Published online Mar 28, 2025. doi: 10.3748/wjg.v31.i12.103991
Revised: January 26, 2025
Accepted: February 24, 2025
Published online: March 28, 2025
Processing time: 110 Days and 15.7 Hours
Keratin 80 (KRT80), a type I intermediate filament protein, is a member of the keratin family with specialized functions in epithelial tissues. While KRT80 has been implicated in both normal physiological processes and various diseases, its role in gastric cancer (GC), particularly its expression and prognostic significance, remains poorly understood. In this study, we investigated the role and underlying molecular mechanisms of KRT80 in oxaliplatin resistance in GC. Our analysis revealed that KRT80 is significantly upregulated in GC tissues and is associated with poor clinical prognosis. The role of KRT80 in GC cell proliferation was assessed through in vitro and in vivo assays.
To explore the expression of KRT80 in GC and its impact on the prognosis of pa
KRT80 expression in GC tissues was analyzed using Western blotting, quan
Integrating our experimental findings with multiple published studies, we found that increased KRT80 expression is associated with poor prognosis in GC and promotes resistance to oxaliplatin. Moreover, we have preliminarily verified the interaction between KRT80 and EEF1A1. Therefore, this study provides a novel perspective on overcoming oxaliplatin resistance in GC.
Increased KRT80 expression predicts poor prognosis and promotes oxaliplatin resistance in GC, suggesting its potential as a novel prognostic biomarker.
Core Tip: In this paper, keratin 80 (KRT80) was found to be overexpressed in gastric cancer (GC) tissues, functioning as an oncogene and correlating with poor patient prognosis. KRT80 promoted GC proliferation, migration, invasion abilities and oxaliplatin resistance by activating the NF-κB signaling pathway. Immunoprecipitation and mass spectrometry analyses identified elongation factor 1-alpha 1 (EEF1A1) as a binding protein of KRT80, enhancing its protein stability. Notably, knockdown of EEF1A1 partially reversed the effects of KRT80 overexpression in GC cells.
- Citation: Yang B, Cao HX, He YW, Ouyang JJ, Lv M, Li YX, Lu YD. Increased keratin 80 expression predicts poor prognosis and promotes oxaliplatin resistance in gastric cancer. World J Gastroenterol 2025; 31(12): 103991
- URL: https://www.wjgnet.com/1007-9327/full/v31/i12/103991.htm
- DOI: https://dx.doi.org/10.3748/wjg.v31.i12.103991
Gastric cancer (GC) is one of the most prevalent gastrointestinal malignancies worldwide. Despite significant research advancements, the five-year survival rate for GC remains low, largely due to metastasis, recurrence, and drug resistance[1]. Current clinical treatments primarily consist of surgical resection and chemotherapy, with platinum-based agents such as oxaliplatin and cisplatin serving as the cornerstone of first-line chemotherapy regimens[2]. The development of treatments oncogenic gene alterations and related signaling pathways, driven by advances in understanding of molecular cancer biology, was a major step in cancer treatment evolution[3]. Platinum-based chemotherapy remains a key treatment for advanced GC, but its effectiveness is limited by many factors. Additionally, the treatment induces severe side effects, including neurotoxicity (peripheral neuropathy), myelosuppression (anemia, neutropenia), and renal toxicity, which often limit its clinical application.
However, resistance to these drugs, whether primary or acquired, pose significant clinical challenges. Prolonged exposure of tumor cells to anticancer agents often leads to reduced drug sensitivity, diminishing therapeutic efficacy[4,5]. Previous studies have implicated various genes and proteins, including B-cell lymphoma-2 (Bcl-2), Caspase-3, Bcl-2-associated X protein (Bax), survivin, and the inhibitor of apoptosis protein family, in the resistance mechanisms of resistance to platinum-based drugs[6-9].
The NF-κB signaling pathway plays a crucial role in regulating various cellular functions and serves as a key mediator of tumor drug resistance[10,11]. This pathway frequently interacts with others, such as the PI3K-AKT pathway, forming a complex regulatory network. During chemotherapy, activation of the PI3K-AKT pathway can enhance IKK activation, amplifying NF-κB activity. This amplification influences tumor cell behaviors, including growth, apoptosis resistance, and migration[12-14]. Numerous studies have demonstrated that chemotherapy-induced activation of NF-κB signaling enhances drug resistance in tumor cells[15-17].
As a member of the keratin family, keratin 80 (KRT80) plays a distinct role in epithelial tissues, contributing not only to normal physiological processes but also being intricately closely linked to the onset of various diseases[18]. Aberrant KRT80 expression has been observed in multiple cancers, including breast[19], colon[20], and lung cancers[21], where its overexpression is associated with increased tumor cell proliferation, invasion, and metastasis. Additionally, KRT80 holds potential as a therapeutic target for hair follicle-related disorders[22]. It is important to highlight that while the keratin family has been recognized for its significant role in tumor chemoresistance, the specific involvement of the involvement of KRT80 in this process remains unexplored.
In our study, we identified elevated KRT80 expression in GC, which correlated with a malignant phenotype and poor prognosis. Elevated KRT80 expression Increased KRT80 Levels enhanced oxaliplatin-induced apoptosis in GC cells both in vitro and in vivo. Mechanistically, KRT80 interacted with elongation factor 1-alpha 1 (EEF1A1), protecting GC cells from oxaliplatin-induced cytotoxicity by activating the NF-κB signaling pathway.
This study included a total of 118 formalin-fixed paraffin-embedded GC tissue samples and 18 randomly selected adjacent normal tissue specimens, all obtained from patients who underwent radical gastrectomy without prior chemotherapy or radiotherapy. The samples were collected from the First Affiliated Hospital of Anhui Medical University between January 2017 and December 2018 for tissue microarray (TMA) construction. The follow-up period for these patients ranged from 6 to 65 months. Clinical and pathological data are presented in Table 1, with GC stages assigned according to the 8th edition of the American Joint Committee on Cancer staging system. Additionally, 6 fresh primary cancer samples and corresponding adjacent normal tissue samples were collected for histamine extraction. The study was approved by the Ethics Committee of Anhui Medical University. To be eligible for inclusion, patients had to: (1) Adult males or females (individuals ≥ 18 years of age); (2) Have a preoperative gastroscopy and biopsy-confirmed diagnosis of gastric adenocarcinoma; (3) Have underwent a thorough preoperative cardiopulmonary function evaluation, which revealed no significant contraindications to surgery or anesthesia; (4) Have undergone open, laparoscopic or robotic gastrectomy in Department of Gastrointestinal Surgery of the First Affiliated Hospital of Anhui Medical University; (5) Did not undergo neoadjuvant chemotherapy or immunotherapy before surgical procedure; and (6) Provide written informed consent before surgery, and are explicitly informed that their surgical specimens would be utilized for scientific research purposes[23-25].
| Parameters | Cases | KRT80 | χ² | P value | |
| High expression | Low expression | ||||
| Gender | |||||
| Male | 71 | 41 (59.4) | 30 (61.2) | 0.039 | 0.844 |
| Female | 47 | 28 (40.6) | 19 (38.8) | ||
| Age (years) | |||||
| < 61 | 55 | 31 (44.9) | 24 (49) | 0.189 | 0.664 |
| ≥ 61 | 63 | 38 (55.1) | 25 (51) | ||
| Tumor location | |||||
| Upper | 64 | 40 (58) | 24 (49) | 0.933 | 0.334 |
| Middle + lower | 54 | 29 (42) | 25 (51) | ||
| Tumor size (cm) | |||||
| < 6 | 63 | 36 (52.2) | 27 (55.1) | 0.099 | 0.753 |
| ≥ 6 | 55 | 33 (47.8) | 22 (44.9) | ||
| Tumor differentiation | |||||
| Well + moderate | 47 | 27 (39.1) | 20 (40.8) | 0.034 | 0.854 |
| Poor + differentiated | 71 | 42 (60.9) | 29 (59.2) | ||
| Lymph nodes metastasis | 58 (84.1) | 38 (77.6) | 0.800 | 0.371 | |
| pTNM stage | |||||
| I + II | 30 | 10 (14.5) | 20 (40.8) | 10.472 | 0.001 |
| III + IV | 88 | 59 (85.5) | 29 (59.2) | ||
Normal gastric epithelial cell line GES-1, along with several GC cell lines (AGS, MGC803, SGC7901, HGC27, and BGC823), were procured from the Chinese Academy of Sciences (Shanghai, China). These cells were cultured in RPMI 1640 medium (R10-040-CV, Corning, United States), supplemented with 10% fetal bovine serum (FBS, Clark, United States) and 1% penicillin-streptomycin (SV30010, HyClone, United States)[26].
Lentiviruses overexpressing KRT80, KRT80-specific small hairpin RNA, and a negative control lentivirus were generated by Jiman Technology (Shanghai, China). Small interfering RNA (siRNA) targeting EEF1A1, along with a control siRNA, were obtained from Tsingke Biotechnology (Beijing, China). Stable GC cell lines with KRT80 knockdown or overexpression were established by transfection, followed by selection with 2 μg/mL puromycin (ST551, Sangon Biotech, China) for 6 days.
