Published online Nov 28, 2022. doi: 10.3748/wjg.v28.i44.6282
Peer-review started: September 2, 2022
First decision: September 30, 2022
Revised: September 30, 2022
Accepted: November 17, 2022
Article in press: November 17, 2022
Published online: November 28, 2022
Processing time: 83 Days and 20.2 Hours
Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) is the pathogen responsible for pandemic coronavirus disease 2019 (COVID-19). It is a highly contagious virus which primarily affects the respiratory tract, nevertheless, the lungs are not the only target organs of the virus. The intestinal tract could represent an additional tropism site for SARS-CoV-2. Several observations have collectively suggested that enteric infections can occur in COVID-19 patients. However, the detection of viral RNA in gastrointestinal (GI) tissue samples has not been adequately investigated and results are conflicting.
To detect the presence of SARS-CoV-2 RNA in intestinal mucosa samples and to evaluate histological features.
The COVID-19 patients hospitalized at an Italian tertiary hospital from April 2020 to March 2021 were evaluated for enrollment in an observational, monocentric trial. The study population was composed of two groups of adult patients. In the first group (biopsy group, 30 patients), patients were eligible for inclusion if they had mild to moderate disease and if they agreed to have a rectal biopsy; in the second group (surgical specimen group, 6 patients), patients were eligible for inclusion if they underwent intestinal resection during index hospitalization. Fifty-nine intestinal mucosal samples were analyzed.
Viral RNA was not detectable in any of the rectal biopsies performed (0/53). Histological examination showed no enterocyte damage, but slight edema of the lamina propria with mild inflammatory lymphoplasmacytic infiltration. There was no difference in inflammatory infiltrates in patients with and without GI symptoms. SARS-CoV-2 RNA was detected in fecal samples in 6 cases out of 14 cases examined (42.9%). In the surgical specimen group, all patients underwent emergency intestinal resection. Viral RNA was detected in 2 surgical specimens of the 6 examined, both of which were from patients with active neoplastic disease. Histological examination also pointed out abundant macrophages, granulocytes and plasma cells infiltrating the muscular layer and adipose tissue, and focal vasculitis.
Mild-moderate COVID-19 may not be associated with rectal infection by the virus. More comprehensive autopsies or surgical specimens are needed to provide histological evidence of intestinal infection.
Core Tip: The detection of viral RNA in gastrointestinal tissue samples has not been adequately investigated. In this trial, 53 rectal biopsies and 6 surgical specimens from coronavirus disease 2019 patients were analyzed, with the primary objective of detecting severe acute respiratory syndrome coronavirus-2 RNA, using real time reverse transcriptase-polymerase chain reaction, and evaluating the histological features. Viral RNA was not detectable in any of the rectal biopsies performed (0/53). Histological examination of rectal biopsies showed no enterocyte damage, but mild inflammatory infiltration. Viral RNA was detected in 2 surgical specimens of the 6 examined, both of which were from patients with active neoplastic disease. Histological examination also pointed out mild inflammatory infiltration and focal vasculitis.
- Citation: Cuicchi D, Gabrielli L, Tardio ML, Rossini G, D’Errico A, Viale P, Lazzarotto T, Poggioli G. Virological and histological evaluation of intestinal samples in COVID-19 patients. World J Gastroenterol 2022; 28(44): 6282-6293
- URL: https://www.wjgnet.com/1007-9327/full/v28/i44/6282.htm
- DOI: https://dx.doi.org/10.3748/wjg.v28.i44.6282
Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) is the pathogen responsible for pandemic coronavirus disease 2019 (COVID-19). It is a highly contagious virus which primarily affects the respiratory tract, typically causing symptoms, such as fever, dry cough and dyspnea, up to respiratory failure and death[1]. Nevertheless, the lungs are not the only target organs of the virus. Several studies have suggested that the intestinal tract could represent an additional tropism site for SARS-CoV-2[2]. Gastrointestinal (GI) symptoms, including diarrhea, nausea, vomiting, anorexia and abdominal pain are present in a substantial number of COVID-19 patients; the incidence can vary from 10% to 55%[3-6]. Many studies have reported that viral nucleic acids are detected in stool samples of COVID-19 patients with rates varying from 15.3% to 66.7%[7-11]. Viral receptor angiotensin-converting enzyme 2 (ACE2) and transmembrane protease serine-type 2 are highly expressed in the epithelial cells of the intestinal mucosa[7,12]. Moreover, some studies have demonstrated that SARS-CoV-2 virus can actively infect and replicate in human enteroids and in vitro models of human intestinal epithelium[13,14]. These observations have collectively suggested that enteric infection can occur in patients with COVID-19. However, more robust evidence is limited. The detection of viral RNA in GI tissue samples or the isolation of the virus in stool samples have been reported in studies which enrolled only a limited number of cases[15-18]. In the present study, a greater number of intestinal mucosal samples from COVID-19 patients (as compared to previous studies) were analyzed, with the primary objective of detecting SARS-CoV-2 RNA and evaluating histological features.
The COVID-19 patients hospitalized at IRCCS Sant’Orsola Hospital, University of Bologna, Bologna, Italy from June 2020 to March 2021 were evaluated for enrollment in a monocentric trial. SARS-CoV-2 infection was diagnosed at admission using real-time reverse transcriptase-polymerase chain reaction (RT-PCR) on pharyngeal swab specimens. The study was approved by the local hospital ethics committee and informed consent was obtained from all of the patients.
The study population was composed of two groups of adult patients (≥ 18 years of age) hospitalized for COVID-19. In the first group (biopsy group), patients were eligible for inclusion if they had mild to moderate disease (mild: Only mild symptoms with no radiological signs; moderate: Characterized by fever, respiratory symptoms and radiological signs of pneumonia)[19] and if they agreed to have a rectal biopsy; in the second group (surgical specimen group), patients were eligible for inclusion if they underwent intestinal resection during index hospitalization.
In the biopsy group, patients who had severe (dyspnea, respiratory frequency ≥ 30 breaths per minute, blood oxygen saturation ≤ 93%, partial pressure of arterial oxygen to fraction of inspired oxygen ratio < 300, and/or lung infiltrates > 50% within 24-48 h) and critical (respiratory failure, septic shock, and/or multiple organ dysfunction or failure) disease[19], contraindications to rectal biopsies (anticoagulant therapy with the exception of therapy with low molecular weight heparin and antiplatelet therapy), rectal disease (e.g., chronic inflammatory disease and proctitis), previous abdominoperineal resection, recent anal surgery, anal stenosis or anal pain were excluded. In the surgical specimen group, those patients undergoing surgical treatment, cases without intestinal resection were excluded.
