Copyright
©The Author(s) 2017.
World J Hepatol. Feb 8, 2017; 9(4): 191-208
Published online Feb 8, 2017. doi: 10.4254/wjh.v9.i4.191
Published online Feb 8, 2017. doi: 10.4254/wjh.v9.i4.191
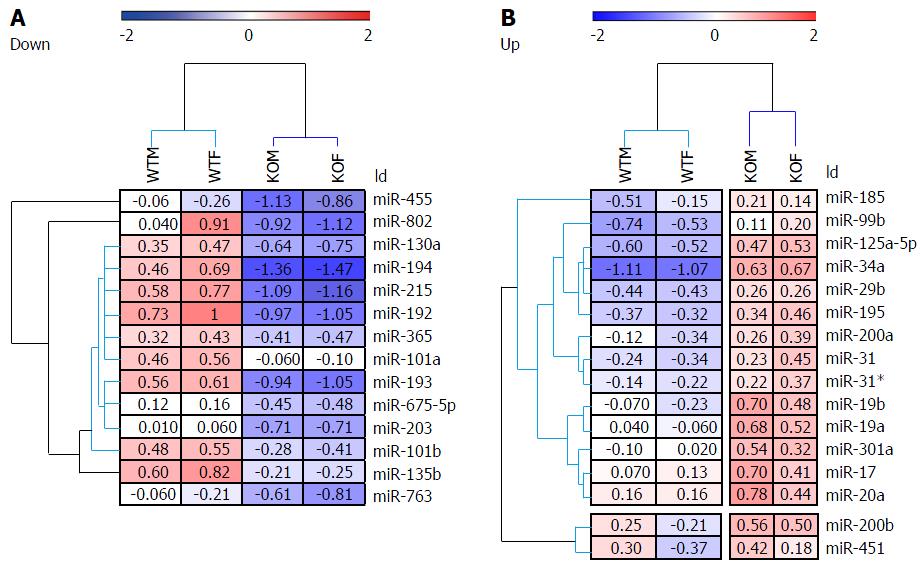
Figure 1 Heat map and unsupervised hierarchical clustering of hepatic microRNAs in male and female Hnf4a-LivKO mice.
The heat map diagram shows the results of the 2-way hierarchical clustering of microRNAs and samples. Each row represents a microRNA and each column represents a pooled liver sample. The microRNA clustering tree is shown on the left, and the sample clustering tree appears at the top. The color scale shown at the top illustrates the relative expression level of a microRNA across all samples: Red color represents an expression level above mean, blue color represents expression lower than the mean. The clustering is performed on log2(Hy3/Hy5) ratios which passed the