Copyright
©The Author(s) 2021.
World J Gastroenterol. Sep 14, 2021; 27(34): 5630-5665
Published online Sep 14, 2021. doi: 10.3748/wjg.v27.i34.5630
Published online Sep 14, 2021. doi: 10.3748/wjg.v27.i34.5630
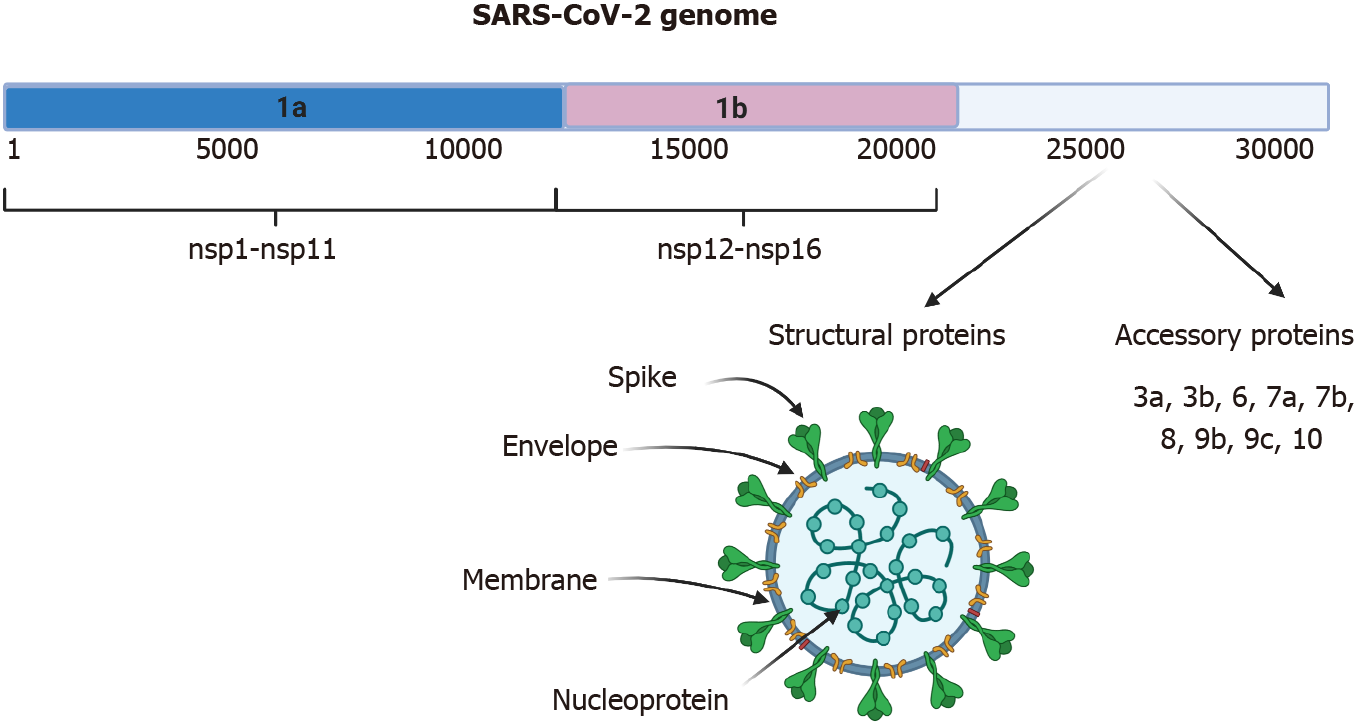
Figure 1 Genome and structure of severe acute respiratory syndrome-coronavirus-2.
The 30-kb severe acute respiratory syndrome-coronavirus-2 virus genome is shown. It possesses different open reading frames (ORFs). ORF 1a generates the nonstructural proteins (nsp) from nsp1-nsp11, ORF 1b generates proteins nsp12-nsp16. The spike, envelope, membrane, and nucleoprotein structural proteins and accessory proteins are generated from the 3’ region. Created with BioRender.com. SARS-CoV-2: Severe acute respiratory syndrome-coronavirus-2.