Patients enrolled in the biopsy group were asked to collect a stool sample and undergo anoscopy with biopsy during their hospital stay. The anoscopy was performed at the bedside with the patient in the left lateral decubitus position using a disposable anoscope 18 mm diameter, lubricated with an anesthetic gel. During each anoscopy, 2 biopsies were performed at different sites of the rectal mucosa for viral RNA detection and stored, one in RNA Preservation Medium (RNAlaterTM Stabilization Solution, Thermo Fisher Scientific, United States) and then fresh frozen, and one in 10% buffered formalin. The biopsy specimen fixed in formalin was subsequently paraffin embedded, cut into 4-μm-thick sections and then stained with hematoxylin and eosin. Multiple sections were obtained to assess the extent of the inflammation. Serial paraffin sections, 3-μm-thick, mounted on precoated slides were processed using standardized automated procedures with prediluted anti-CD68 antibody (clone PG-M1; DBS, Pleasanton, CA). The stool samples and all the biopsies were tested for the detection of SARS-CoV-2 RNA using real-time RT-PCR. The extraction of nucleic acids, reverse transcription reaction and real-time PCR amplification were performed using a SARS-CoV-2 ELITe MGB® Kit (ELITechGroup, Italy) on an ELITeInGenius® instrument. The assay detects the RNA of two SARS-CoV-2 specific genomic regions: RdRp gene and ORF8 gene. The tissue viral load was reported as number of copies/microgram RNA. The limit of detection of the test is 2 copies/reaction (10 copies/microgram RNA). Positive results below the lower limit of quantification (5 copies/reaction) were reported as < 25 copies/microgram RNA. Qualitative data were provided for the fecal samples.
Patients enrolled in the surgical specimen group were asked for consent in order to take a tissue sample from the surgical specimen to identify the presence of both inflammatory cells infiltrated and virus SARS-CoV-2 RNA using the same methodology described above. For each patient, data were collected regarding sex, age, comorbidities, disease severity, symptoms on admission, radiological features and clinical outcomes.
The primary aim was to evaluate the presence of SARS-CoV-2 RNA in intestinal mucosa samples using real-time RT-PCR. The secondary aims were to detect both SARS-CoV-2 RNA in stool samples using RT-PCR and the inflammatory state in all tissue samples.
Sample size analysis was not carried out due to the exploratory nature of the study. Statistical analysis was carried out using SPSS 20 software (SPSS Inc, Chicago, IL). Continuous data were expressed as means ± SDs (for data with normal distribution), or median and range (for data with nonnormal distribution); discrete data were expressed as percentages. The descriptive analyses were carried out using parametric methods, depending on the distribution of the variables under examination. Variables between the 2 groups were compared using the following tests as appropriate: t-test and Fisher’s test. The differences were considered statistically significant for values of < 0.05.
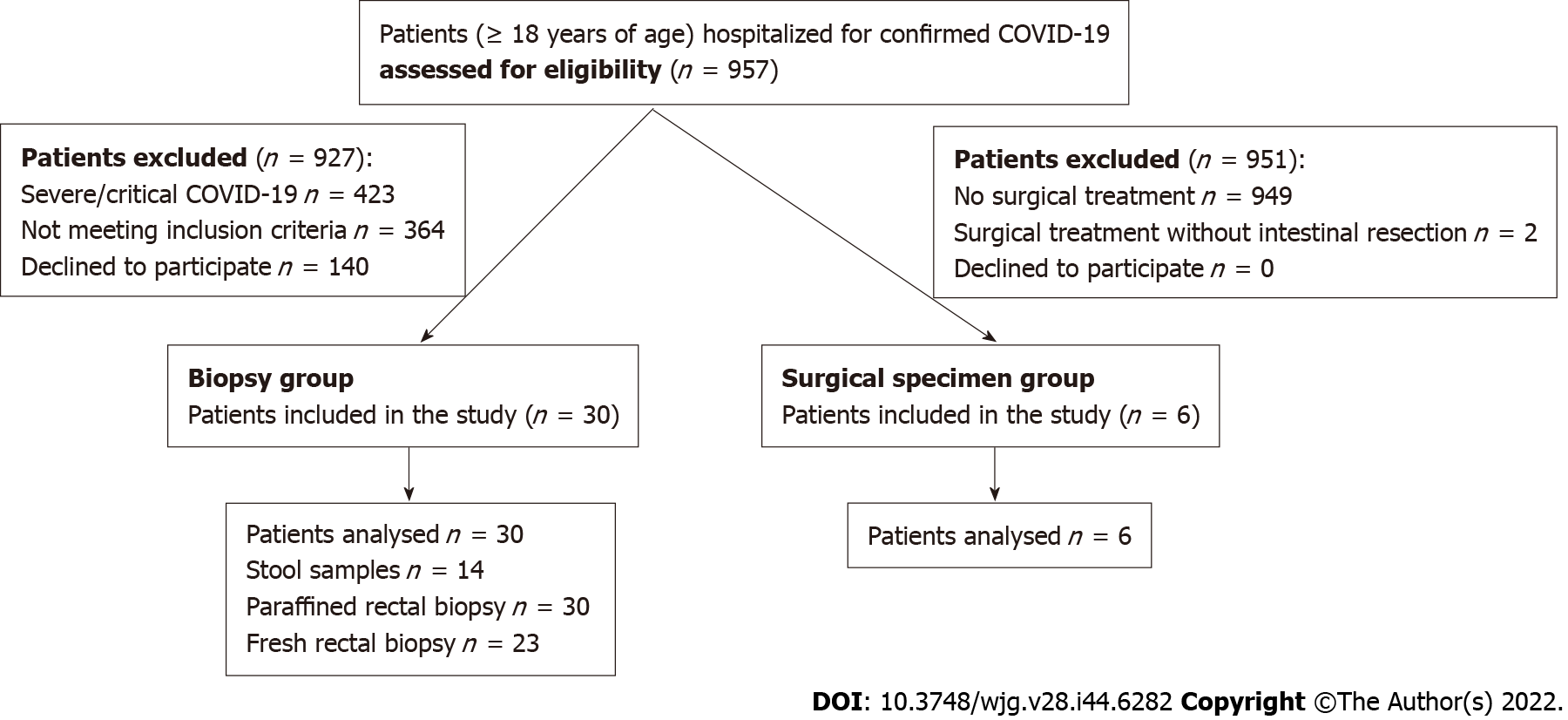
From June 2020 to March 2021, 957 patients hospitalized for confirmed COVID-19 were screened. The diagram in Figure 1 shows the flow of participants in the trial. The biopsy group consisted of 30 patients and the surgical specimen group had 6 patients. The clinical characteristics of the patients in both groups are shown in Tables 1 and 2. In the biopsy group, GI symptoms were present in 19 patients (63.3%). Diarrhea was the most common GI symptom; it was reported in 14 cases (73.7%), anorexia in 8 cases (42.1%), nausea and vomiting in 7 cases (36.8%) and abdominal pain in 4 cases (21.0%). There was no statistically significant difference in the general demographics or clinical outcomes between patients with and without GI symptoms (Table 1); SARS-CoV-2 RNA was detected in the stool in 36.4% of the patients with GI symptoms and in 66.7% of those without GI symptoms. Nevertheless, the number of positive fecal cases did not show significant difference between the two groups. Considering only patients who had diarrhea on admission, viral RNA was found in half of the fecal samples examined. Overall, SARS-CoV-2 RNA was detected in fecal samples in 6 cases out of 14 cases examined (42.9%). The greatest number of positive cases was found in the fecal samples collected in the 2nd wk after the first positive nasopharyngeal swab (NPS) (3/3, 100%), 2 cases in the 1st wk (2/5, 40%) and the remaining case in the 3rd wk (1/4, 25%). There was no significant difference in time interval between sampling and the first positive NPS in positive and negative viral RNA fecal samples (9.0 ± 4.6 vs 16.4 ± 14.7 respectively, P = 0.26). Viral RNA was not detectable in any of the 53 rectal biopsies performed. The histological examination of the rectal samples showed that the mucosal epithelium of the rectum did not have any major damage in patients with and without GI symptoms. The glandular architecture was always normal. In no case was there any enterocyte damage. Moreover, microscopy revealed slight expansion of the lamina propria by moderate edema in 26 cases out of 30 (86.7%) and an inflammatory lymphoplasmacytic infiltration in the lamina propria, varying from mild to moderate, in 28 and 2 cases (93.3% and 6.7%), respectively. There was no difference in inflammatory infiltrates in patients with and without GI symptoms (Table 1). Rare eosinophilic and neutrophil granulocytes were identified in the lamina propria in 20 and 2 cases (66.7% and 6.7%), respectively.
