stem and progenitor cells expressing a mutant p53 displayed a growth advantage vs wild type counterparts when transplanted in mice following exposure to mild irradiation, while no such advantage was evident upon transplantation of the same cell populations in untreated recipients[15]. Furthermore, we have proposed that aging is associated with a generalized decrease in the efficiency of mechanisms overlooking maintenance of cell fitness, possibly including cell competition[17,45,46]. In addition, specific genetic alterations might be positively selected, as opposed to eliminated, under environmental conditions favoring their phenotype. This could partly account for the pervasive presence of aberrant clonal expansions in aged human tissues[47] and/or in association with disease states such as ulcerative colitis[48,49]. It was shown that organoids derived from ulcerative colitis patients are populated by genetically altered cell clones that are adapted to an inflammatory microenvironment, i.e., they are fitter to that environment compared with wild type intestinal cells[48,49]. Whether in these instances cell competition is at play is an important question that remains to be addressed. Relevant to this issue, a mechanistic association was reported between alterations in intestinal barrier integrity, aging, dietary regimen and the efficiency of cell competition[46].
Direct evidence that cell competition is indeed mechanistically exploited by cancer cells during growth and metastatic spread was recently presented[32,50]. Human cancer cells were shown to express the Flower code of a winner cell phenotype and inhibition of the latter resulted in reduced tumor growth and increased response to chemotherapy[32].
CELL COMPETITION IN LIVER CARCINOGENESIS
Given the limited amount of information available regarding the role of cell competition in liver under normal conditions, it is not surprising that a similar consideration also applies to the process of liver cancer development. Several years ago we have pointed out the existence of intriguing analogies between the process of liver repopulation by normal hepatocytes and carcinogenesis[34]. For example, several of the available experimental models of massive liver repopulation are also prone to develop neoplastic disease, including the RS-based model developed by our research group and referred to above. Thus, pre-neoplastic hepatocytes grow and progress to cancer upon transplantation into retrorsine treated rat liver, while the same cells are unable to expand when delivered to normal untreated host[51]. As discussed above for liver repopulation by transplanted normal hepatocytes, the biological coordinates of this phenomenon suggest that cell competition might be involved (Figure 2)[27], albeit formal proof of this linkage is not available yet.
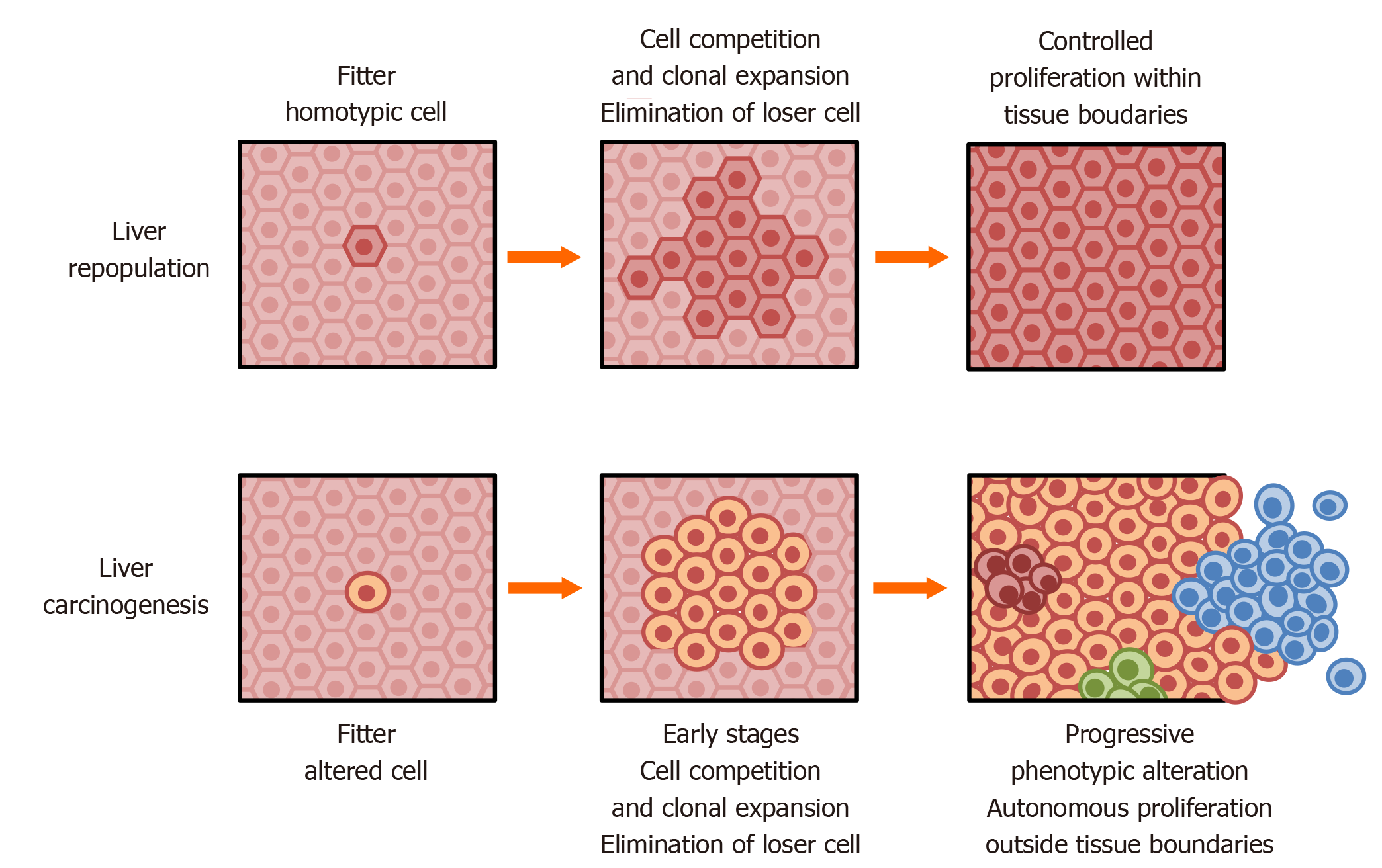
Figure 2 Liver repopulation and liver carcinogenesis.