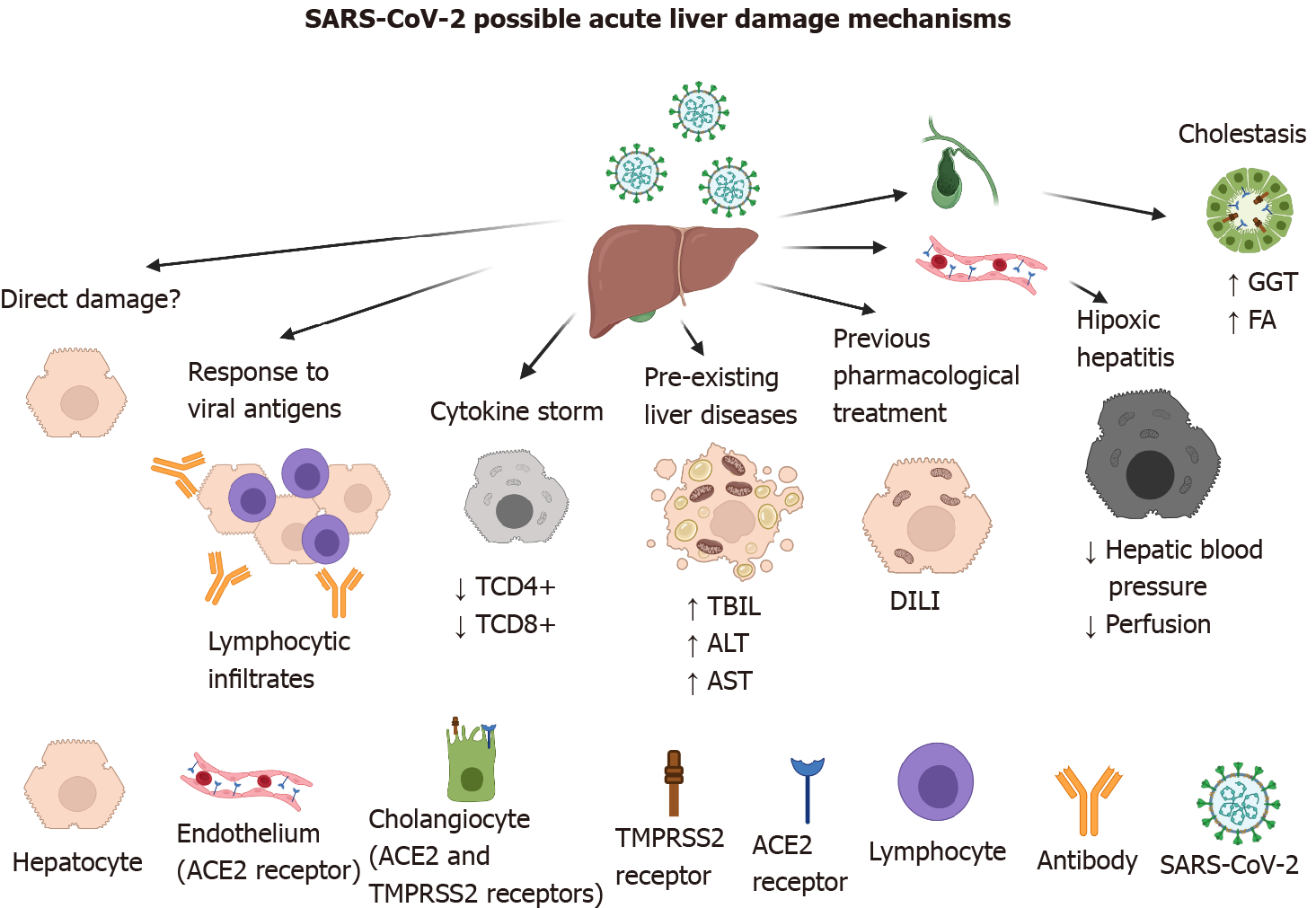
Figure 2 Possible mechanisms of acute liver damage by severe acute respiratory syndrome-coronavirus-2 infection.