| Characteristic | All patients, n = 30 | Patients with GI symptoms at onset, n = 19 | Patients without GI symptoms at onset, n = 11 | P value |
| Median age in yr, mean (range) | 65 (41-90) | 60 (44-90) | 78 (41-89) | NS |
| Sex as women:men (%) | 7:23 (23.3:76.7) | 4:15 (21.0:79) | 3:8 (27.3:72.7) | NS |
| COVID-19 classification | ||||
| Mild, n (%) | 0 | 0 | 0 | NS |
| Moderate, n (%) | 30 (100) | 19 (100) | 11 (100) | NS |
| Coexisting illness | ||||
| Hypertension | 9 (30.0) | 6 (31.6) | 3 (27.3) | |
| Diabetes mellitus | 4 (13.3) | 3 (15.8) | 1 (9.1) | |
| Cardio-cerebrovascular disease | 5 (16.7) | 3 (15.8) | 2 (18.2) | |
| Previous malignant tumor | 2 (6.7) | 1 (5.3) | 1 (9.1) | |
| Chronic obstructive pulmonary disease | 3 (10.0) | 2 (10.5) | 1 (9.1) | |
| Chronic kidney disease | 3 (10.0) | 2 (10.5) | 1 (9.1) | |
| Obesity | 2 (6.7) | 2 (10.5) | 0 | |
| Chest CT | ||||
| Negative, n (%) | 4 (13.3) | 3 (15.8) | 1 (9.1) | NS |
| Bilateral distribution of GGO with or without consolidation, n (%) | 16 (53.3) | 9 (47.4) | 7 (63.6) | NS |
| Unilateral distribution of GGO, n (%) | 2 (6.7) | 1 (5.2) | 1 (9.1) | NS |
| Bilateral interlobular septal thickening, n (%) | 8 (26.7) | 6 (31.6) | 2 (18.2) | NS |
| Median time from first positive NPS (range), d | 10 (1-25) | 8.0 (1-25) | 12.0 (3-24) | NS |
| SARS-CoV-2 RNA in feces1, n (%) | 6/14 (42.9) | 4/11 (36.4) | 2/3 (66.7) | NS |
| SARS-CoV-2 RNA in RNA-preservation medium rectal mucosa biopsy2, n (%) | 0 | 0 | 0 | - |
| SARS-CoV-2 RNA in FFPE rectal mucosa biopsy3, n (%) | 0 | 0 | 0 | - |
| Histological examination | ||||
| Glandular architecture: Normal/altered, n (%) | 30/0 (100/0) | 19/0 (100/0) | 11/0 (100/0) | NS |
| Edema of the lamina propria: Absent/slight, n (%) | 4/26 (13.3/86.7) | 3/16 (15.8/84.2) | 1/10 (9.1/90.9) | NS |
| Inflammatory lymphoplasmacytic infiltration in the lamina propria: Mild/moderate, n (%) | 28/2 (93.3/6.7) | 18/1 (94.7/5.3) | 10/1 (90.9/9.1) | NS |
| Eosinophilic granulocytes in the lamina propria: Absent/occasional/scattered, n (%) | 10/18/2 (33.3/60.0/6.7) | 7/10/2 (36.8/52.6/10.7) | 3/8/0 (27.3/72.2/0) | NS |
| Neutrophil granulocytes in the lamina propria: Absent/rare, n (%) | 28/2 (93.3/6.7) | 18/1 | 10/1 (90.9/9.1) | NS |
| Enterocyte damage: Absent/present, n (%) | 30/0 (100/0) | 19/0 | 11/0 (100/0) | NS |
| Clinical outcome, n (%) | ||||
| Discharged | 30 (100) | 19 (100) | 11 (100) | NS |
| Died | 0 | 0 | 0 | NS |
| Characteristic | Patient no. 1 | Patient no. 2 | Patient no. 3 | Patient no. 4 | Patient no. 5 | Patient no. 6 |
| Age in yr | 93 | 76 | 78 | 96 | 56 | 62 |
| Coexisting illness | COPD, CVD | COPD, metastatic colon cancer | COPD, CVD, chronic kidney disease | COPD, hypertension | Psychiatric disease, obesity | Acute myeloid leukemia |
| COVID-19 classification | Moderate | Moderate | Mild | Mild | Moderate | Moderate |
| Thorax CT imaging | Bilateral distribution of GGO with consolidation | Unilateral distribution of GGO with consolidation | Bilateral distribution of GGO with consolidation | Unilateral distribution of GGO with consolidation | Bilateral distribution of GGO with consolidation | Bilateral distribution of GGO with consolidation |
| Time from first positive NPS | 10 | 1 | 8 | 1 | 20 | 7 |
| Reason for admission | COVID-19 | Spontaneous pneumothorax | COVID-19 | Intestinal occlusion | COVID-19 | Chemotherapy for acute myeloid leukemia |