filtering criteria on variation across samples; LogMedianDRatios differencies > 0.58, corresponding to 50% differential expression. WTM: Wild-type male; WTF: Wild-type female; KOM: Knockout male; KOF: Knockout female.
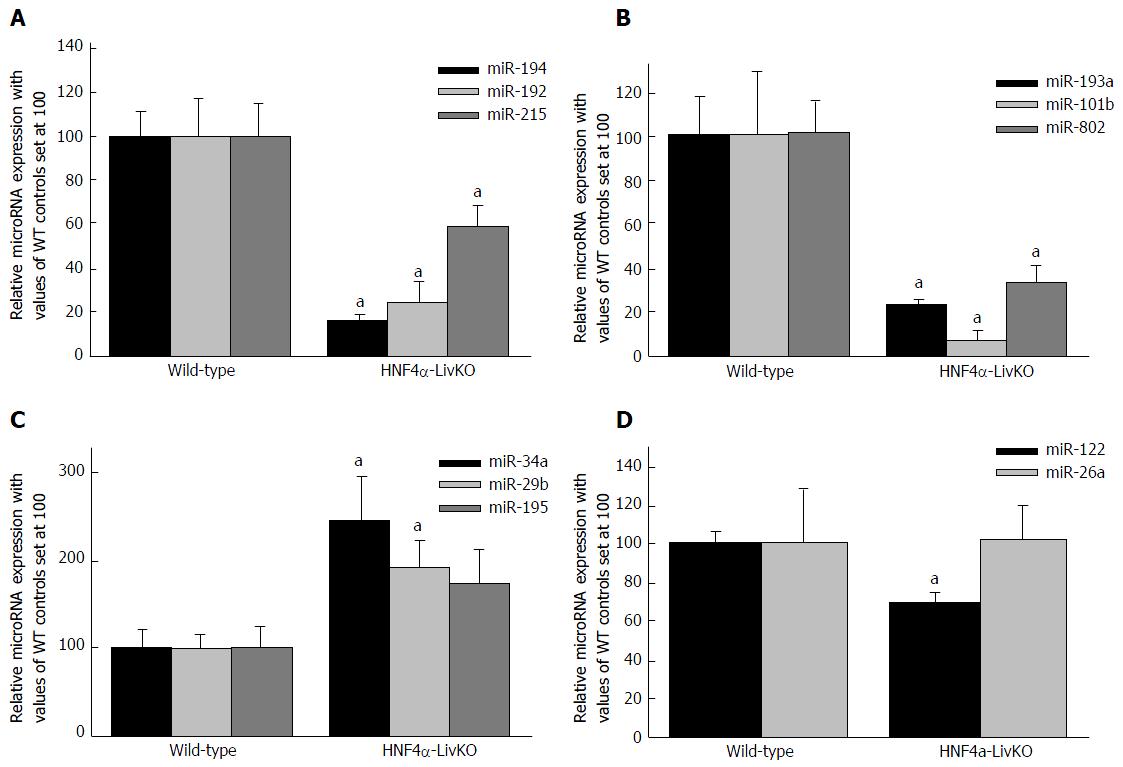
Figure 2 Hepatic microRNA expression in young-adult male mice with liver-specific deletion of Hnf4a (Hnf4a-LivKO) (A-D).
microRNAs in total RNA from livers of Hnf4a-LivKO and wild-type (WT) control mice (n = 5-6) were determined by miRCURY LNA™ Universal RT microRNA PCR (Exiqon). Mean ± SE. aP < 0.05 compared to WT control. HNF4α: Hepatocyte nuclear factor 4 alpha.
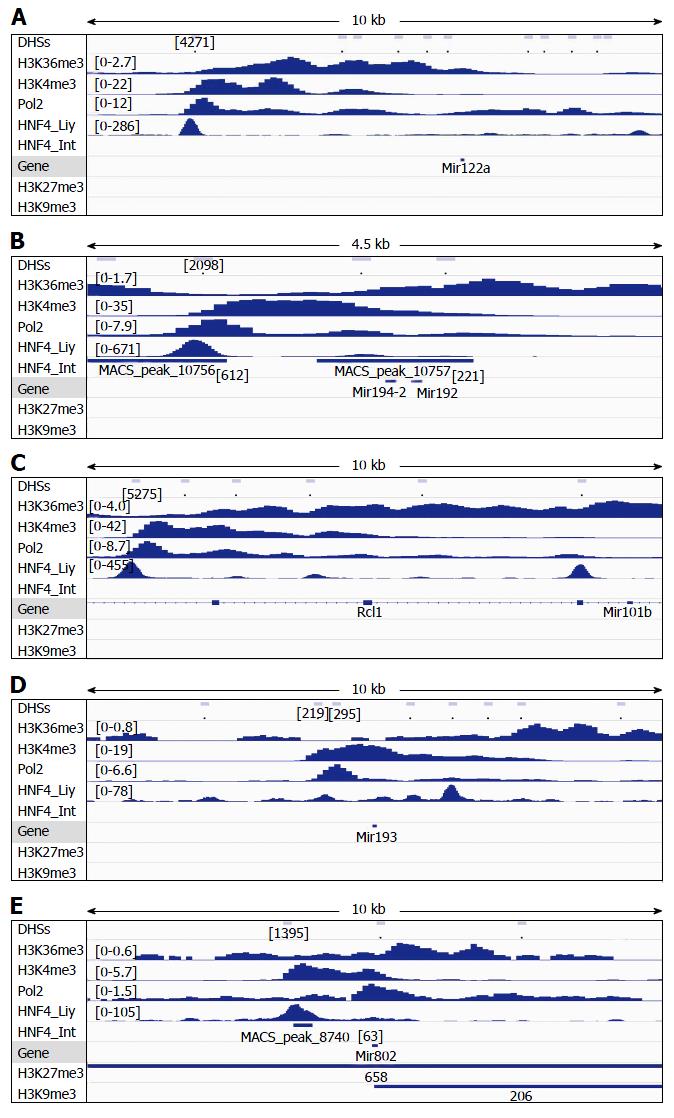
Figure 3 Analysis of DNAse-I hypersensitive sites as well as DNA-binding of HNF4α, RNA polymerase II (Pol2), and methylated histones to loci of miR-122a (A), miR-194-2/miR-192 (B), miR-101b (C), miR-193 (D) and miR-802 (E) in wildtype mouse liver.
