Copyright
©2011 Baishideng Publishing Group Co.
World J Gastroenterol. Feb 14, 2011; 17(6): 681-690
Published online Feb 14, 2011. doi: 10.3748/wjg.v17.i6.681
Published online Feb 14, 2011. doi: 10.3748/wjg.v17.i6.681
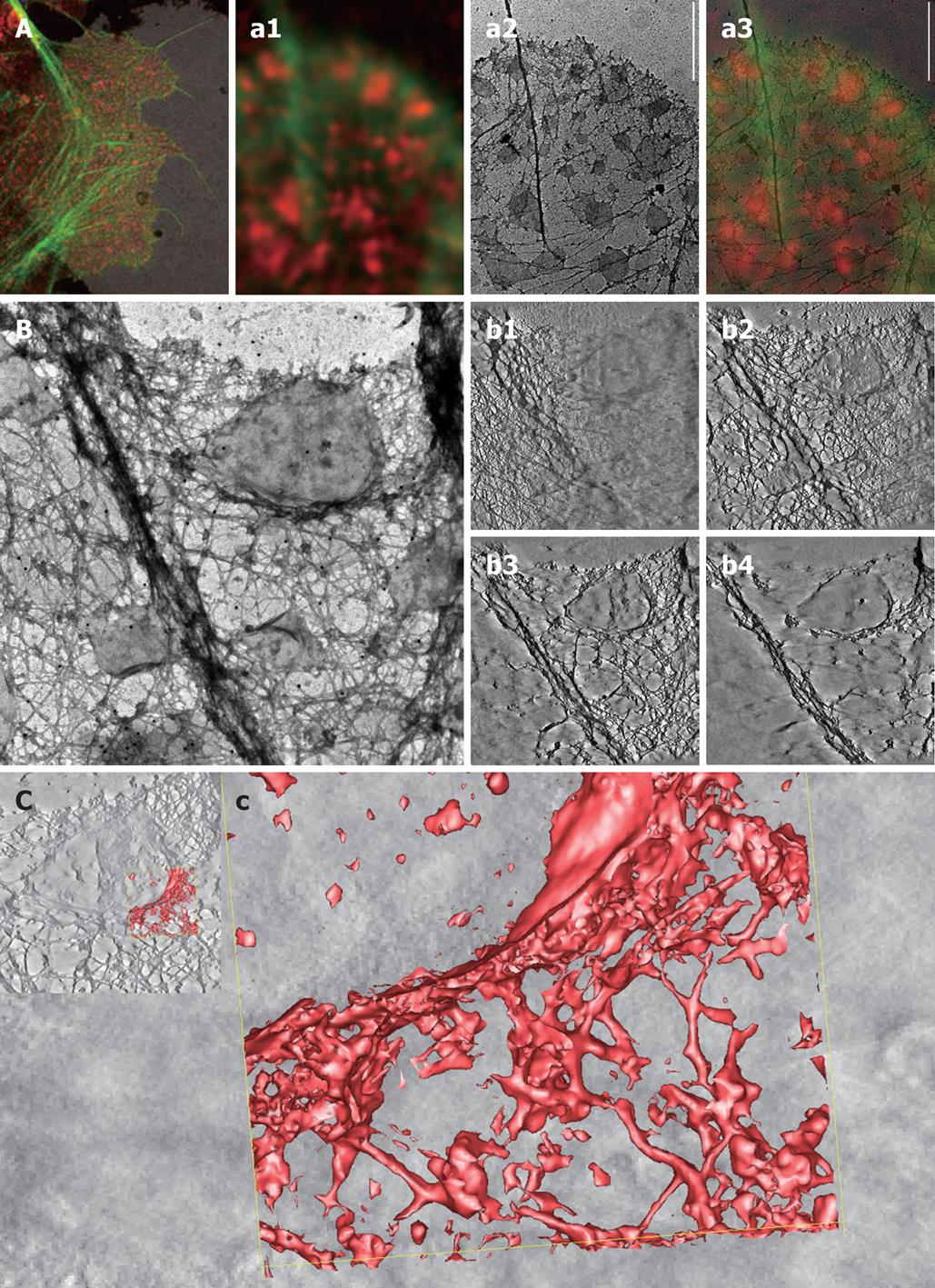
Figure 1 High-resolution imaging of membrane domains in human Caco-2 colorectal cancer cells.