| Reason for surgical treatment | Rectal perforation after enema | Fecal peritonitis from perforated colon cancer | Fecal peritonitis from perforated diverticulitis | Small bowel obstruction secondary to bridle | Ischemic colitis | Acute abdomen |
| Surgical treatment | Colorectal resection and end-colostomy | Right colectomy and end-ileostomy | Sigmoid resection and end-colostomy | Segmental intestinal resection | Total colectomy and ileostomy | Appendectomy and sigmoid resection |
| GI symptoms at onset | No | No | No | No | No | Diarrhea |
| SARS-CoV-2 RNA in FFPE tissue/viral load (n° copies/microg) | No | 29 | No | No | No | < 25 |
| Histological examination | ||||||
| Glandular architecture | Normal | Normal | Normal | Normal | Normal | Normal |
| Edema of the lamina propria | Slight | Absent | Slight | Slight | Slight | Absent |
| Inflammatory lymphoplasmacytic infiltration in the lamina propria | Mild | Mild | Mild | Mild | Mild | Moderate |
| Eosinophilic granulocytes in the lamina propria | Occasional | Occasional | Occasional | Occasional | Occasional | Occasional |
| Enterocyte damage | Absent | Absent | Absent | Absent | Absent | Absent |
| Vasculitis | Absent | Focal | Absent | Absent | Absent | Focal |
| Granulocyte, macrophage and plasma cell infiltrate in the muscle wall and adipose tissue | Absent | Moderate | Absent | Absent | Absent | Severe |
| Clinical outcome | Died | Died | Died | Died | Died | Died |
| Time from surgical treatment in d | 6 | 4 | 4 | 35 | 17 | 20 |
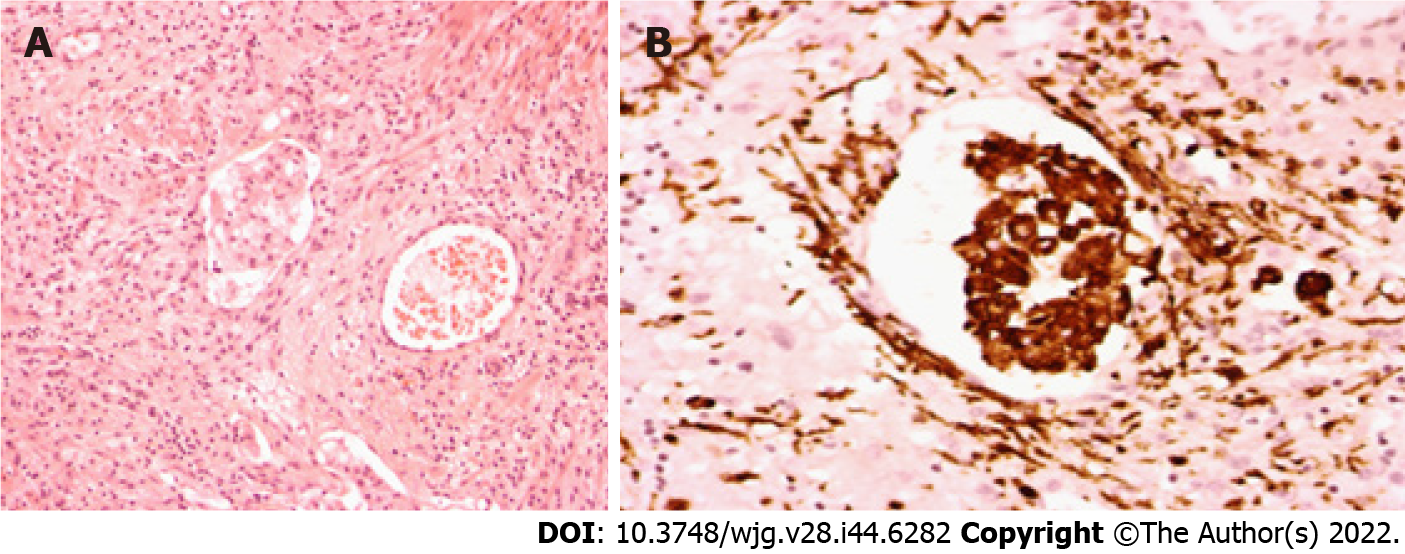
In the surgical specimen group, all patients underwent emergency intestinal resection (Table 2). Three patients were hospitalized for COVID-19 symptoms and, 8-20 d after the first positive NPS, underwent bowel resection for iatrogenic perforation of the rectum, diverticular perforation and ischemic colitis, respectively. One patient was hospitalized for intestinal obstruction secondary to bridles and underwent segmental resection of the small intestine within 1 d after a positive pharyngeal swab for SARS-CoV-2. One patient with plurimetastic colon cancer (liver, lung, peritoneum and brain), during hospitalization for a spontaneous pneumothorax developed symptoms of COVID-19 and, soon after, had an intestinal perforation for which the patient underwent a right hemicolectomy. The remaining patient affected by acute myeloid leukemia developed COVID-19 symptoms while hospitalized for chemotherapy; a week later, the patient underwent an appendectomy and colonic resection for peritonitis secondary to an inflammatory mass englobing the appendix and the sigmoid colon. All the patients died during hospitalization. SARS-CoV-2 RNA was detected in 2 cases out of 6 (33.3%). In both cases, the virus RNA was positive in the colonic tissue of the 2 patients with active neoplastic disease. In both cases, the viral load was very low. In one intestinal specimen, the viral load was less than the limit of quantification of the test (< 25 copies/microg RNA) and, in the second, it was 29 copies/microgr RNA. Histological examination of the apparently healthy tissue of all the cases showed normal glandular architecture, no enterocyte damage, a slight expansion of the lamina propria by edema and inflammatory lymphoplasmacytic infiltration in the lamina propria varying from mild to moderate. However, in the two cases positive for viral RNA, histological examination also pointed out abundant macrophages, granulocytes and plasma cells infiltrating the muscular layer, and adipose tissue, focal vasculitis and some macrophages in the vascular lumen (Figures 2A and B).