Direct damage to the hepatocyte is does not occur because it does not possess the viral entry receptor. However, indirect damage-associated with the immune response against viral antigens, cytokine storm, pre-existing liver diseases such as hepatitis A, B, and C viruses infection, and treatment with hepatotoxic drugs like acetaminophen, remdesivir, lopinavir, etc. has been characterized. In severely affected patients, oxygen deficiency can give rise to hypoxic hepatitis. Severe acute respiratory syndrome-coronavirus-2 damage to cholangiocytes has been documented, because they both possess the angiotensin-converting enzyme 2 and transmembrane protease serine 2 receptors. Created with BioRender.com. ACE2: Angiotensin-converting enzyme 2; ALT: Alanine aminotransferase; AST: Aspartate aminotransferase; DILI: Drug-induced liver injury; FA: Alkaline phosphatase; GGT: Gamma-glutamyl transferase;SARS-CoV-2: Severe acute respiratory syndrome-coronavirus-2; TBIL: Total bilirubin; TMPRSS2: Transmembrane protease serine 2.
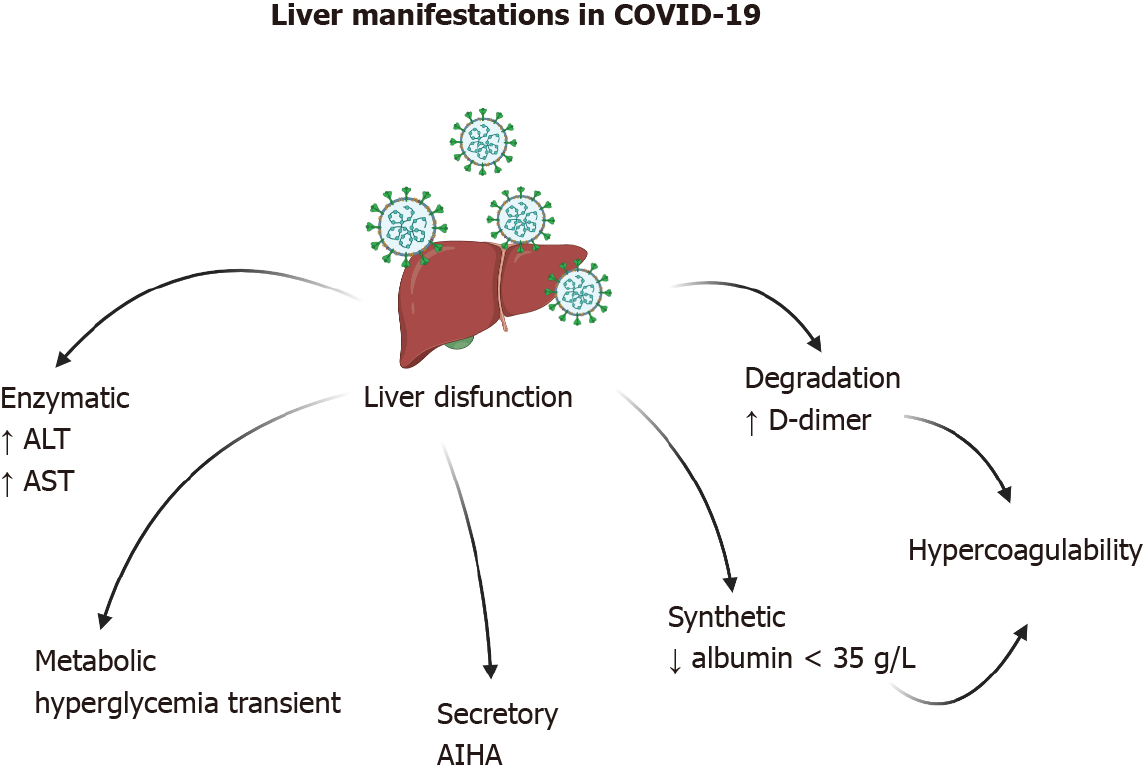
Figure 3 Liver manifestations in coronavirus disease 2019 are quite varied.
Increased transaminases and hypercoagulability have been observed in the majority of patients. In rare cases, autoimmune hemolytic anemia has been observed. Created with BioRender.com. AIHA: autoimmune hemolytic anemia; ALT: Alanine aminotransferase; AST: Aspartate aminotransferase; COVID-19: Coronavirus disease 2019.