Total RNA of cells and tissues were extracted by using TRIzol reagent (Invitrogen, United States), and then used the Hifair® Ⅲ 1st Strand cDNA Synthesis SuperMix with gDNA Remover (11141ES, Yeasen, China) according to the protocol. Quantitative real-time PCR (qRT-PCR) was performed with the Hifair® III One Step RT-qPCR Probe Kit (11145ES, Yeasen, China) on the QuantStudio 5 qPCR Platform (Thermo Fisher, United States). Gene expression was analyzed using the 2-ΔΔCt method, with GAPDH as an internal control.
Proteins from cells or tissues were extracted using lysis buffer (78501, Thermo Fisher Scientific, United States), and the protein concentration was determined using a BCA protein assay kit (P0012S, Beyotime, China). Equal amounts of protein were separated on a 12% SDS-PAGE gel and transferred to a PVDF membrane. The membrane was blocked with 5% non-fat milk (A600669, Sangon Biotech, China) and incubated overnight at 4 °C with primary antibodies. After a one-hour incubation with secondary antibodies, protein bands were detected using enhanced chemiluminescence (Bridgen, Beijing, China) and visualized with a Tanon 5200 Multi chemiluminescence detection system (Tanon Science, Shanghai, China). The protein expression levels were quantitatively analyzed using ImageJ software 7.0 (National Institutes of Health, United States). The relative expression levels of the target proteins were normalized to those of the housekeeping protein, and the data were presented as fold changes relative to the control group.
TMA sections were incubated with anti-KRT80 antibody and subjected to immunohistochemical staining. The staining was scored based on the extent (0, no staining; 1, 0%-25%; 2, 26%-50%; 3, 51%-75%; 4, 76%-100%) and intensity (0, negative; 1, weak; 2, moderate; 3, strong). Immunohistochemical scores were evaluated by two pathologists, with a score of 5 or higher considered indicative of high expression and a score of 0-4 representing low expression.
GC cells were plated in 12-well plates at a density of 3000 cells per well and cultured at 37 °C for 7 days, with medium changes every 48 hours. On each day after plating, one set of cells was digested and counted, generating a cell growth curve based on the recorded cell numbers[27].
GC cells were seeded into the upper chamber of a 24-well Transwell plate at a density of 5 × 105 cells. For the invasion assay, the upper chamber was coated with Matrigel (BD Biosciences, United States), while no Matrigel was applied for the migration assay. The upper chamber was filled with RPMI-1640 containing 1%-2% FBS, while the lower chamber was filled with RPMI-1640 supplemented with 20% FBS. After 24-48 hours, cells were fixed in 4% paraformaldehyde (PFA) and stained with 0.1% crystal violet. Images were taken using a Leica DMi1 microscope (Leica, Germany), and cell migration or invasion was quantified based on the number of stained cells[28,29].
Cells were cultured on plates with pre-positioned cover glasses (Nest, China) and treated with varying concentrations of oxaliplatin. After 48 hours, cells attached to the cover glasses were fixed with 4% PFA for 15 minutes at room temperature, followed by three washes with PBS. The cells were then permeabilized with 0.1% PBS-TX (0.1% Triton X-100 in PBS) for 5 minutes and washed three times with PBS. Blocking was performed using 2% BSA (bovine serum albumin) diluted in 0.1% PBS-TX for 30 minutes. Next, 80 μL of a 1X Phalloidin-iFluor™ 594 Conjugate (AAT Bioquest, United States) working solution was added and incubated for 1 hour. Cells were washed 3-5 times with PBS to remove any excess solution. Subsequently, nuclei were stained with Antifade Mounting Medium containing DAPI (P0131, Beyotime Institute of Biotechnology) for 15 minutes at room temperature. Images were captured using a Live Cell Imaging System (Cell Discoverer 7, Carl Zeiss, Germany).
Transfected cells were seeded in 6-well plates and cultured until reaching 90% confluence. A linear wound was created in each well by scratching the confluent cell monolayer using a 200 μL pipette tip. The cells were washed twice with PBS to remove any floating cells and debris. The wounds were photographed and measured at 0, 24, and 48 hours using the Live Cell Imaging System (Cell Discoverer 7, Carl Zeiss, Germany).
GC cells were lysed in ice-cold lysis buffer supplemented with protease and phosphatase inhibitor cocktails. Following preclearing with 20 μL of protein A/G plus-agarose (sc-2003; Santa Cruz Biotechnology, United States) at 4 °C for 30 minutes, the lysates were incubated overnight at 4 °C with anti-KRT80 antibody or IgG, on a gyratory shaker. The mixture was then incubated with 20 μL of protein A/G plus-agarose at 4 °C for 6 hours. Immunoprecipitated beads were washed three times with ice-cold lysis buffer, then boiled in 50 μL of 1X SDS sample buffer for 8 minutes at 95 °C. The immunoprecipitated protein complexes were analyzed by Western blotting. To minimize interference from the heavy or light chains, a specific secondary antibody against the mouse anti-rabbit IgG light chain (M21008, Abmart, China) was used[30].
Cell proliferation was assessed using the BeyoClick™ EdU Cell Proliferation Kit (C0078S, Beyotime, China). Approximately 5 × 104 cells were seeded into 12-well plates and cultured overnight at 37 °C. After 48 hours of incubation with DMSO or various reagents, 20 μM EdU was added to the culture medium, and cells were incubated for 120 minutes. Cells were then fixed with 4% PFA for 20 minutes, rinsed three times with 3% BSA, and permeabilized for 15 minutes using 0.3% Triton X-100 in PBS. Cells were incubated with BeyoClick™ Click Additive Solution at room temperature, protected from light, for 30 minutes. Nuclei were stained with Hoechst 33342 for 15 minutes at room temperature. Cells were imaged using the Live Cell Imaging System (Cell Discoverer 7, Carl Zeiss, Germany), and the number of EdU-positive cells in each field was quantified.
KRT80 protein stability was evaluated using a cycloheximide (CHX) chase assay. Cells were treated with 20 μg/mL CHX (S7418, Selleck Chemicals, United States) for various time points (0, 2, 4, 6, and 8 hours). Protein samples were collected and analyzed by Western blotting, with GAPDH serving as a loading control.
Animal experiments were approved by the Ethics Committee for Animal Studies of Anhui Medical University. Four- to six-week-old male BALB/c nude mice, purchased from GemPharmatech (Jiangsu, China), were fed in laminar flow cabinets under specific pathogen-free (SPF) conditions. The mice were randomly divided into several groups. Approximately 3 × 106 KRT80 overexpress or control GC cells were subcutaneously injected into the right axilla of the nude mice. Starting on day 7, mice in the oxaliplatin treatment group received 8 mg/kg oxaliplatin by intraperitoneal injection every 5 days for 20 days. Tumor volume was measured starting from day 10 after injection, and measurements were taken every 3 days in three dimensions using a digital caliper. Tumor volume was calculated using the formula: Tumor volume (mm³) = π/6 × (W)² × (L), where L is the long diameter and W is the short diameter. After 28 days, the mice were euthanized and tumors were excised and weighed[31-33].
The survival curve of GC patients with high and low KRT80 expression was generated using Kaplan-Meier Plotter (https://www.kmplot.com).
All animal experiments adhered to the guidelines set forth by the Animal Center of Anhui Medical University. Experimental protocols were approved by the Experimental Animal Ethical Committee of Anhui Medical University. All animals were humanely euthanized at the conclusion of the study.
Statistical analyses were performed using SPSS 22.0 (SPSS Inc., United States) and GraphPad Prism 9.0 (GraphPad Software Inc., United States). Data are presented as mean ± SD and were compared using Student’s t-test or one-way ANOVA. The relationship between KRT80 expression and pathological variables was evaluated using Pearson’s χ2 test. Survival analysis was conducted using the Cox proportional hazards regression model and Kaplan-Meier method with log-rank testing. P values < 0.05 were considered statistically significant (aP < 0.05; bP < 0.01; cP < 0.001).