In the present study, SARS-CoV-2 RNA was identified in only two cases out of the 59 intestinal samples (3.4%). In both cases the viral load was very low; the intestinal samples consisted of colonic tissue of patients with active neoplastic disease (a patient with acute myeloid leukemia who was receiving immunosuppressive therapy and a patient with metastatic colon cancer), and there was a nosocomial transmission of SARS-CoV-2. Interestingly, in the biopsy group, two patients had previously undergone surgery for cancer (prostate and cervix) and in both cases, no viral RNA was identified in the rectal samples. Several studies have suggested that people with neoplastic disease are more likely to contract COVID-19 and to develop more severe disease or die from it than the general population. A recent systematic review on COVID-19 patients with active malignancy, defined as current malignant disease or treatment for malignancy within the last 12 mo, showed that cancer constitutes a co-morbidity in 2.6% [95% confidence interval (CI): 1.8%-3.5%, I2: 92.0%] of hospitalized COVID-19 patients and that the pooled in-hospital mortality risk was 14.1% (95%CI: 9.1%-19.8%, I2: 52.3%)[20]. Nahshon et al[21] suggested a severe clinical course of 50.6% and a mortality rate of 34.5% in COVID-19 patients with cancer. The worst COVID-19 outcomes include acute respiratory distress syndrome, septic shock, acute myocardial ischemia and death[22]. These severe events occurred more frequently in patients with stage IV cancer as compared to those with non-stage IV cancer (70.0% vs 44.4%, respectively) and if the last antitumor treatment was within 14 d (hazard ratio = 4.079, 95%CI: 1.086-15.322, P = 0.037)[22]. The pathogenesis of severe COVID-19 in cancer patients may be due to the aggravation of inflammatory cytokine storms, the imbalance of immune responses, and multiple organ damage[23]. In a multicenter retrospective cohort study on 232 patients with cancer and 519 statistically matched patients without cancer, Tian et al[23] identified elevated levels of interleukin (IL)-6, tumor necrosis factor α (TNF-α) and N-terminal pro-B-Type natriuretic peptide (NT-proBNP) and a reduced level of CD4+T cells and albumin-globulin ratio as risk factors of COVID-19 severity in patients with cancer. Similarly, in a retrospective cohort study which included 2052 patients hospitalized with COVID-19 (cancer, n = 93; non-cancer, n = 1959), Cai et al[24] reported that immune dysregulation was an important feature in cancer patients with COVID-19, which might account for their poorer prognosis; they found that COVID-19 patients with cancer had ongoing and significantly elevated inflammatory factors and cytokines (C-reactive protein, procalcitonin, IL-2 receptor, IL-6, IL-8) as well as a decreased number of immune cells (CD8 + T cells, CD4 + T cells, B cells, nature killer cells, T-helper and T- suppressor cells) than those without cancer. In patients with weakened immune systems, SARS-CoV-2 could infect vascular epithelial cells and organs, such as the lungs, heart, kidneys, liver, and intestine, expressing high levels of ACE2[25]. Autopsy data have reported viral infection in several organs, indicating hematogenic spread of the virus[26,27]. Moreover, serum SARS-CoV-2 nucleic acid (RNAemia) was associated with COVID-19 severity [odds ratio (OR) = 5.43, 95%CI: 3.46-8.53], increased risk of multiorgan failure (OR = 7.33, 95%CI: 2.46-21.88) and mortality (OR = 11.07, 95%CI: 5.60-22.88)[28]. Notably, viral RNA was undetectable in any of 53 rectal biopsies performed on the 30 patients hospitalized for moderate COVID-19 (biopsy group). The inability of the authors to detect viral RNA in the rectal samples contrasted with some data which have identified SARS-CoV-2 in intestinal samples[8,9]. Xiao et al[8] evaluated the viral nucleocapsid protein in the GI tissues of a COVID-19 patient who developed severe respiratory distress and an upper GI bleed. At endoscopy, they observed mucosal damage in the esophagus and found viral nucleocapsid protein in the cytoplasm of gastric, duodenal, and rectal glandular epithelial cells with immunofluorescent staining[10]. Lin et al[9] detected SARS-CoV-2 RNA at endoscopy in esophageal, gastric, duodenal, and rectal specimens in 2 out of 6 COVID-19 patients having GI symptoms. In this study, the presence of viral RNA in the GI tissue was also associated with severe disease; in fact, SARS-CoV-2 RNA was found in the samples of 2 patients with severe disease but not in those of 4 patients with non-severe disease[9]. Therefore, although the authors’ failure to detect SARS-CoV-2 in the rectal samples could have been due to the mild disease course of the cases selected, the possibility that the viral load was below the detection limit of their RT-PCR assay cannot be excluded. This could justify the presence of a mild to moderate inflammatory infiltrate in the lamina propria in all the rectal samples at histological examination. In fact, plasma cells, lymphocytes, and granulocytes can migrate to the extravascular space to reach the possibly infected tissues. These findings are in line with the low number of endoscopic and histological examinations of intestinal samples of COVID-19 patients which showed inflammatory infiltration in the lamina propria. Endoscopy and biopsy samples of the esophagus, stomach, duodenum and rectum were taken from a 78-year-old patient with COVID-19 who showed symptoms of upper GI bleeding. Numerous infiltrating plasma cells and lymphocytes with interstitial edema were found in the lamina propria of the stomach, duodenum and rectum of this patient[8]. In a surgical rectal specimen obtained during the incubation period in a COVID-19 patient with rectal adenocarcinoma, histological examination showed prominent lymphocytes and macrophages infiltrating the lamina propria without significant mucosal damage. T lymphocytes and macrophages were found to be more numerous than B lymphocytes in the lamina propria[29].
Six patients out of the 14 cases examined (42.9%) tested positive for SARS-CoV-2 RNA in the stool. Multiple studies have reported the positive detection of viral nucleic acids in the fecal samples of COVID-19 patients, finding rates varying from 15.3% to 66.7%[6,14]. In a meta-analysis, the authors showed that viral RNA may be present in the feces in 48.1% of patients[30]. The mechanism of diarrhea in patients with COVID-19 is still largely unknown. Various etiopathogenetic hypotheses have been advanced to explain the occurrence of diarrhea in COVID-19 patients including alterations in gut microbiota, osmotic diarrhea due to malabsorption or inflammation, release of virulent proteins or toxins, and viral-induced intestinal fluid and electrolyte secretion by activation of the enteric nervous system[31,32].