DNA-binding of HNF4α to these microRNA loci in the mouse small intestine (HNF4α _Int) was compared to those in the mouse liver (HNF4α_Liv). Data of DHSs (determined by DNAse-seq) and DNA-binding of proteins (determined by ChIP-seq) were retrieved from the public database of GEO DataSets and visualized in the IGV software. The peak values/ranges for each mark were shown in square brackets or under the line mark. DHSs: DNAse-I hypersensitive sites; HNF4α: Hepatocyte nuclear factor 4 alpha; H3K36me3: H3 trimethylation at lysine-36; H3K4me3: H3 trimethylation at lysine-4; H3K27me3: H3 trimethylation at lysine-27; H3K9me3: H3 trimethylation at lysine-9; Pol2: Polymerase 2; HNF4α: Hepatocyte nuclear factor 4 alpha; ChIP-seq: Chromatin immunoprecipitation-sequencing; IGV: Integrative genomics viewer.
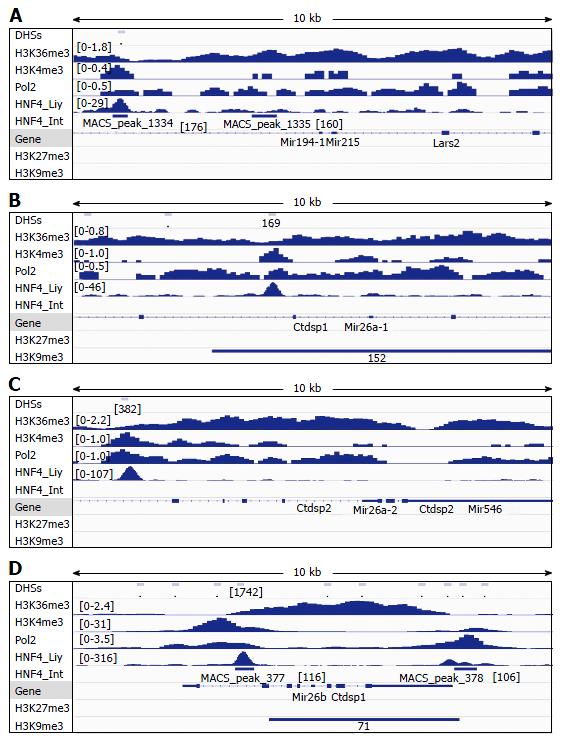
Figure 4 Analysis of DNAse-I hypersensitive sites as well as DNA-binding of HNF4α, RNA polymerase II (Pol2), and methylated histones to loci of miR-194-1/miR-215 (A), miR-26a-1 (B), miR-26a-2 (C) and miR-26b (D) in wildtype mouse liver.
DNA-binding of HNF4α to these microRNA loci in the mouse small intestine (HNF4α _Int) was compared to those in the mouse liver (HNF4α_Liv). Data of DHSs (determined by DNAse-seq) and DNA-binding of proteins (determined by ChIP-seq) were retrieved from the public database of GEO DataSets and visualized in the IGV software. The peak values/ranges for each mark were shown in square brackets or under the line mark. DHSs: DNAse-I hypersensitive sites; H3K36me3: H3 trimethylation at lysine-36; H3K4me3: H3 trimethylation at lysine-4; H3K27me3: H3 trimethylation at lysine-27; H3K9me3: H3 trimethylation at lysine-9; Pol2: Polymerase 2; HNF4α: Hepatocyte nuclear factor 4 alpha; ChIP-seq: Chromatin immunoprecipitation-sequencing; IGV: Integrative genomics viewer.