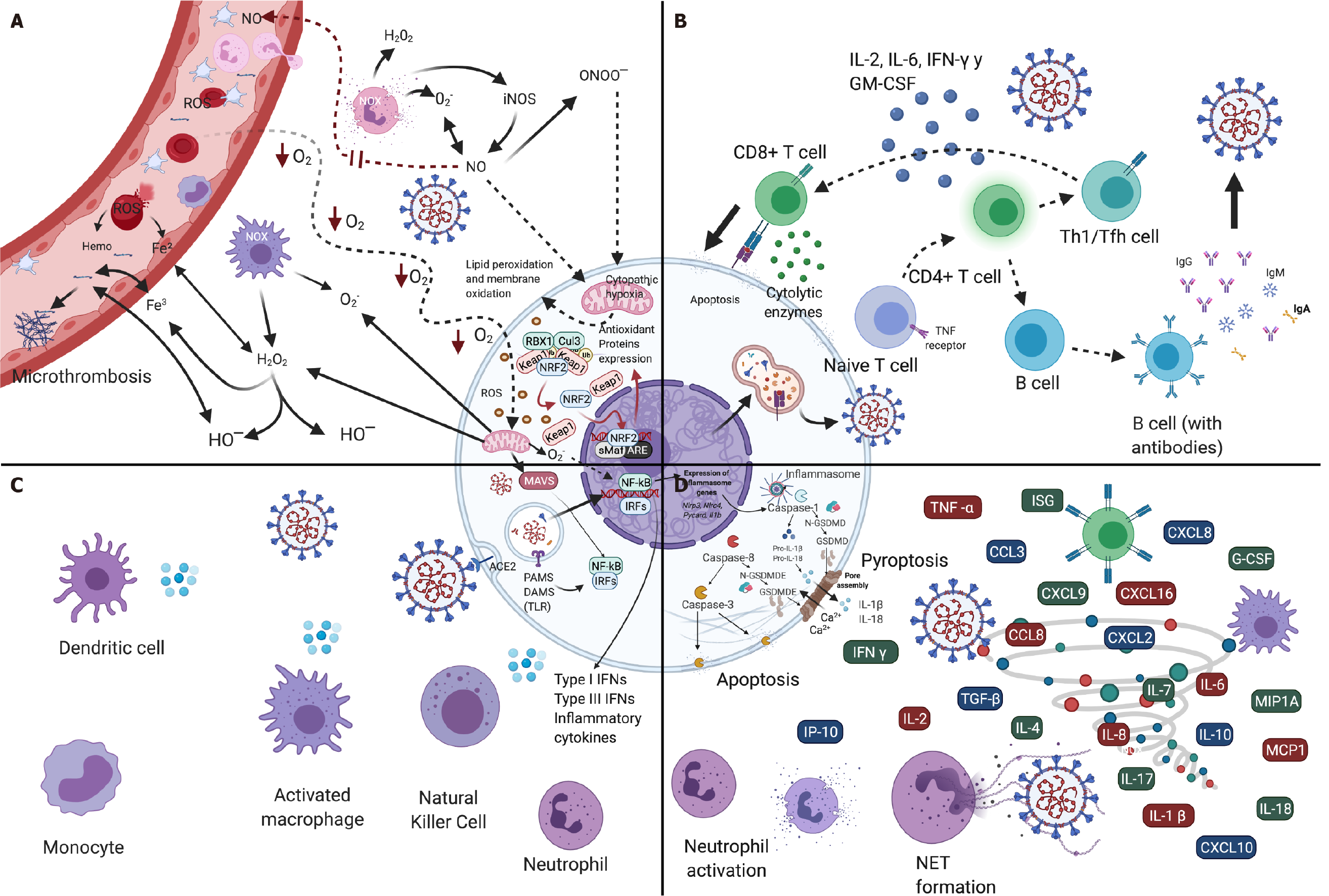
Figure 4 Immune response and oxidative stress in severe acute respiratory syndrome-coronavirus-2 infection.
A: Oxidative stress; B: Adaptive immune response; C: Innate immune response; D: Cytokine storm. Created with BioRender.com.