Currently, the exact mechanism of intestinal involvement in COVID-19 is not yet well understood. Intestinal epithelial cells could be primarily infected by SARS-CoV-2 via the oral-fecal route or SARS-CoV-2 may invade the enteric cells after respiratory infection via lympho hematogenic spread. As COVID-19 is also associated with the involvement of different organs and systems, such as the liver, kidneys, heart, blood, and nervous system; it has been hypothesized that in severe COVID-19 patients and in those with compromised immunity, SARS-CoV-2 has not been successfully eradicated and can spread from the lungs to target organs, such as the intestine[33]. Although the present data are unable to support the observations suggesting that enteric infection can occur in COVID-19 patients, in the two cases positive for viral RNA, histological examination showed an inflammatory infiltrate characterized by the presence of macrophages, granulocytes, plasma cells, and focal vasculitis. Thus, it could be hypothesized that in these cases, there were both a direct viral infection and immune hyperactivation. Hyperactivation of the immune system in response to infection can cause severe complications and organ damage. The host immune response is thought to play a vital role in the pathogenesis of COVID-19[34-36]. The present study has several limitations. The biopsies were performed only in the rectum of patients with moderate COVID-19. On the other hand, due to the high risk of viral spreading during endoscopic procedures, it is difficult to obtain samples from the gastric, intestinal and colic mucosa in patients who do not complain of GI symptoms, especially in cases of severe illness. Moreover, it is not possible to exclude that the two positive colon samples could be contaminated by SARS-CoV-2 positive stool or blood. Another limitation of the present study was its observational nature which made it difficult to identify the causes of the observed phenomena. Nonetheless, the detection of the viral RNA observed and the inflammatory cell infiltration to the colonic tissue of patients with active cancer could serve as hypothesis generators, leading to the analyzing of more comprehensive autopsy or surgical specimens in order to assess the potential link between SARS-CoV-2 and enteric infections in this population. The strengths of this study include the use of RT-PCR which is the gold standard for detecting SARS-CoV-2 infection. Moreover, the rectal biopsies were performed in two different sites and stored in both RNA preservation medium and in 10% buffered formalin to reduce the risk of false negatives.
SARS-CoV-2 RNA was found in only a small percentage of the intestinal samples analyzed (3.4%). Nevertheless, more comprehensive autopsy or surgical specimens are needed to provide histological evidence of intestinal infection.
Although some observations provide evidence for intestinal infection of severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2), the mechanisms leading to this infection are not known.
The detection of viral RNA in gastrointestinal (GI) tissue samples has not been adequately investigated and results are conflicting. More GI tissue samples, comprehensive autopsy and surgical specimens are needed to provide histological evidence of intestinal infection.
Intestinal mucosal samples from mild-moderate coronavirus disease 2019 (COVID-19) patients were analyzed with the primary objective of detecting SARS-CoV-2 RNA and evaluating histological features.
This is a monocentric trial in which real time reverse transcriptase-polymerase chain reaction and histological features were used to detect SARS-CoV-2 RNA in intestinal mucosal samples. The study population was composed of two groups of patients hospitalized for COVID-19. In the first group (biopsy group), the patients were eligible for inclusion if they had mild to moderate disease and if they agreed to have a rectal biopsy regardless of the presence or absence of GI symptoms; in the second group (surgical specimen group), patients were eligible for inclusion if they underwent intestinal resection during index hospitalization. The data obtained in this study are valuable because rectal biopsies were carried out on 30 patients who did not need the procedure to frame their disease status. The study therefore provides data that are not only more numerous but also qualitatively different from those available up to now.
Overall, we analyzed 53 rectal biopsies and 6 surgical specimens. Viral RNA was not detectable in any of the rectal biopsies performed (0/53). Histological examination showed no enterocyte damage, but slight edema of the lamina propria with mild inflammatory lymphoplasmacytic infiltration. Viral RNA was detected in 2 surgical specimens of the 6 examined, both of which were from patients with active neoplastic disease. Histological examination also pointed out abundant macrophages, granulocytes and plasma cells infiltrating the muscular layer and adipose tissue, and focal vasculitis.
Mild-moderate COVID-19 may not be associated with rectal infection by the virus. Although the present data are unable to support the observations suggesting that enteric infection can occur in COVID-19 patients, the detection of the viral RNA observed and the inflammatory cell infiltration to the colonic tissue of patients with active cancer could serve as hypothesis generators, leading to the analyzing more comprehensive autopsy or surgical specimens in order to assess the potential link between SARS-CoV-2 and enteric infections in this population.
Does intestinal infection lead to increased expression of inflammatory cytokines in the intestine and/or serum? Since the two positive samples were both from patients with active cancer, could a weakened immune system, induced by the neoplastic disease, increase the risk of the intestinal infection of SARS-CoV-2?
Provenance and peer review: Unsolicited article; Externally peer reviewed.
Peer-review model: Single blind
Specialty type: Gastroenterology and hepatology
Country/Territory of origin: Italy
Peer-review report’s scientific quality classification
Grade A (Excellent): A
Grade B (Very good): B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Ait Addi R, Morocco; Nooripour R, Iran S-Editor: Wang JJ L-Editor: Filipodia P-Editor: Wang JJ
| 1. | Wiersinga WJ, Rhodes A, Cheng AC, Peacock SJ, Prescott HC. Pathophysiology, Transmission, Diagnosis, and Treatment of Coronavirus Disease 2019 (COVID-19): A Review. JAMA. 2020;324:782-793. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2691] [Cited by in RCA: 3176] [Article Influence: 635.2] [Reference Citation Analysis (0)] |