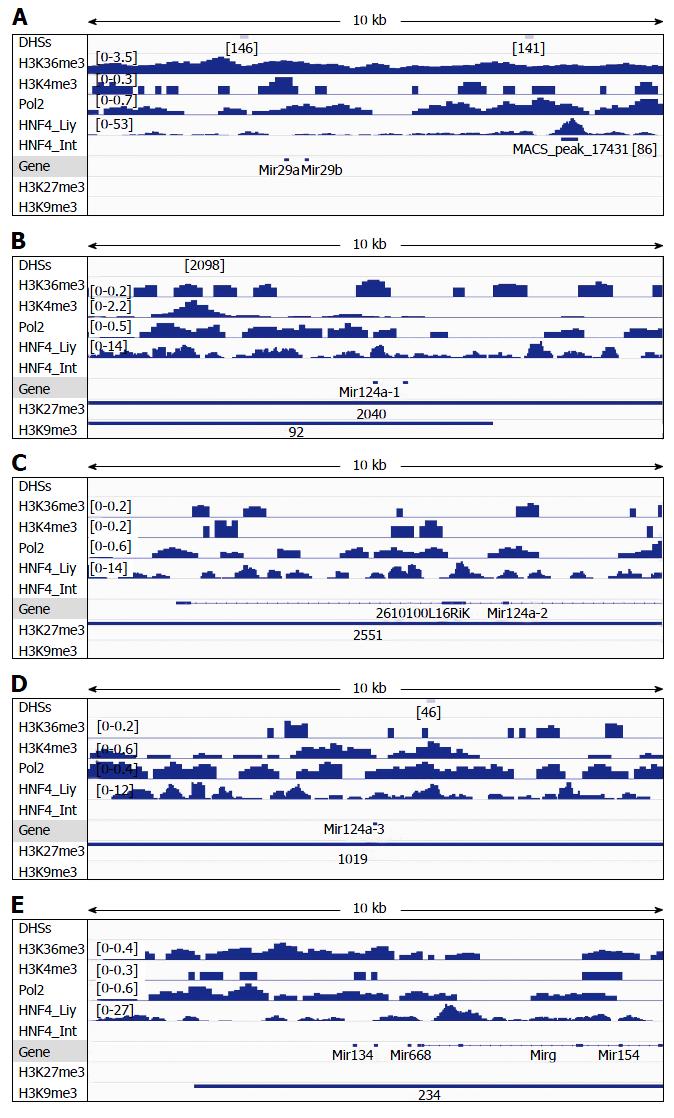
Figure 5 Analysis of DNAse-I hypersensitive sites as well as DNA-binding of HNF4α, RNA polymerase II (Pol2), and methylated histones to loci of miR-29a/miR-29b (A), miR-124a-1 (B), miR-124a-2 (C), miR-124a-3 (D) and miR-134 (E) in wildtype mouse liver.
DNA-binding of HNF4α to these microRNA loci in the mouse small intestine (HNF4α _Int) was compared to those in the mouse liver (HNF4α_Liv). Data of DHSs (determined by DNAse-seq) and DNA-binding of proteins (determined by ChIP-seq) were retrieved from the public database of GEO DataSets and visualized in the IGV software. The peak values/ranges for each mark were shown in square brackets or under the line mark. DHSs: DNAse-I hypersensitive sites; H3K36me3: H3 trimethylation at lysine-36; H3K4me3: H3 trimethylation at lysine-4; H3K27me3: H3 trimethylation at lysine-27; H3K9me3: H3 trimethylation at lysine-9; Pol2: Polymerase 2; HNF4α: Hepatocyte nuclear factor 4 alpha; ChIP-seq: Chromatin immunoprecipitation-sequencing; IGV: Integrative genomics viewer.
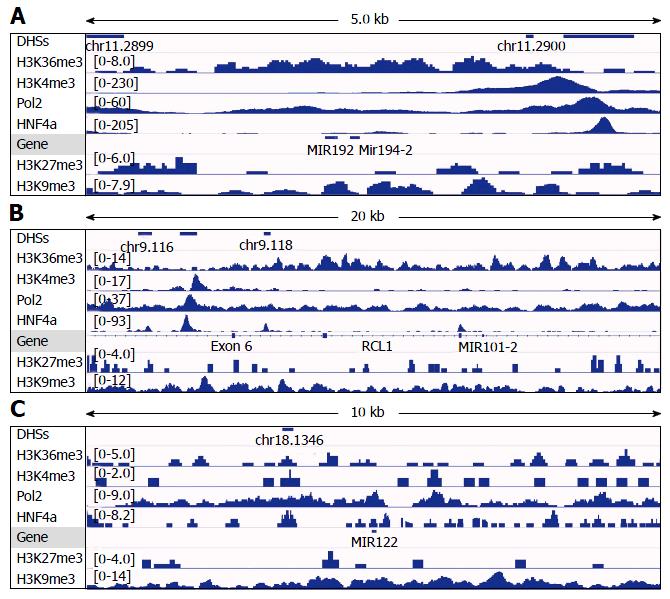
Figure 6 Analysis of DNAse-I hypersensitive sites as well as DNA-binding of HNF4α, RNA polymerase II (Pol2), and methylated histones to loci of miR-194-2/miR-192 (A), miR-101-2 (B) and miR-122 (C) in human hepatoma HepG2 cells.