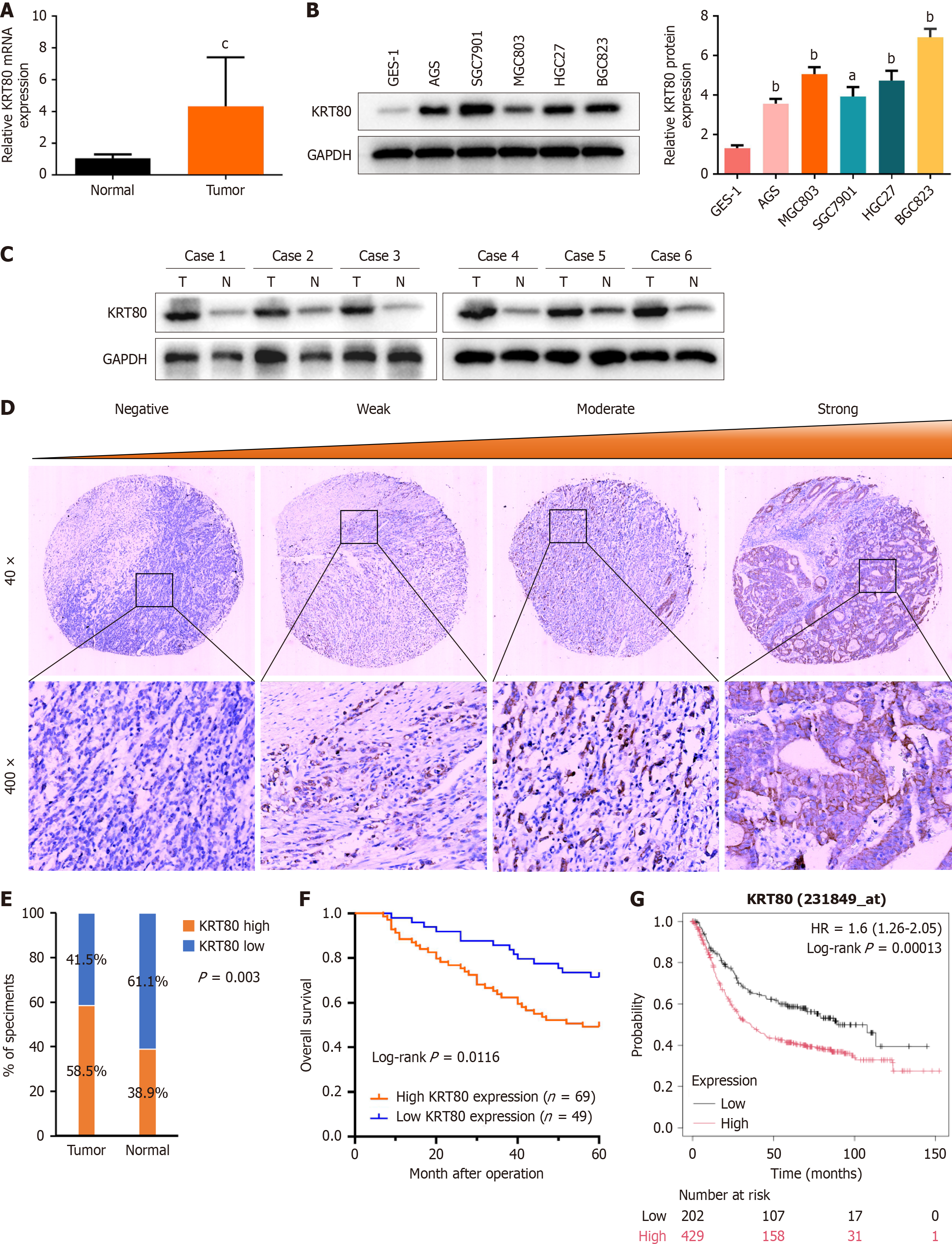
To investigate the expression pattern of KRT80 in GC, we performed qRT-PCR on 20 paired samples of GC and adjacent normal tissues. The results revealed that KRT80 mRNA expression was significantly elevated in GC tissues compared to normal tissues (Figure 1A). Similarly, KRT80 expression was higher in GC cell lines (AGS, MGC803, SGC7901, HGC27, and BGC823) than in the nonmalignant GES-1 cell line (Figure 1B). To further validate these findings, we analyzed KRT80 protein expression in six paired samples of fresh GC and adjacent normal tissues, confirming increased KRT80 expression in GC tissues (Figure 1C).
Immunohistochemistry was conducted on a TMA containing 118 GC tissues and 18 adjacent normal tissues. Representative images illustrating varying KRT80 expression levels are shown in Figure 1D. High KRT80 protein expression was detected in 58.5% (69/118) of GC tissues, while 61.1% (11/18) of adjacent normal tissues exhibited low KRT80 expression (Figure 1E). To assess the clinical relevance of KRT80, its correlation with clinicopathological parameters was analyzed. As shown in Table 1, increased KRT80 expression was significantly associated with distant metastasis (P = 0.001). Additionally, overall survival (OS) analysis indicated that GC patients with elevated KRT80 expression had shorter survival times than those with lower expression (log-rank P = 0.0116; Figure 1F). This association was further validated in a larger cohort using the Kaplan-Meier plotter database (log-rank P = 0.00013; Figure 1G).
Cox regression analysis, including univariate and multivariate models, was conducted to evaluate prognostic factors such as sex, age, tumor location, tumor size, depth of invasion, lymph node metastasis, differentiation, tumor-node-metastasis stage, and KRT80 expression. Univariate analysis identified lymph node metastasis (P = 0.025) and KRT80 expression (P = 0.014) as significantly associated with OS in GC patients (Table 2). Multivariate analysis confirmed that both lymph node metastasis (P = 0.03) and KRT80 expression (P = 0.018) were independent prognostic factors for OS (Table 2). Collectively, these findings indicate that KRT80 is upregulated in GC and serves as an independent prognostic biomarker for OS.
| Parameters | Univariate analysis | Multivariate analysis | ||
| HR (95%CI) | P value | HR (95%CI) | P value | |
| Gender (male vs female) | 1.230 (0.682-2.216) | 0.491 | ||
| Age (< 60 years vs ≥ 60 years) | 1.475 (0.834-2.609) | 0.181 | ||
| Tumor location (upper vs middle + lower) | 0.827 (0.468-1.462) | 0.514 | ||
| Tumor size (< 6 cm vs ≥ 6 cm) | 0.879 (0.501-1.544) | 0.653 | ||
| Differentiation (well + moderate vs poor + undifferentiated) | 0.758 (0.432-1.330) | 0.335 | ||
| pT stage (T1-T2 vs T3-T4 ) | 1.047 (0.597-1.836) | 0.872 | ||
| Lymph node metastasis (absent vs present) | 3.212 (1.155-8.938) | 0.025 | 3.110 (1.117-8.657) | 0.03 |
| pTNM stage (I-II vs III-IV) | 1.409 (0.703-2.842) | 0.333 | ||
| KRT80 expression (low vs high) | 2.174 (1.169-4.044) | 0.014 | 2.119 (1.139-3.943) | 0.018 |
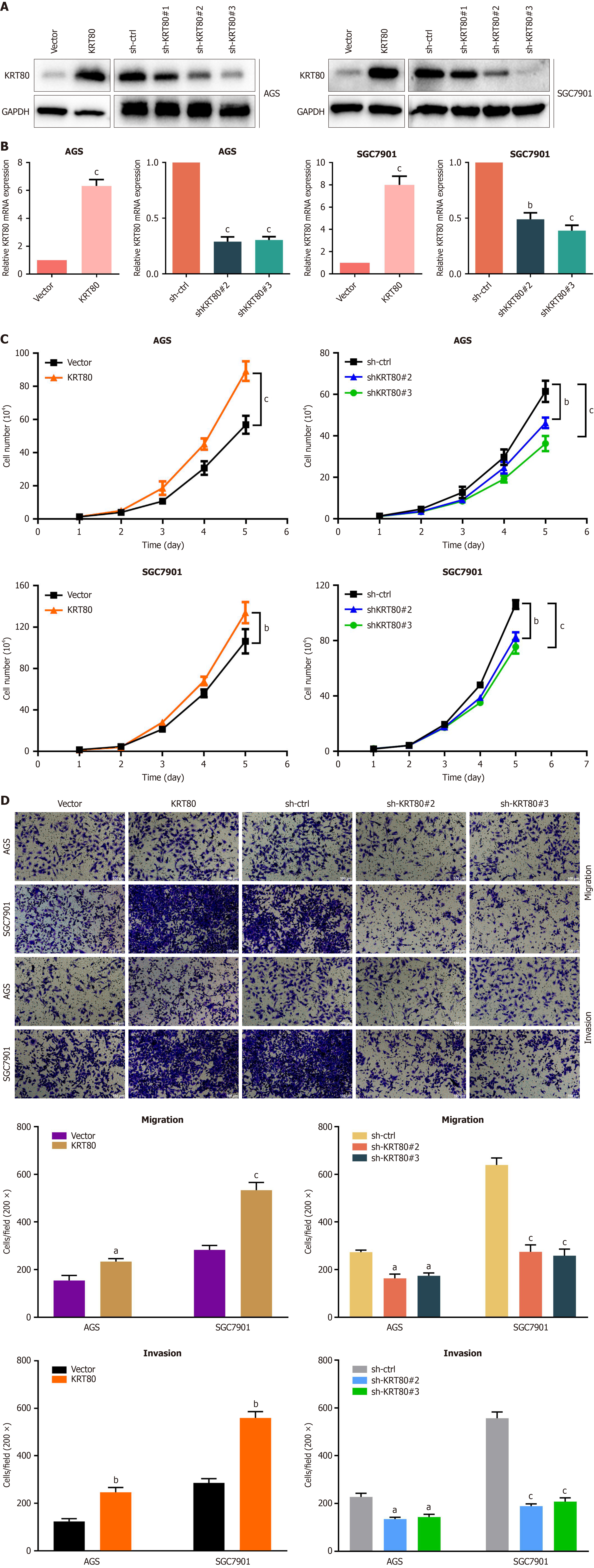
To elucidate the role of KRT80 in GC, a series of functional assays were performed. Based on KRT80 expression levels in GC cell lines, the AGS and SGC7901 cell lines were selected for KRT80 overexpression and knockdown experiments. The efficiency of these manipulations was confirmed using qRT-PCR and Western blot analysis (Figure 2A and B). Cell growth curve analysis demonstrated that KRT80 overexpression significantly enhanced cell proliferative capacity, while KRT80 knockdown reduced proliferation (Figure 2C). Furthermore, as shown in Figure 2D, KRT80 overexpression markedly increased the migration and invasion abilities of GC cells, whereas KRT80 knockdown diminished these capacities. Collectively, these findings indicate that KRT80 plays a pivotal role in regulating GC cell malignant phenotypes in vitro.