| 2. | Cuicchi D, Lazzarotto T, Poggioli G. Fecal-oral transmission of SARS-CoV-2: review of laboratory-confirmed virus in gastrointestinal system. Int J Colorectal Dis. 2021;36:437-444. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 36] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 3. | Trypsteen W, Van Cleemput J, Snippenberg WV, Gerlo S, Vandekerckhove L. On the whereabouts of SARS-CoV-2 in the human body: A systematic review. PLoS Pathog. 2020;16:e1009037. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 167] [Cited by in RCA: 144] [Article Influence: 28.8] [Reference Citation Analysis (0)] |
| 4. | Tariq R, Saha S, Furqan F, Hassett L, Pardi D, Khanna S. Prevalence and Mortality of COVID-19 Patients With Gastrointestinal Symptoms: A Systematic Review and Meta-analysis. Mayo Clin Proc. 2020;95:1632-1648. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 95] [Cited by in RCA: 81] [Article Influence: 16.2] [Reference Citation Analysis (0)] |
| 5. | Mao R, Qiu Y, He JS, Tan JY, Li XH, Liang J, Shen J, Zhu LR, Chen Y, Iacucci M, Ng SC, Ghosh S, Chen MH. Manifestations and prognosis of gastrointestinal and liver involvement in patients with COVID-19: a systematic review and meta-analysis. Lancet Gastroenterol Hepatol. 2020;5: 667-678. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 741] [Cited by in RCA: 752] [Article Influence: 150.4] [Reference Citation Analysis (0)] |
| 6. | Pan L, Mu M, Yang P, Sun Y, Wang R, Yan J, Li P, Hu B, Wang J, Hu C, Jin Y, Niu X, Ping R, Du Y, Li T, Xu G, Hu Q, Tu L. Clinical Characteristics of COVID-19 Patients With Digestive Symptoms in Hubei, China: A Descriptive, Cross-Sectional, Multicenter Study. Am J Gastroenterol. 2020;115:766-773. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1160] [Cited by in RCA: 1204] [Article Influence: 240.8] [Reference Citation Analysis (0)] |
| 7. | Wu Y, Guo C, Tang L, Hong Z, Zhou J, Dong X, Yin H, Xiao Q, Tang Y, Qu X, Kuang L, Fang X, Mishra N, Lu J, Shan H, Jiang G, Huang X. Prolonged presence of SARS-CoV-2 viral RNA in faecal samples. Lancet Gastroenterol Hepatol. 2020;5:434-435. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1050] [Cited by in RCA: 1150] [Article Influence: 230.0] [Reference Citation Analysis (0)] |
| 8. | Xiao F, Tang M, Zheng X, Liu Y, Li X, Shan H. Evidence for Gastrointestinal Infection of SARS-CoV-2. Gastroenterology. 2020;158:1831-1833.e3. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1985] [Cited by in RCA: 1993] [Article Influence: 398.6] [Reference Citation Analysis (1)] |
| 9. | Lin L, Jiang X, Zhang Z, Huang S, Fang Z, Gu Z, Gao L, Shi H, Mai L, Liu Y, Lin X, Lai R, Yan Z, Li X, Shan H. Gastrointestinal symptoms of 95 cases with SARS-CoV-2 infection. Gut. 2020;69:997-1001. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 630] [Cited by in RCA: 657] [Article Influence: 131.4] [Reference Citation Analysis (0)] |
| 10. | Zhang Y, Chen C, Zhu S, Shu C, Wang D, Song J, Song Y, Zhen W, Feng Z, Wu G, Xu J, Xu W. Isolation of 2019-nCoV from a Stool Specimen of a Laboratory-Confirmed Case of the Coronavirus Disease 2019 (COVID-19). China CDC Wkly. 2020;2:123-124. [PubMed] |
| 11. | Wölfel R, Corman VM, Guggemos W, Seilmaier M, Zange S, Müller MA, Niemeyer D, Jones TC, Vollmar P, Rothe C, Hoelscher M, Bleicker T, Brünink S, Schneider J, Ehmann R, Zwirglmaier K, Drosten C, Wendtner C. Virological assessment of hospitalized patients with COVID-2019. Nature. 2020;581:465-469. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4682] [Cited by in RCA: 4809] [Article Influence: 961.8] [Reference Citation Analysis (0)] |
| 12. | Zhang H, Kang Z, Gong H, Xu D, Wang J, Li Z, Cui X, Xiao J, Meng T, Zhou W, Liu J, Xu H. The digestive system is a potential route of 2019-nCov infection: a bioinformatics 2 analysis based on single-cell transcriptomes. 2020 Preprint. Available from: bioRxiv: 2020.01.30.927806. [DOI] [Full Text] |
| 13. | Zhou J, Li C, Liu X, Chiu MC, Zhao X, Wang D, Wei Y, Lee A, Zhang AJ, Chu H, Cai JP, Yip CC, Chan IH, Wong KK, Tsang OT, Chan KH, Chan JF, To KK, Chen H, Yuen KY. Infection of bat and human intestinal organoids by SARS-CoV-2. Nat Med. 2020;26:1077-1083. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 326] [Cited by in RCA: 430] [Article Influence: 86.0] [Reference Citation Analysis (0)] |
| 14. | Lamers MM, Beumer J, van der Vaart J, Knoops K, Puschhof J, Breugem TI, Ravelli RBG, Paul van Schayck J, Mykytyn AZ, Duimel HQ, van Donselaar E, Riesebosch S, Kuijpers HJH, Schipper D, van de Wetering WJ, de Graaf M, Koopmans M, Cuppen E, Peters PJ, Haagmans BL, Clevers H. SARS-CoV-2 productively infects human gut enterocytes. Science. 2020;369:50-54. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1245] [Cited by in RCA: 1310] [Article Influence: 262.0] [Reference Citation Analysis (0)] |
| 15. | Chen L, Lou J, Bai Y, Wang M. COVID-19 Disease With Positive Fecal and Negative Pharyngeal and Sputum Viral Tests. Am J Gastroenterol. 2020;115:790. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 97] [Cited by in RCA: 105] [Article Influence: 21.0] [Reference Citation Analysis (0)] |
| 16. | Wang W, Xu Y, Gao R, Lu R, Han K, Wu G, Tan W. Detection of SARS-CoV-2 in Different Types of Clinical Specimens. JAMA. 2020;323:1843-1844. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1894] [Cited by in RCA: 2658] [Article Influence: 531.6] [Reference Citation Analysis (0)] |
| 17. | Zhang J, Wang S, Xue Y. Fecal specimen diagnosis 2019 novel coronavirus-infected pneumonia. J Med Virol. 2020;92:680-682. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 278] [Cited by in RCA: 301] [Article Influence: 60.2] [Reference Citation Analysis (0)] |
| 18. | Han C, Duan C, Zhang S, Spiegel B, Shi H, Wang W, Zhang L, Lin R, Liu J, Ding Z, Hou X. Digestive Symptoms in COVID-19 Patients With Mild Disease Severity: Clinical Presentation, Stool Viral RNA Testing, and Outcomes. Am J Gastroenterol. 2020;115:916-923. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 413] [Cited by in RCA: 383] [Article Influence: 76.6] [Reference Citation Analysis (0)] |
| 19. | Wu Z, McGoogan JM. Characteristics of and Important Lessons From the Coronavirus Disease 2019 (COVID-19) Outbreak in China: Summary of a Report of 72 314 Cases From the Chinese Center for Disease Control and Prevention. JAMA. 2020;323:1239-1242. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11409] [Cited by in RCA: 11504] [Article Influence: 2300.8] [Reference Citation Analysis (0)] |