These data come from high-resolution correlative fluorescence and transmission electron microscopy (CFEM) studies on whole mounts. Human colorectal cancer (CRC) cells (Caco-2) were cultured on formvar-coated nickel grids and then treated with Triton X-100 in cytoskeleton stabilization buffer, leaving detergent-resistant membranes behind[37]. These membrane fractions were labeled with the membrane raft marker GM1 CTxB-594 (red) and the actin cytoskeleton was stained with phalloidin-488 (green)[48]. A: Low-magnification overview of the peripheral cytoskeleton reveals actin-rich lamellipodia and associated filopodial extensions. Note the numerous GM1-positive lipid rafts interspersed throughout the extracted cytoplasm. The corresponding high-magnification CFEM analysis (a1-a3) of one of these lamellipodium-filopodial regions reveals the complex architectural nature of the leading edge, showing an intricate cytoskeleton-rich matrix and the close structural relationship with the lipid rafts (a1). Subsequent TEM analysis of exactly the same region not only allowed us to determine the exact size and shape of lipid rafts, but also provided an accurate idea of whether the rafts were located on top or beneath the lamellipodium: i.e. apical or basal (a2). The corresponding merged information (a3) clearly shows the additional value of applying different imaging techniques on the same cell: the electron-microscope data reveals small detergent-resistant membrane islands that could not be resolved by advanced confocal microscopy. Scale bar, 2 μm; For the electron tomographic analysis of a detergent-resistant membrane island and the surrounding cytoskeletal matrix (show in B and C), whole mounts of Caco-2 cells were prepared as outlined under A. Then we performed transmission electron tomography imaging at 1.5º incremental steps under dual-axis tilting (B) and then carried out subsequent segmenting (b1-b4) of the XYZ-tilting series. The single-image slices obtained via the IMOD tomography software show the sample at different heights from bottom to top (b1-b4) spanning a height of about 300 nm; C: An area of the entire tomogram was next selected (c) and a three-dimensional model generated of the membrane raft-cytoskeleton interface (c), showing the close interaction of fine cytoskeletal fibers at the rim of the detergent-resistant membrane.
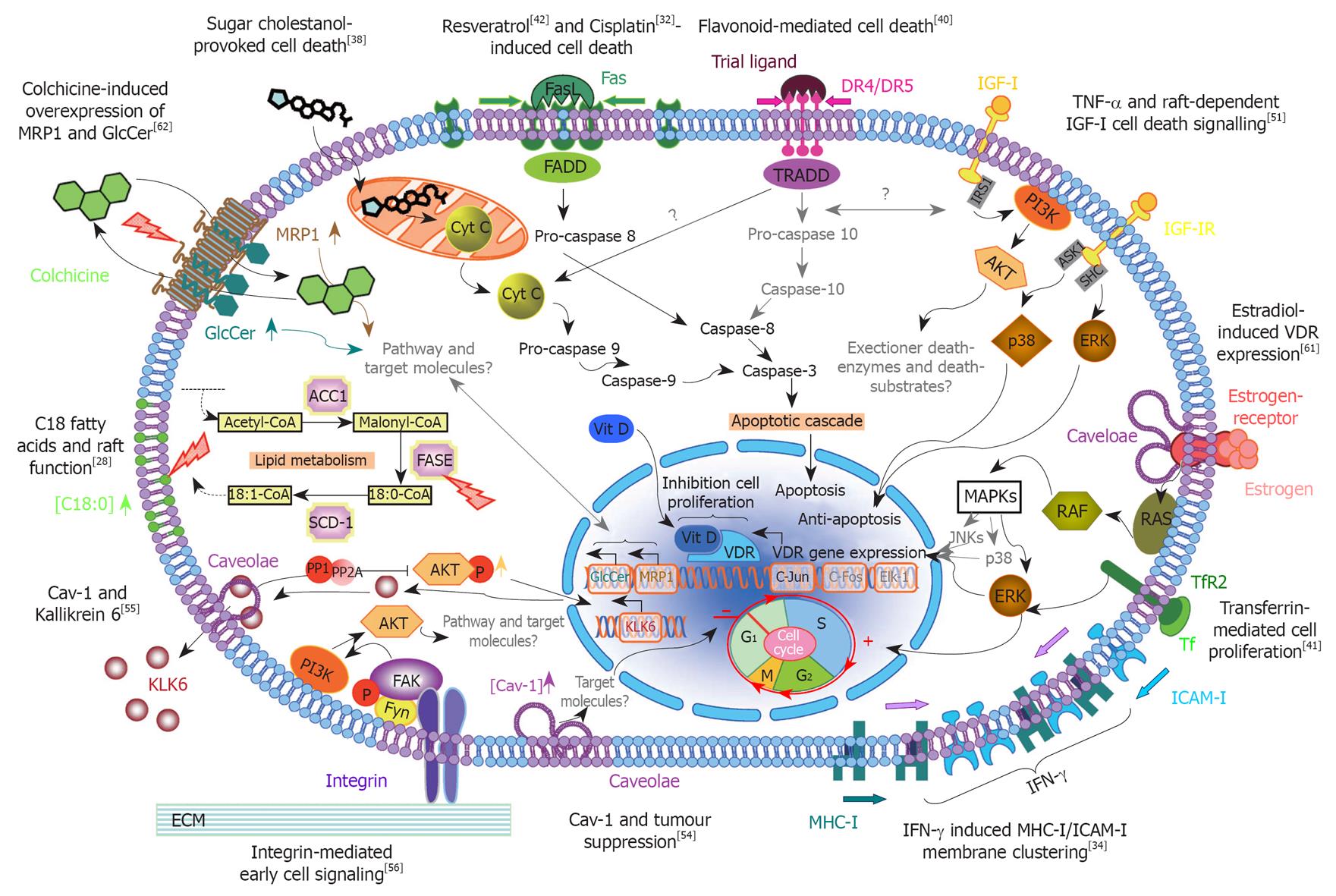