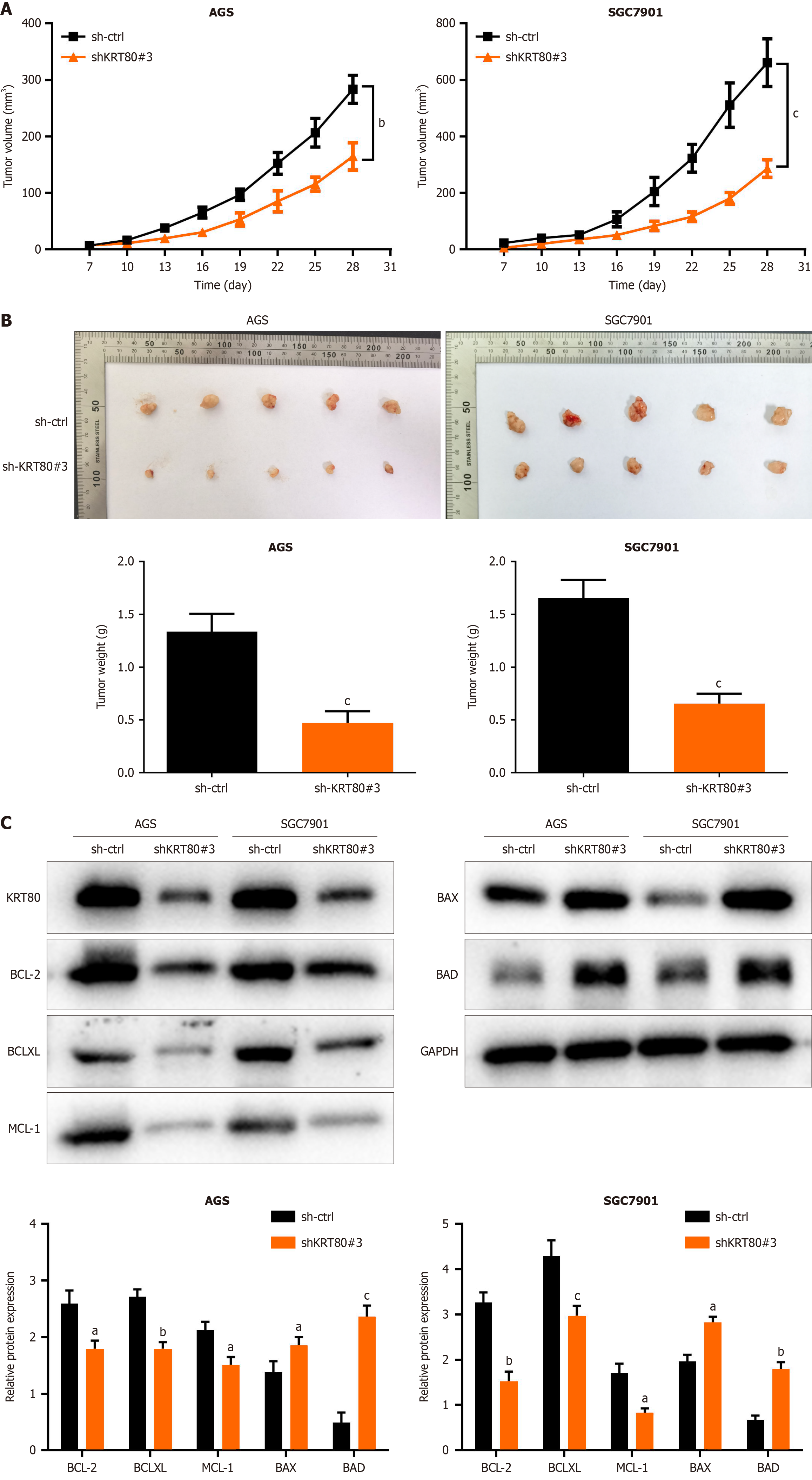
To further investigate the impact of KRT80 on GC tumor growth in vivo, a subcutaneous xenograft model was established in nude mice. Four-week-old male Balb/c nude mice were acclimatized in SPF conditions for one week. Subsequently, KRT80 knockdown cells and their corresponding control cells (AGS and SGC7901) were injected subcutaneously into the posterior axillary region of the right flank of the mice. Tumor volume measurements began on day 10 post-injection and were conducted every three days (Figure 3A). On day 18, the mice were euthanized, and the tumors were excised for volume and weight measurement. The results showed a significant reduction in tumor growth rate, volume, and mass in the KRT80 knockdown group (P < 0.01; Figure 3B).
Additionally, proteins were extracted from the subcutaneous tumor tissues, and the expression of apoptosis-related proteins was evaluated. These included anti-apoptotic proteins (BCL-2, BCL-XL, and MCL-1) and pro-apoptotic proteins (BAX and BAD). KRT80 knockdown significantly downregulated the expression of anti-apoptotic proteins while upregulating pro-apoptotic proteins (Figure 3C). These findings indicate that KRT80 depletion may promote apoptosis in GC cells.
Previous studies suggest that KRT80 expression may influence the efficacy of chemotherapy in cancer patients. Given that oxaliplatin is a widely used first-line chemotherapeutic agent for GC, this part aimed to investigate whether KRT80 expression affects its therapeutic efficacy.
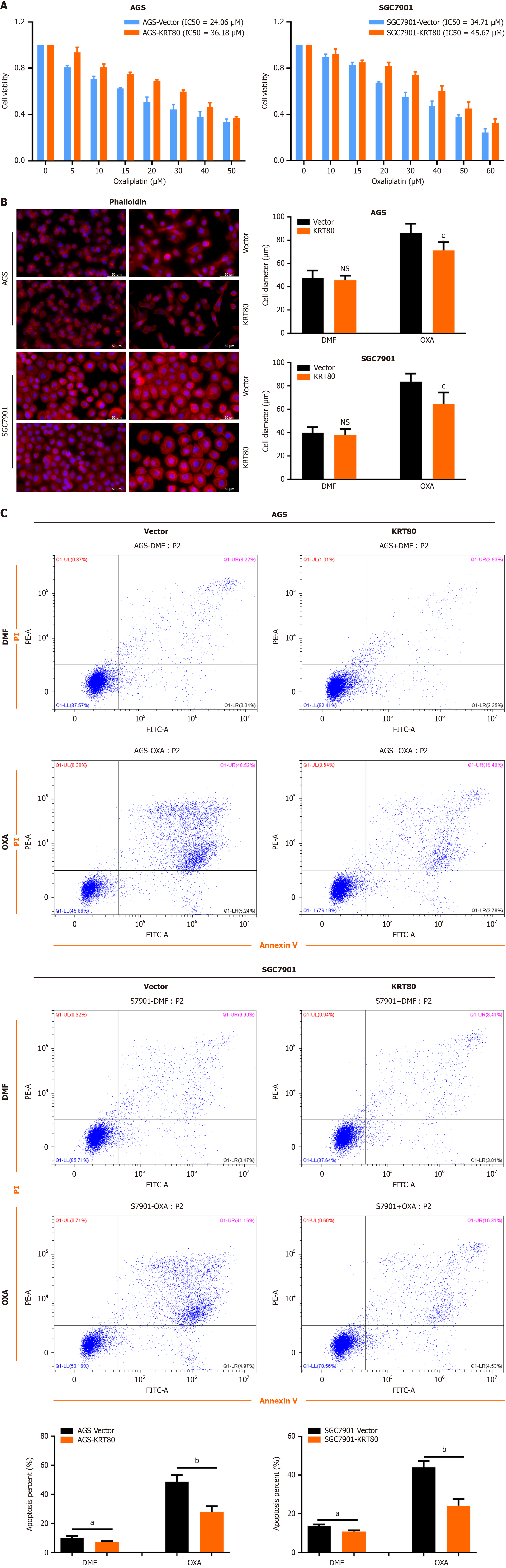
A Cell Counting Kit-8 assay was performed to assess oxaliplatin cytotoxicity in KRT80-overexpressing and control cells. The results showed that KRT80 overexpression reduced the sensitivity of GC cells to oxaliplatin. Specifically, the IC50 of oxaliplatin increased from 24.06 μM in control AGS cells to 36.18 μM in AGS-KRT80 cells. Similarly, in the SGC7901 group, the IC50 increased from 34.71 μM to 45.67 μM (Figure 4A).
Treatment with 10 μg/mL oxaliplatin induced morphological changes in GC cells, including spindle-shaped structures and cell scattering. To determine whether KRT80 regulates actin filament reassembly, fluorescent phalloidin staining was used to visualize filamentous actin distribution. Oxaliplatin treatment disrupted microfilament formation and increased the area covered by microfilaments. However, KRT80 overexpression mitigated oxaliplatin-induced damage to microfilaments, significantly reducing microfilament disruption (Figure 4B). These findings suggest that KRT80 overexpression protects against microfilament damage caused by oxaliplatin.
To further explore KRT80's role in oxaliplatin response, AGS and SGC7901 cells were treated with low concentrations of oxaliplatin for 48 hours. Flow cytometry analysis revealed that elevated KRT80 expression decreased oxaliplatin-induced apoptosis in GC cells (Figure 4C).