| 20. | Zarifkar P, Kamath A, Robinson C, Morgulchik N, Shah SFH, Cheng TKM, Dominic C, Fehintola AO, Bhalla G, Ahillan T, Mourgue d'Algue L, Lee J, Pareek A, Carey M, Hughes DJ, Miller M, Woodcock VK, Shrotri M. Clinical Characteristics and Outcomes in Patients with COVID-19 and Cancer: a Systematic Review and Meta-analysis. Clin Oncol (R Coll Radiol). 2021;33:e180-e191. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 56] [Cited by in RCA: 55] [Article Influence: 13.8] [Reference Citation Analysis (0)] |
| 21. | Nahshon C, Segev Y, Schmidt M, Bar-Noy T, Ostrovsky L, Lavie O. Outcomes of diagnosed COVID-19 cancer patients: concerning results of a systematic review. J Chemother. 2021;33:528-538. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 22. | Zhang L, Zhu F, Xie L, Wang C, Wang J, Chen R, Jia P, Guan HQ, Peng L, Chen Y, Peng P, Zhang P, Chu Q, Shen Q, Wang Y, Xu SY, Zhao JP, Zhou M. Clinical characteristics of COVID-19-infected cancer patients: a retrospective case study in three hospitals within Wuhan, China. Ann Oncol. 2020;31:894-901. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 819] [Cited by in RCA: 1016] [Article Influence: 203.2] [Reference Citation Analysis (0)] |
| 23. | Tian J, Yuan X, Xiao J, Zhong Q, Yang C, Liu B, Cai Y, Lu Z, Wang J, Wang Y, Liu S, Cheng B, Zhang M, Wang L, Niu S, Yao Z, Deng X, Zhou F, Wei W, Li Q, Chen X, Chen W, Yang Q, Wu S, Fan J, Shu B, Hu Z, Wang S, Yang XP, Liu W, Miao X, Wang Z. Clinical characteristics and risk factors associated with COVID-19 disease severity in patients with cancer in Wuhan, China: a multicentre, retrospective, cohort study. Lancet Oncol. 2020;21:893-903. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 388] [Cited by in RCA: 389] [Article Influence: 77.8] [Reference Citation Analysis (0)] |
| 24. | Cai G, Gao Y, Zeng S, Yu Y, Liu X, Liu D, Wang Y, Yu R, Desai A, Li C, Gao Q. Immunological alternation in COVID-19 patients with cancer and its implications on mortality. Oncoimmunology. 2021;10:1854424. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 16] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 25. | Qin C, Zhou L, Hu Z, Zhang S, Yang S, Tao Y, Xie C, Ma K, Shang K, Wang W, Tian DS. Dysregulation of Immune Response in Patients With Coronavirus 2019 (COVID-19) in Wuhan, China. Clin Infect Dis. 2020;71:762-768. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2495] [Cited by in RCA: 3336] [Article Influence: 667.2] [Reference Citation Analysis (0)] |
| 26. | Puelles VG, Lütgehetmann M, Lindenmeyer MT, Sperhake JP, Wong MN, Allweiss L, Chilla S, Heinemann A, Wanner N, Liu S, Braun F, Lu S, Pfefferle S, Schröder AS, Edler C, Gross O, Glatzel M, Wichmann D, Wiech T, Kluge S, Pueschel K, Aepfelbacher M, Huber TB. Multiorgan and Renal Tropism of SARS-CoV-2. N Engl J Med. 2020;383:590-592. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1254] [Cited by in RCA: 1423] [Article Influence: 284.6] [Reference Citation Analysis (0)] |
| 27. | Wichmann D, Sperhake JP, Lütgehetmann M, Steurer S, Edler C, Heinemann A, Heinrich F, Mushumba H, Kniep I, Schröder AS, Burdelski C, de Heer G, Nierhaus A, Frings D, Pfefferle S, Becker H, Bredereke-Wiedling H, de Weerth A, Paschen HR, Sheikhzadeh-Eggers S, Stang A, Schmiedel S, Bokemeyer C, Addo MM, Aepfelbacher M, Püschel K, Kluge S. Autopsy Findings and Venous Thromboembolism in Patients With COVID-19: A Prospective Cohort Study. Ann Intern Med. 2020;173:268-277. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1577] [Cited by in RCA: 1748] [Article Influence: 349.6] [Reference Citation Analysis (0)] |
| 28. | Tang K, Wu L, Luo Y, Gong B. Quantitative assessment of SARS-CoV-2 RNAemia and outcome in patients with coronavirus disease 2019. J Med Virol. 2021;93:3165-3175. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 37] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 29. | Qian Q, Fan L, Liu W, Li J, Yue J, Wang M, Ke X, Yin Y, Chen Q, Jiang C. Direct Evidence of Active SARS-CoV-2 Replication in the Intestine. Clin Infect Dis. 2021;73:361-366. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 123] [Cited by in RCA: 119] [Article Influence: 29.8] [Reference Citation Analysis (0)] |
| 30. | Cheung KS, Hung IFN, Chan PPY, Lung KC, Tso E, Liu R, Ng YY, Chu MY, Chung TWH, Tam AR, Yip CCY, Leung KH, Fung AY, Zhang RR, Lin Y, Cheng HM, Zhang AJX, To KKW, Chan KH, Yuen KY, Leung WK. Gastrointestinal Manifestations of SARS-CoV-2 Infection and Virus Load in Fecal Samples From a Hong Kong Cohort: Systematic Review and Meta-analysis. Gastroenterology. 2020;159:81-95. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1113] [Cited by in RCA: 1131] [Article Influence: 226.2] [Reference Citation Analysis (1)] |
| 31. | Guo M, Tao W, Flavell RA, Zhu S. Potential intestinal infection and faecal-oral transmission of SARS-CoV-2. Nat Rev Gastroenterol Hepatol. 2021;18:269-283. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 198] [Cited by in RCA: 213] [Article Influence: 53.3] [Reference Citation Analysis (0)] |
| 32. | Scaldaferri F, Ianiro G, Privitera G, Lopetuso LR, Vetrone LM, Petito V, Pugliese D, Neri M, Cammarota G, Ringel Y, Costamagna G, Gasbarrini A, Boskoski I, Armuzzi A. The Thrilling Journey of SARS-CoV-2 into the Intestine: From Pathogenesis to Future Clinical Implications. Inflamm Bowel Dis. 2020;26:1306-1314. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 27] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 33. | Gupta A, Madhavan MV, Sehgal K, Nair N, Mahajan S, Sehrawat TS, Bikdeli B, Ahluwalia N, Ausiello JC, Wan EY, Freedberg DE, Kirtane AJ, Parikh SA, Maurer MS, Nordvig AS, Accili D, Bathon JM, Mohan S, Bauer KA, Leon MB, Krumholz HM, Uriel N, Mehra MR, Elkind MSV, Stone GW, Schwartz A, Ho DD, Bilezikian JP, Landry DW. Extrapulmonary manifestations of COVID-19. Nat Med. 2020;26:1017-1032. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2419] [Cited by in RCA: 2046] [Article Influence: 409.2] [Reference Citation Analysis (2)] |
| 34. | Bhalerao A, Raut S, Noorani B, Mancuso S, Cucullo L. Molecular Mechanisms of Multi-Organ Failure in COVID-19 and Potential of Stem Cell Therapy. Cells. 2021;10. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 5] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 35. | Zaim S, Chong JH, Sankaranarayanan V, Harky A. COVID-19 and Multiorgan Response. Curr Probl Cardiol. 2020;45:100618. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 636] [Cited by in RCA: 641] [Article Influence: 128.2] [Reference Citation Analysis (0)] |
| 36. | Koçak Tufan Z, Kayaaslan B, Mer M. COVID-19 and Sepsis. Turk J Med Sci. 2021;51:3301-3311. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29] [Cited by in RCA: 56] [Article Influence: 14.0] [Reference Citation Analysis (0)] |