Top: In the context of persistent, widespread tissue damage, transplanted homotypic cells with normal fitness (dark brown) can outcompete resident damaged cells (light brown) and slowly repopulate nearly the entire tissue, without exceeding tissue boundaries. However, in the absence of normal transplanted hepatocytes, endogenous cells persist for at least several months; Bottom: In a similar context of persistent, widespread tissue damage, transplanted homotypic cells with normal fitness and with a pre-neoplastic phenotype (dark yellow) can outcompete resident damaged cells and form hepatic nodules that progress to cancer. The same pre-neoplastic cells do not grow following transplantation in a context of normal cell fitness in vivo (see text for details).
Along these lines, an important step forward was the report by Moya et al[45], who have studied the role of Hippo signaling pathway in the growth of primary liver tumors and liver metastases from melanoma cells in mice. It was found that the relative level of activation of Hippo pathway in normal surrounding vs tumor cells was critical in determining the growth rate of the latter; specifically, inhibition of this pathway in peritumoral cells increased proliferation in nodular lesions, while tumors regressed when Hippo activity was up-regulated in surrounding normal tissue. In addition, tumor survival in wild type mice was dependent on the presence of an active Hippo pathway in cancer cells, while the activity of the pathway was dispensable when tumors were growing in a Hippo-deficient liver background[52]. These findings were interpreted to suggest that Hippo pathway-driven cell competition is an important determinant in controlling the growth of (pre)-neoplastic cell populations in the liver. It is intriguing to note that Hippo pathway activity was also shown to be essential for the maintenance of the differentiated state in hepatocytes and its inhibition correlated with the appearance of a progenitor cell phenotype[53].
CONCLUSION
Cell competition has emerged as an important quality control mechanism overlooking tissue functional proficiency during development and in post-natal life. In essence, such mechanisms entails the elimination and replacement of a less fit (loser) cell population by a fitter cell type (winner). Evidence is also accruing that this process might be involved in the pathogenesis of neoplastic disease at different steps. On one side of the coin is the first and possibly firmest conclusion so far, i.e., cell competition protects from the risk of cancer via identification and purging of single altered cells expressing a loser code, which flags a decreased fitness relative to surrounding counterparts. On the other hand, several studies suggest that the winner/loser code associated with cell competition can be exploited or hijacked by (pre)-cancerous cell populations to outcompete normal neighbors in the same tissue, thereby fueling their progression towards increasing malignancy. The prototypic example of such scenario is the super-competitor phenotype of Myc-overexpressing cells, which can be a winner phenotype over wild type cells. However, this apparently straightforward sequence of events appears difficult to reconcile with the defining biological features of cell competition, as it was aptly pointed out long ago[23]. A critical attribute of cell competition is in fact that it operates within discrete tissue boundaries that cannot be overridden by winner cells (see above). Thus, any cell type engaging in this process should stop expanding once the appropriate tissue compartment has been fully replenished, and this is also the case for the Myc super-competitor phenotype[1,21]. However, cancer cells do not obey these rules and their growth beyond set compartmental boundaries cannot be explained by cell competition per se (Figure 2). Stated otherwise, while canonical cell competition occurs within tissue homeostatic control mechanisms, the type of cell competition engaged by neoplastic cell populations is clearly outside such boundaries and is the expression of an eminently selfish phenotype[54,55].
A second feature of canonical cell competition which appears at odds with the phenotype of cancer cells is the type of cell interaction that is required to define relative fitness and hence the winner vs loser cell fate. According to the currently accepted paradigm, cell competition occurs among homotypic cells, in that it compares fitness levels within a cell population performing the same function in a tissue. By contrast, it is almost axiomatic that cancer cells have often departed significantly from the phenotype of the tissue of origin, including its functional proficiency. Therefore, a comparison of cell fitness between neoplastic cells and their normal tissue counterparts would likely favor survival of the latter, not the former.
Based on the above considerations, we propose that canonical cell competition is possibly involved in the initial stages of carcinogenesis leading, to discrete clonal expansions which are still within tissue homeostatic control[56]. However, the molecular machinery of cell competition may be co-opted by overtly neoplastic cell populations endowed with additional phenotypic features to sustain their uncontrolled growth.
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country/Territory of origin: Italy
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): 0
Grade C (Good): C, C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Hashimoto M, Li W S-Editor: Zhang L L-Editor: A P-Editor: Li JH