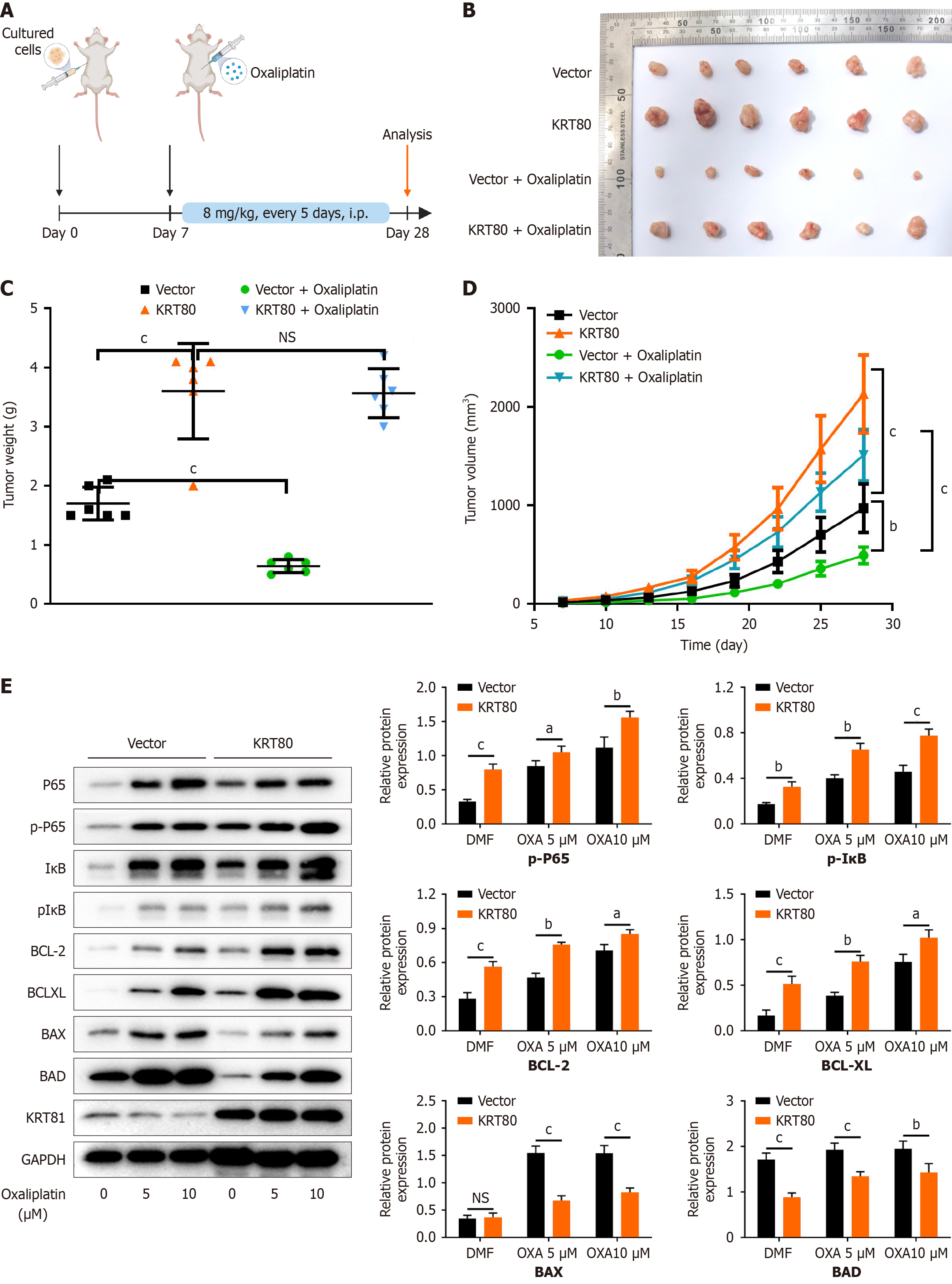
To validate the findings in vivo, a subcutaneous xenograft model was established using SGC7901 cells overexpressing KRT80 and corresponding control cells. The mice were treated with intraperitoneal injections of oxaliplatin to assess the impact of KRT80 on oxaliplatin's therapeutic efficacy (Figure 5A). The results showed that KRT80 overexpression significantly enhanced GC cell growth in vivo, with tumor volume and weight significantly greater than those in the control group. In the control group, oxaliplatin treatment effectively suppressed tumor growth. However, in the KRT80-overexpressing group, tumors showed no significant reduction in size following oxaliplatin treatment (Figure 5B-D). These findings indicates that KRT80 overexpression promotes tumor growth in nude mice and substantially diminishes the antitumor efficacy of oxaliplatin.
Subsequently, KRT80-overexpressing GC cells and their corresponding control cells were treated with varying concentrations of oxaliplatin. After 48 hours of treatment, the expression levels of related proteins were analyzed by Western blot. The results showed that the phosphorylation levels of NF-κB-p65 and IκB were significantly upregulated in the KRT80-overexpressing group. This effect was further enhanced by oxaliplatin treatment in a concentration-dependent manner (Figure 5E).
These findings suggest that KRT80 overexpression, in response to oxaliplatin, significantly activates the NF-κB signaling pathway, thereby enhancing the resistance of GC cells to oxaliplatin.
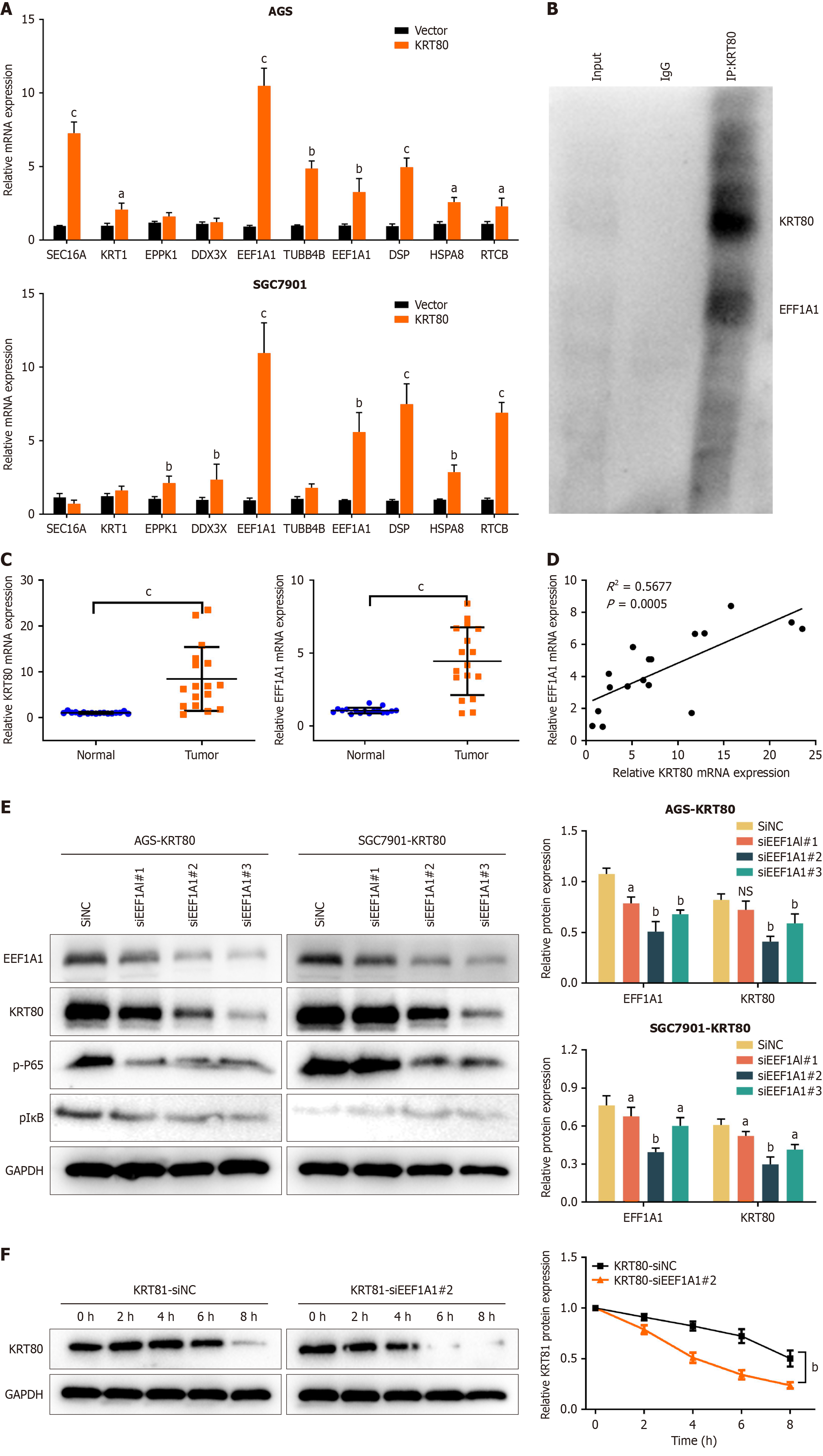
To identify potential KRT80-interacting proteins, co-immunoprecipitation was performed on SGC7901 cells overexpressing KRT80 and corresponding control cells, followed by mass spectrometry analysis of the total proteins. After comparing the mass spectrometry results between the two groups and eliminating redundant data, differentially expressed genes with the most significant variations were selected for further analysis (Figure 6A).
The mRNA expression levels of these target genes were quantified using real-time PCR in both AGS and SGC7901 cell lines. Co-immunoprecipitation experiments further confirmed that KRT80 physically interacted with the EEF1A1 protein, suggesting a potential interaction between KRT80 and EEF1A1 (Figure 6B).