Data of DHSs (determined by DNAse-seq) and DNA-binding of proteins (determined by ChIP-seq) were retrieved from the public database of GEO DataSets and visualized in the IGV software. The peak values/ranges for each mark were shown in square brackets or under the line mark. DHSs: DNAse-I hypersensitive sites; H3K36me3: H3 trimethylation at lysine-36; H3K4me3: H3 trimethylation at lysine-4; H3K27me3: H3 trimethylation at lysine-27; H3K9me3: H3 trimethylation at lysine-9; Pol2: Polymerase 2; HNF4α: Hepatocyte nuclear factor 4 alpha; ChIP-seq: Chromatin immunoprecipitation-sequencing; IGV: Integrative genomics viewer.
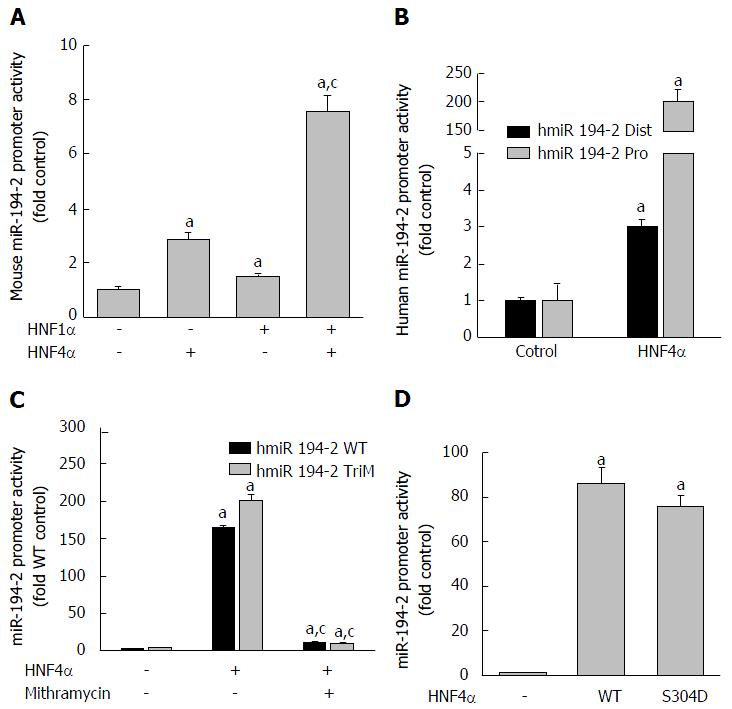
Figure 7 Activation of mouse (A) and human (B-D) miR-194-2/miR-192 promoter by HNF4α.
Human hepatoma HepG2 cells were transfected with firefly luciferase vectors containing wild-type and mutant miR-194-2 promoter, pRL-CMV, and an expression vector for HNF4α/HNF1α. Dual-luciferase reporter assay was conducted 24 h after transfection. The y-axis represents relative luciferase activity for microRNA promoter normalized by the renilla luciferase. n = 4, Mean ± SE. aP < 0.05 compared to vector control; cP < 0.05 compared to HNF4α alone group. HNF4α: Hepatocyte nuclear factor 4 alpha.