Figure 2 Scheme outlining the various membrane microdomain-mediated intracellular signaling pathways in colorectal cancer.
This diagram summarizes what has been reported to date in the literature about the different intracellular signaling pathways that are mediated by lipid rafts and the implications of these paths for the colon cancer cells’ state and fate. Briefly, the different observations of colorectal cancer (CRC) lipid rafts can be generally categorized under the following main topics of investigation: cell death-mediated mechanisms, caveolae in cancer cell growth and function, unique structure-function molecular associations, and intervention studies with bioactive compounds. Note that the text and connector arrows as shown in black are confirmed observations, whereas the gray denotes postulated signaling pathways and/or unknown molecular targets. The lipid bilayer of the cell membrane is depicted in light blue, membrane microdomains or lipid rafts in light purple, and the pear-shaped caveolae associated with these rafts in dark purple. For detailed descriptions of each of the individual cell signaling pathways, refer to the corresponding sections in this paper. The numbers in superscript refer directly to the original published papers. MRP: Multidrug-resistance protein; GlcCer: Glucosylceramide; FADD: Fas-associated protein with death domain; TRADD: Tumor necrosis factor receptor type 1-associated DEATH domain protein; PI3K: Phosphoinositide 3-kinase; AKT: Serine/threonine protein kinase; ERK: Extracellular signal-regulated kinase; MAPK: Mitogen-activated protein kinase; IRS1: Insulin receptor substrate 1; ASK1: Apoptosis signal-regulating kinase 1; SHC: Src homology 2 domain; TNF-α: Tumor necrosis factor-α; IGF-I: Insulin-like growth factor-I; VDR: Vitamin D receptor; Vit D: Vitamin D; RAF: Proto-oncogene serine/threonine-protein kinase; RAS: RAt sarcoma; TfR2: The second transferrin receptor; Tf: Transferrin; JNKs: c-Jun N-terminal kinases; ICAM-I: Intercellular adhesion molecule I; IFN-γ: Interferon-γ; MHC-I: Major histocompatibility complex I; FAK: Focal adhesion kinase; ECM: Extracellular matrix; FASE: Fatty acid synthase; SCD-1: Stearoyl-coenzyme A desaturase 1; ACC1: Acetyl-CoA carboxylase; Cav: Caveolin.
- Citation: Jahn KA, Su Y, Braet F. Multifaceted nature of membrane microdomains in colorectal cancer. World J Gastroenterol 2011; 17(6): 681-690
- URL: https://www.wjgnet.com/1007-9327/full/v17/i6/681.htm
- DOI: https://dx.doi.org/10.3748/wjg.v17.i6.681