Next, we performed qRT-PCR to examine the mRNA expression levels of KRT80 and EEF1A1 in GC tissues and their paired adjacent normal tissues. The results revealed that both genes were significantly upregulated in GC tissues compared to normal tissues (Figure 6C). Furthermore, correlation analysis demonstrated a positive relationship between the expression of KRT80 and EEF1A1 in tumor samples (R² = 0.5677, P = 0.0005; Figure 6D).
To investigate the regulatory relationship between KRT80 and EEF1A1, we employed siRNA to knockdown EEF1A1 in AGS and SGC7901 cells, resulting in a marked reduction in KRT80 protein expression. This suggests that EEF1A1 may positively regulate KRT80 protein levels. We also detected changes in protein levels of the NF-κB pathway, but no significant changes were found (Figure 6E).
To assess changes in protein stability, we inhibited intracellular protein synthesis using CHX and examined the impact of EEF1A1 knockdown on KRT80 (Figure 6F). The results indicated that EEF1A1 knockdown significantly accelerated the degradation of KRT80 in SGC7901 cells. Taken together, these findings suggest that the complex formed between KRT80 and EEF1A1 plays a critical role in enhancing KRT80 stability.
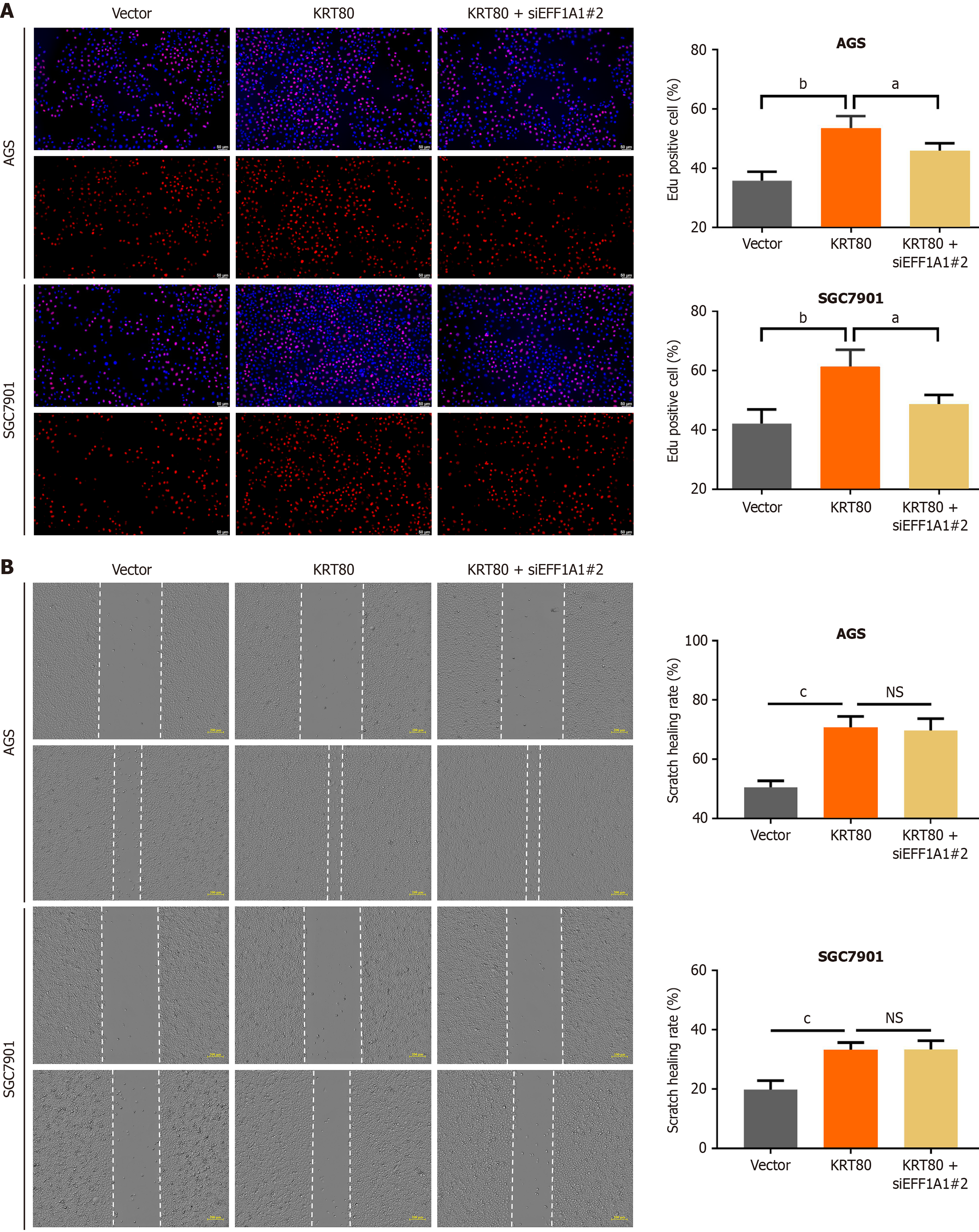
To investigate whether EEF1A1 expression influences the malignant phenotypes of KRT80-overexpressing cells, we knocked down EEF1A1 in these cells and assessed changes in proliferation using the EdU assay. The results showed that EEF1A1 knockdown partially reversed the increased proliferation induced by KRT80 overexpression (Figure 7A). Next, we performed a scratch wound assay to evaluate alterations in cell migration. However, silencing EEF1A1 had no significant effect on cell migration capacity (Figure 7B).
Despite significant advancements in GC research, the five-year survival rate remains suboptimal. Tumor metastasis, recurrence, and chemoresistance are the primary factors contributing to the poor prognosis of this malignancy[34]. Platinum-based chemotherapeutic agents are commonly used in the treatment of GC. However, both intrinsic and acquired resistance present major challenges[4,35]. Given the difficulty in developing novel antitumor drugs, a more practical approach is to focus on strategies that reduce tumor resistance and enhance the efficacy of existing therapies.
KRT80 is predominantly expressed in epithelial tissues and is closely associated with the development and progression of epithelial-derived tumors. In breast cancer, KRT80 expression correlates with tumor differentiation and metastatic potential; in colorectal cancer, it promotes tumor invasion and metastasis[20]; and in ovarian cancer, KRT80 has been identified as a potential diagnostic and prognostic biomarker[36].
This study represents the first systematic investigation of the effects of KRT80 expression on the malignant phenotype of GC cells, including proliferation, migration, and invasion. Our results show that KRT80 is upregulated in both GC cells and tumor tissues, and its upregulation significantly enhances the malignant characteristics of GC cells. Conversely, KRT80 knockdown substantially suppresses these malignant phenotypes. Furthermore, elevated KRT80 expression is strongly associated with unfavorable clinicopathological features and poor prognosis in GC patients. This suggests that KRT80 may serve as a potential GC marker.
Chemotherapy remains a cornerstone of treatment for advanced GC, with oxaliplatin-a second-generation platinum-based agent-widely recommended for both adjuvant and palliative therapy[37]. However, intrinsic or acquired resistance to oxaliplatin can lead to poor clinical outcomes. The antitumor action of oxaliplatin primarily involves the disruption of DNA replication and transcription, resulting in DNA damage and activation of cellular repair pathways. When DNA damage exceeds the cell's repair capacity, apoptotic pathways are activated, ultimately leading to tumor cell death[9,38].
Nevertheless, tumor cells can develop resistance to oxaliplatin through various mechanisms, including enhanced DNA repair, increased drug efflux, activation of nucleotide excision repair pathways, and inhibition of apoptotic signaling[39,40]. Overcoming chemoresistance in GC is crucial for improving patient outcomes, especially given the aggressive nature of advanced stages and the limited treatment options once resistance develops. KRT80 may offer promising clinical implications in this regard.
In this study, GC cells overexpressing KRT80 exhibited markedly enhanced resistance to the inhibitory effects of oxaliplatin, including reduced suppression of cell proliferation and apoptosis. In vivo, xenograft tumors derived from KRT80-overexpressing cells showed significantly larger tumor volumes, greater weight, and faster growth rates compared to control tumors, accompanied by a reduced tumor-suppressive effect of oxaliplatin. These findings suggest that KRT80 plays a critical role in modulating oxaliplatin-induced apoptosis and may contribute to the development of resistance, highlighting its potential as a therapeutic target to enhance oxaliplatin sensitivity in GC.
The NF-κB signaling pathway plays a pivotal role in regulating a wide range of cellular functions and is recognized as a key mediator of tumor chemoresistance[10]. Transcription factors within the NF-κB family regulate the expression of numerous target genes by binding to κB sites on DNA, thereby influencing cellular processes such as survival, proliferation, and apoptosis. Among the critical components of this pathway, the interactions between NF-κB, IκB, and the IKK complex are particularly important[11,16].
As an inhibitor, IκB binds to NF-κB and suppresses its transcriptional activity. However, upon external stimulation, such as exposure to chemotherapeutic agents, the IKK complex is activated. Activated IKK phosphorylates IκB, marking it for degradation. This degradation releases NF-κB, allowing it to translocate from the cytoplasm to the nucleus, where it activates the expression of target genes[15,41].