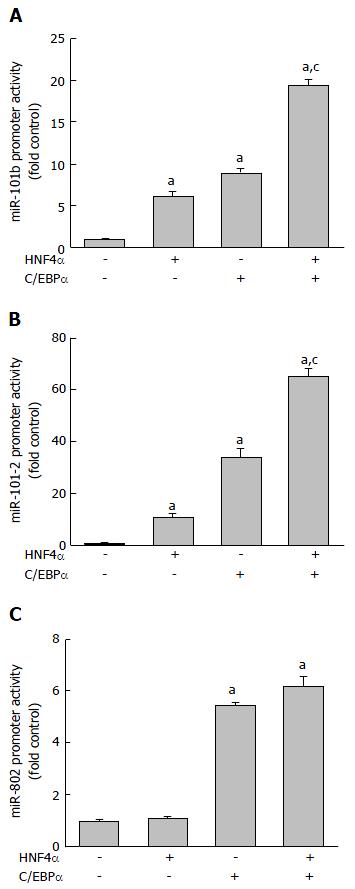
Figure 8 Activation of (A) mouse miR-101b, (B) human miR-101-2, and (C) mouse miR-802 promoter by HNF4α.
Human hepatoma HepG2 cells were transfected with firefly luciferase vectors containing microRNA promoter, pRL-CMV, and an expression vector for HNF4α and/or C/EBPα. Dual-luciferase reporter assay was conducted 24 h after transfection. The Y-axis represents relative luciferase activity for microRNA promoter normalized by the renilla luciferase. n = 4, Mean ± SE. aP < 0.05 compared to vector control; cP < 0.05 compared to HNF4α alone group. HNF4α: Hepatocyte nuclear factor 4 alpha; C/EBPα: CCAAT/enhancer-binding protein α.
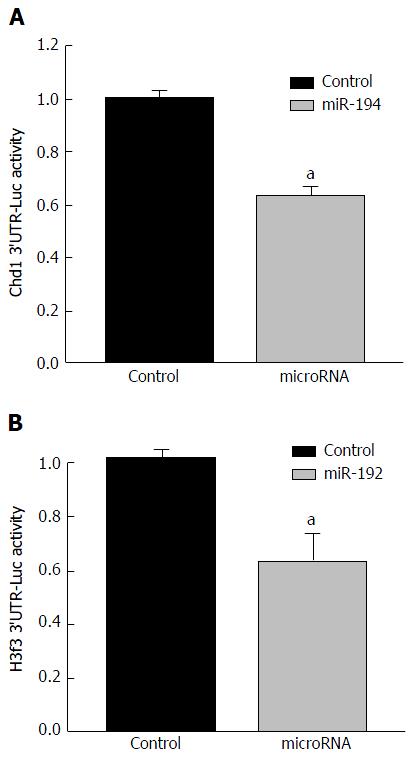
Figure 9 Effects of miR-194 and miR-192 on the activities of luciferase reporter vectors for the 3’UTR of mouse Chd1 and H3f3.
Human hepatoma HepG2 cells were transfected with plasmid DNA including pmiR-Chd1 (or pmiR-H3f3), the pRL-CMV luciferase, and a synthetic mimic of miR-194/miR-192, or AllStars Negative Control siRNA (as negative control for microRNAs) using Lipofectamine 2000. Dual-luciferase reporter assay was conducted 24 h after transfection. The Y-axis represents relative luciferase activity for the 3’UTR of Chd1 or H3f3 normalized by the renilla luciferase. n = 4, Mean ± SE. aP < 0.05 compared to control (AllStars Negative Control siRNA). 3’UTR: Untranslated regions.
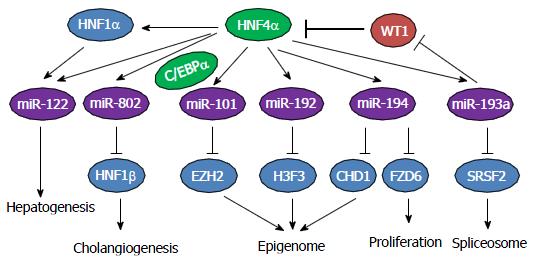
Figure 10 Diagram that illustrates the regulation of hepatic microRNA expression by Hnf4α in mouse liver.
HNF4α: Hepatocyte nuclear factor 4 alpha; C/EBPα: CCAAT/enhancer-binding protein α.
- Citation: Lu H, Lei X, Liu J, Klaassen C. Regulation of hepatic microRNA expression by hepatocyte nuclear factor 4 alpha. World J Hepatol 2017; 9(4): 191-208
- URL: https://www.wjgnet.com/1948-5182/full/v9/i4/191.htm
- DOI: https://dx.doi.org/10.4254/wjh.v9.i4.191