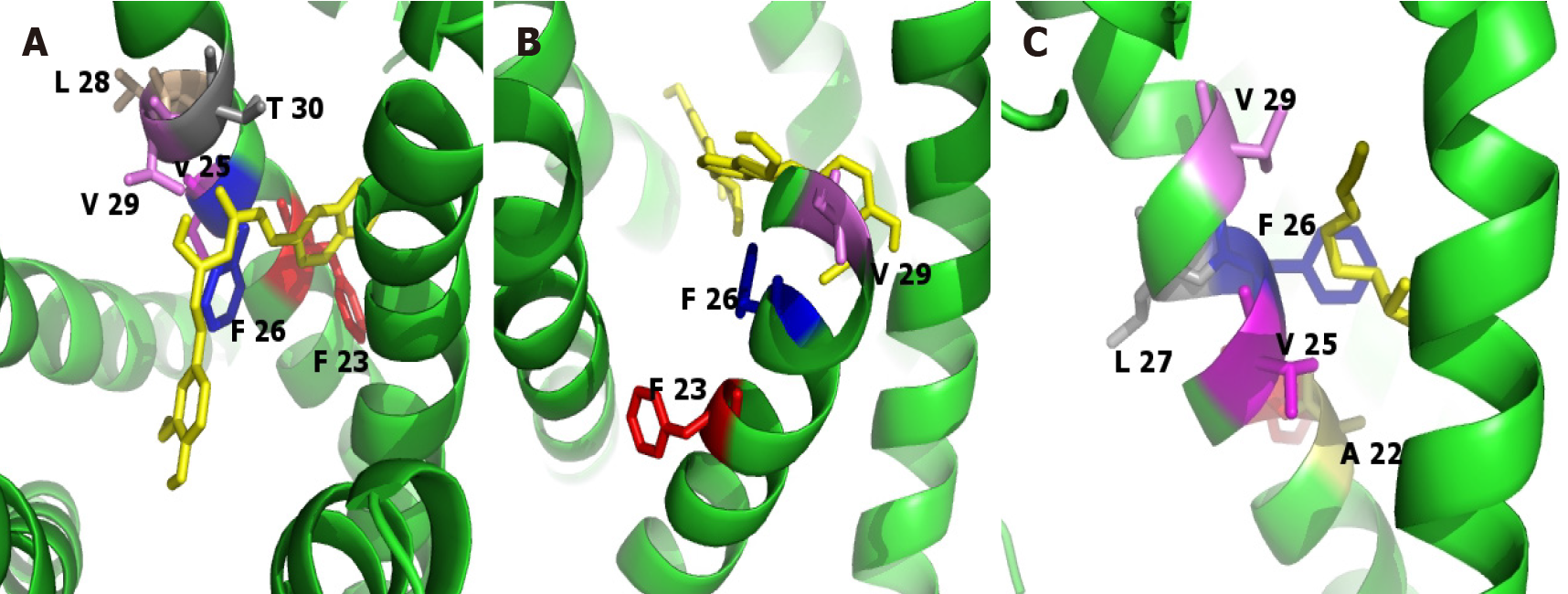
Figure 5 Graphic representation of the binding poses of the ligands on the envelope protein of severe acute respiratory syndrome-coronavirus-2.
Images were carried out using Pymol. Homo-pentamer of the E protein of severe acute respiratory syndrome-coronavirus-2 is shown in green, and the ligand is shown in yellow. Surrounded residues are shown in different colors and labeled as indicated. A: Curcumine; B: Silybin; C: Sulforaphane.
- Citation: Vargas-Mendoza N, García-Machorro J, Angeles-Valencia M, Martínez-Archundia M, Madrigal-Santillán EO, Morales-González Á, Anguiano-Robledo L, Morales-González JA. Liver disorders in COVID-19, nutritional approaches and the use of phytochemicals. World J Gastroenterol 2021; 27(34): 5630-5665
- URL: https://www.wjgnet.com/1007-9327/full/v27/i34/5630.htm
- DOI: https://dx.doi.org/10.3748/wjg.v27.i34.5630