The NF-κB pathway often interacts with other signaling cascades, such as the PI3K-AKT pathway, forming an extensive regulatory network. Specifically, in chemotherapy-induced tumor treatment, activation of the PI3K-AKT pathway can stimulate IKK activity, thereby amplifying NF-κB signaling[13,42]. This crosstalk profoundly affects tumor cell biology, enhancing growth, resistance to apoptosis, and migratory capabilities.
Numerous studies have shown that activation of the NF-κB signaling pathway during chemotherapy contributes to heightened drug resistance in tumor cells. For instance, while paclitaxel induces apoptosis, it concurrently activates NF-κB, leading to the increased expression of the multidrug resistance gene MDR1 and its product, P-glycoprotein (P-gp)[14,15]. This upregulation enhances the ability of tumor cells to resist chemotherapeutic agents. In addition to the MDR1 gene, NF-κB also regulates the expression of anti-apoptotic proteins such as Bcl-XL, which, in concert with P-gp, fortifies cellular defenses against apoptotic signals[16,43]. These mechanisms underscore the critical role of NF-κB signaling in driving tumor chemoresistance.
The upregulation of the NF-κB signaling pathway is not only associated with tumorigenesis but also with chemo
We integrated mass spectrometry data with experimental results to confirm the interaction between EEF1A1 and KRT80. EEF1A1 is a key elongation factor involved in the elongation phase of protein synthesis. Its primary role is to facilitate the binding of aminoacyl-tRNA to ribosomes, thereby promoting protein synthesis. Increasing evidence suggests that EEF1A1 plays a pivotal role in tumor progression through interactions with various proteins[46]. For example, the interaction between EEF1A1 and TCTP has been shown to enhance proliferation and inhibit apoptosis in hepatocellular carcinoma cells[47]. Similarly, in breast cancer, EEF1A1 interacts with HSP90, contributing to the regu
This study confirmed the specific interaction between EEF1A1 and KRT80. Knockdown of EEF1A1 resulted in a shortened half-life of KRT80 protein in GC cells and partially abrogated the malignant phenotypic effects induced by KRT80 overexpression.
Integrating our experimental findings with multiple published studies, we found that increased KRT80 expression is associated with poor prognosis in GC and promotes resistance to oxaliplatin. Moreover, we have preliminarily verified the interaction between KRT80 and EEF1A1. Therefore, this study provides a novel perspective on overcoming oxaliplatin resistance in GC.
| 1. | Siegel RL, Giaquinto AN, Jemal A. Cancer statistics, 2024. CA Cancer J Clin. 2024;74:12-49. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2279] [Cited by in RCA: 4534] [Article Influence: 4534.0] [Reference Citation Analysis (3)] |
| 2. | Qiu H, Cao S, Xu R. Cancer incidence, mortality, and burden in China: a time-trend analysis and comparison with the United States and United Kingdom based on the global epidemiological data released in 2020. Cancer Commun (Lond). 2021;41:1037-1048. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 349] [Cited by in RCA: 572] [Article Influence: 143.0] [Reference Citation Analysis (0)] |
| 3. | Sonkin D, Thomas A, Teicher BA. Cancer treatments: Past, present, and future. Cancer Genet. 2024;286-287:18-24. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 160] [Article Influence: 160.0] [Reference Citation Analysis (0)] |
| 4. | Chia DKA, Sundar R, Kim G, Ang JJ, Lum JHY, Nga ME, Goh GH, Seet JE, Chee CE, Tan HL, Ho J, Ngoi NYL, Lee MXW, Muthu V, Chan GHJ, Pang ASL, Ang YLE, Choo JRE, Lim JSJ, Teh JL, Lwin A, Soon Y, Shabbir A, So JBY, Yong WP. Outcomes of a Phase II Study of Intraperitoneal Paclitaxel plus Systemic Capecitabine and Oxaliplatin (XELOX) for Gastric Cancer with Peritoneal Metastases. Ann Surg Oncol. 2022;29:8597-8605. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 8] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 5. | Yin Y, Lin Y, Yang M, Lv J, Liu J, Wu K, Liu K, Li A, Shuai X, Cai K, Wang Z, Wang G, Shen J, Zhang P, Tao K. Neoadjuvant tislelizumab and tegafur/gimeracil/octeracil (S-1) plus oxaliplatin in patients with locally advanced gastric or gastroesophageal junction cancer: Early results of a phase 2, single-arm trial. Front Oncol. 2022;12:959295. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 62] [Article Influence: 20.7] [Reference Citation Analysis (0)] |
| 6. | Zanardelli M, Micheli L, Nicolai R, Failli P, Ghelardini C, Di Cesare Mannelli L. Different apoptotic pathways activated by oxaliplatin in primary astrocytes vs. colo-rectal cancer cells. Int J Mol Sci. 2015;16:5386-5399. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 16] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 7. | Shen X, Zhang Y, Xu Z, Gao H, Feng W, Li W, Miao Y, Xu Z, Zong Y, Zhao J, Lu A. KLF5 inhibition overcomes oxaliplatin resistance in patient-derived colorectal cancer organoids by restoring apoptotic response. Cell Death Dis. 2022;13:303. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 32] [Article Influence: 10.7] [Reference Citation Analysis (0)] |
| 8. | Martinez-Balibrea E, Martínez-Cardús A, Ginés A, Ruiz de Porras V, Moutinho C, Layos L, Manzano JL, Bugés C, Bystrup S, Esteller M, Abad A. Tumor-Related Molecular Mechanisms of Oxaliplatin Resistance. Mol Cancer Ther. 2015;14:1767-1776. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 160] [Cited by in RCA: 234] [Article Influence: 23.4] [Reference Citation Analysis (0)] |
| 9. | Greenlee JD, Lopez-Cavestany M, Ortiz-Otero N, Liu K, Subramanian T, Cagir B, King MR. Oxaliplatin resistance in colorectal cancer enhances TRAIL sensitivity via death receptor 4 upregulation and lipid raft localization. Elife. 2021;10:e67750. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 33] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 10. | Choi MC, Jo J, Park J, Kang HK, Park Y. NF-κB Signaling Pathways in Osteoarthritic Cartilage Destruction. Cells. 2019;8:734. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 175] [Cited by in RCA: 360] [Article Influence: 60.0] [Reference Citation Analysis (0)] |
| 11. | Oeckinghaus A, Ghosh S. The NF-kappaB family of transcription factors and its regulation. Cold Spring Harb Perspect Biol. 2009;1:a000034. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1512] [Cited by in RCA: 2131] [Article Influence: 142.1] [Reference Citation Analysis (0)] |
| 12. | Napetschnig J, Wu H. Molecular basis of NF-κB signaling. Annu Rev Biophys. 2013;42:443-468. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 594] [Cited by in RCA: 759] [Article Influence: 63.3] [Reference Citation Analysis (0)] |
| 13. | Yang L, Hu X, Mo YY. Acidosis promotes tumorigenesis by activating AKT/NF-κB signaling. Cancer Metastasis Rev. 2019;38:179-188. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 25] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 14. | Lee ST, Wong PF, Hooper JD, Mustafa MR. Alpha-tomatine synergises with paclitaxel to enhance apoptosis of androgen-independent human prostate cancer PC-3 cells in vitro and in vivo. Phytomedicine. 2013;20:1297-1305. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 28] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 15. | Yuan Z, Liang X, Zhan Y, Wang Z, Xu J, Qiu Y, Wang J, Cao Y, Le VM, Ly HT, Xu J, Li W, Yin P, Xu K. Targeting CD133 reverses drug-resistance via the AKT/NF-κB/MDR1 pathway in colorectal cancer. Br J Cancer. 2020;122:1342-1353. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 48] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 16. | Bailey ST, Miron PL, Choi YJ, Kochupurakkal B, Maulik G, Rodig SJ, Tian R, Foley KM, Bowman T, Miron A, Brown M, Iglehart JD, Debajit KB. NF-κB activation-induced anti-apoptosis renders HER2-positive cells drug resistant and accelerates tumor growth. Mol Cancer Res. 2014;12:408-420. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 42] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 17. | Ryan SL, Beard S, Barr MP, Umezawa K, Heavey S, Godwin P, Gray SG, Cormican D, Finn SP, Gately KA, Davies AM, Thompson EW, Richard DJ, O'Byrne KJ, Adams MN, Baird AM. Targeting NF-κB-mediated inflammatory pathways in cisplatin-resistant NSCLC. Lung Cancer. 2019;135:217-227. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 24] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 18. | Wei XY, Zhao J, Tong HB, Cheng SJ, He N, Song FX. Characters of KRT80 and its roles in neoplasms diseases. Cancer Med. 2023;12:13991-14003. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 5] [Reference Citation Analysis (0)] |
| 19. | Shao MM, Chan SK, Yu AM, Lam CC, Tsang JY, Lui PC, Law BK, Tan PH, Tse GM. Keratin expression in breast cancers. Virchows Arch. 2012;461:313-322. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 71] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 20. | Li C, Liu X, Liu Y, Liu X, Wang R, Liao J, Wu S, Fan J, Peng Z, Li B, Wang Z. Keratin 80 promotes migration and invasion of colorectal carcinoma by interacting with PRKDC via activating the AKT pathway. Cell Death Dis. 2018;9:1009. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 46] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 21. | Huang S, Tong W, Yang B, Ma L, Zhang J, Wang C, Xu L, Mei J. KRT80 Promotes Lung Adenocarcinoma Progression and Serves as a Substrate for VCP. J Cancer. 2024;15:2229-2244. [RCA] [PubMed] [DOI] [Full Text] [Reference Citation Analysis (0)] |
| 22. | Misago N, Inoue T, Koba S, Narisawa Y. Keratoacanthoma and other types of squamous cell carcinoma with crateriform architecture: classification and identification. J Dermatol. 2013;40:443-452. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 32] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 23. | Li R, Liu H, Dilger JP, Lin J. Effect of Propofol on breast Cancer cell, the immune system, and patient outcome. BMC Anesthesiol. 2018;18:77. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 76] [Cited by in RCA: 67] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 24. | Li R, Mukherjee MB, Jin Z, Liu H, Lin K, Liu Q, Dilger JP, Lin J. The Potential Effect of General Anesthetics in Cancer Surgery: Meta-Analysis of Postoperative Metastasis and Inflammatory Cytokines. Cancers (Basel). 2023;15:2759. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 19] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 25. | Jin Z, Zhang W, Liu H, Ding A, Lin Y, Wu SX, Lin J. Potential Therapeutic Application of Local Anesthetics in Cancer Treatment. Recent Pat Anticancer Drug Discov. 2022;17:326-342. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Reference Citation Analysis (0)] |
| 26. | Liu H, Dong A, Rasteh AM, Wang P, Weng J. Identification of the novel exhausted T cell CD8 + markers in breast cancer. Sci Rep. 2024;14:19142. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 59] [Article Influence: 59.0] [Reference Citation Analysis (0)] |
| 27. | Li R, Xiao C, Liu H, Huang Y, Dilger JP, Lin J. Effects of local anesthetics on breast cancer cell viability and migration. BMC Cancer. 2018;18:666. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 90] [Cited by in RCA: 81] [Article Influence: 11.6] [Reference Citation Analysis (0)] |
| 28. | Ou L, Liu H, Peng C, Zou Y, Jia J, Li H, Feng Z, Zhang G, Yao M. Helicobacter pylori infection facilitates cell migration and potentially impact clinical outcomes in gastric cancer. Heliyon. 2024;10:e37046. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 18] [Article Influence: 18.0] [Reference Citation Analysis (0)] |
| 29. | Liu H, Dilger JP, Lin J. Lidocaine Suppresses Viability and Migration of Human Breast Cancer Cells: TRPM7 as a Target for Some Breast Cancer Cell Lines. Cancers (Basel). 2021;13:234. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 54] [Cited by in RCA: 52] [Article Influence: 13.0] [Reference Citation Analysis (0)] |
| 30. | Lu Y, Yang B, Shen A, Yu K, Ma M, Li Y, Wang H. LncRNA UCA1 promotes vasculogenic mimicry by targeting miR-1-3p in gastric cancer. Carcinogenesis. 2024;45:658-672. [RCA] [PubMed] [DOI] [Full Text] [Reference Citation Analysis (0)] |
| 31. | Lu Y, Wang H, Chen S, Yang B, Li Y, Li Y. Cystatin SA attenuates gastric cancer cells growth and increases sensitivity to oxaliplatin via PI3K/AKT signaling pathway. J Cancer Res Clin Oncol. 2024;150:244. [RCA] [PubMed] [DOI] [Full Text] [Reference Citation Analysis (0)] |
| 32. | Li R, Huang Y, Liu H, Dilger JP, Lin J. Abstract 2162: Comparing volatile and intravenous anesthetics in a mouse model of breast cancer metastasis. Cancer Res. 2018;78:2162-2162. [RCA] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 29] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 33. | Liu H, Dilger JP, Lin J. Effects of local anesthetics on cancer cells. Pharmacol Ther. 2020;212:107558. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 66] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 34. | Joshi SS, Badgwell BD. Current treatment and recent progress in gastric cancer. CA Cancer J Clin. 2021;71:264-279. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 382] [Cited by in RCA: 1070] [Article Influence: 267.5] [Reference Citation Analysis (0)] |
| 35. | Kang L, Tian Y, Xu S, Chen H. Oxaliplatin-induced peripheral neuropathy: clinical features, mechanisms, prevention and treatment. J Neurol. 2021;268:3269-3282. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 108] [Article Influence: 21.6] [Reference Citation Analysis (0)] |
| 36. | Liu O, Wang C, Wang S, Hu Y, Gou R, Dong H, Li S, Li X, Lin B. Keratin 80 regulated by miR-206/ETS1 promotes tumor progression via the MEK/ERK pathway in ovarian cancer. J Cancer. 2021;12:6835-6850. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 17] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 37. | Fischel JL, Formento P, Ciccolini J, Rostagno P, Etienne MC, Catalin J, Milano G. Impact of the oxaliplatin-5 fluorouracil-folinic acid combination on respective intracellular determinants of drug activity. Br J Cancer. 2002;86:1162-1168. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 36] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 38. | Limagne E, Thibaudin M, Nuttin L, Spill A, Derangère V, Fumet JD, Amellal N, Peranzoni E, Cattan V, Ghiringhelli F. Trifluridine/Tipiracil plus Oxaliplatin Improves PD-1 Blockade in Colorectal Cancer by Inducing Immunogenic Cell Death and Depleting Macrophages. Cancer Immunol Res. 2019;7:1958-1969. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 102] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 39. | Shi Y, Niu Y, Yuan Y, Li K, Zhong C, Qiu Z, Li K, Lin Z, Yang Z, Zuo D, Qiu J, He W, Wang C, Liao Y, Wang G, Yuan Y, Li B. PRMT3-mediated arginine methylation of IGF2BP1 promotes oxaliplatin resistance in liver cancer. Nat Commun. 2023;14:1932. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 54] [Reference Citation Analysis (0)] |
| 40. | Li Y, Gan Y, Liu J, Li J, Zhou Z, Tian R, Sun R, Liu J, Xiao Q, Li Y, Lu P, Peng Y, Peng Y, Shu G, Yin G. Downregulation of MEIS1 mediated by ELFN1-AS1/EZH2/DNMT3a axis promotes tumorigenesis and oxaliplatin resistance in colorectal cancer. Signal Transduct Target Ther. 2022;7:87. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 100] [Article Influence: 33.3] [Reference Citation Analysis (0)] |
| 41. | Kanarek N, Ben-Neriah Y. Regulation of NF-κB by ubiquitination and degradation of the IκBs. Immunol Rev. 2012;246:77-94. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 227] [Cited by in RCA: 266] [Article Influence: 20.5] [Reference Citation Analysis (0)] |
| 42. | Yu M, Qi B, Xiaoxiang W, Xu J, Liu X. Baicalein increases cisplatin sensitivity of A549 lung adenocarcinoma cells via PI3K/Akt/NF-κB pathway. Biomed Pharmacother. 2017;90:677-685. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 74] [Cited by in RCA: 170] [Article Influence: 21.3] [Reference Citation Analysis (0)] |
| 43. | Haselager M, Thijssen R, West C, Young L, Van Kampen R, Willmore E, Mackay S, Kater A, Eldering E. Regulation of Bcl-XL by non-canonical NF-κB in the context of CD40-induced drug resistance in CLL. Cell Death Differ. 2021;28:1658-1668. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 58] [Article Influence: 14.5] [Reference Citation Analysis (0)] |
| 44. | Lei ZN, Teng QX, Tian Q, Chen W, Xie Y, Wu K, Zeng Q, Zeng L, Pan Y, Chen ZS, He Y. Signaling pathways and therapeutic interventions in gastric cancer. Signal Transduct Target Ther. 2022;7:358. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 32] [Cited by in RCA: 157] [Article Influence: 52.3] [Reference Citation Analysis (1)] |
| 45. | Wang Y, Fu Q, Zhao W. Tetramethylpyrazine inhibits osteosarcoma cell proliferation via downregulation of NF-κB in vitro and in vivo. Mol Med Rep. 2013;8:984-988. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 26] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 46. | Duman M, Vaquié A, Nocera G, Heller M, Stumpe M, Siva Sankar D, Dengjel J, Meijer D, Yamaguchi T, Matthias P, Zeis T, Schaeren-Wiemers N, Hayoz A, Ruff S, Jacob C. EEF1A1 deacetylation enables transcriptional activation of remyelination. Nat Commun. 2020;11:3420. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 38] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 47. | Leclercq TM, Moretti PA, Pitson SM. Guanine nucleotides regulate sphingosine kinase 1 activation by eukaryotic elongation factor 1A and provide a mechanism for eEF1A-associated oncogenesis. Oncogene. 2011;30:372-378. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 40] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 48. | Tash JS, Chakrasali R, Jakkaraj SR, Hughes J, Smith SK, Hornbaker K, Heckert LL, Ozturk SB, Hadden MK, Kinzy TG, Blagg BS, Georg GI. Gamendazole, an orally active indazole carboxylic acid male contraceptive agent, targets HSP90AB1 (HSP90BETA) and EEF1A1 (eEF1A), and stimulates Il1a transcription in rat Sertoli cells. Biol Reprod. 2008;78:1139-1152. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 77] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 49. | Chen SL, Lu SX, Liu LL, Wang CH, Yang X, Zhang ZY, Zhang HZ, Yun JP. eEF1A1 Overexpression Enhances Tumor Progression and Indicates Poor Prognosis in Hepatocellular Carcinoma. Transl Oncol. 2018;11:125-131. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 31] [Article Influence: 3.9] [Reference Citation Analysis (0)] |