Copyright
©The Author(s) 2025.
World J Clin Oncol. Apr 24, 2025; 16(4): 104182
Published online Apr 24, 2025. doi: 10.5306/wjco.v16.i4.104182
Published online Apr 24, 2025. doi: 10.5306/wjco.v16.i4.104182
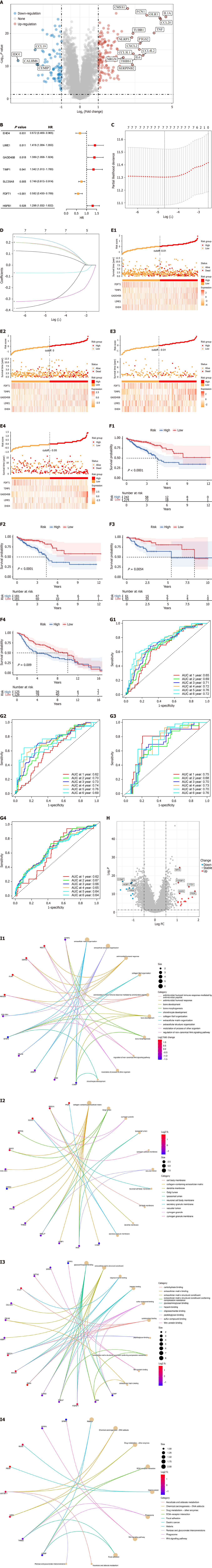
Figure 1 Establishment of prognostic model and enrichment analysis.
A: Volcano plot depicting differential gene expression; B: Forest plot highlighting notable mRNAs; C and D: Least absolute shrinkage and selection operator model selection and feature identification; E: Distribution of risk scores and heatmap of mRNA expression in The Cancer Genome Atlas colorectal cancer dataset. E1: Overall risk score distribution. E2: Training set. E3: Test set. E4: Gene Expression Omnibus (GEO) datasets; F: Kaplan-Meier survival analysis. F1: Total risk score. F2: Training set. F3: Test set. F4: GEO datasets; G: Time-dependent receiver operating characteristic curves. G1: Entire dataset. G2-G4: Training set, test set, and GEO datasets, respectively; H: Volcano plot of differential genes; I: Enrichment results. I1: Biological processes. I2: Molecular function. I3: Cellular component. I4: Kyoto Encyclopedia of Genes and Genomes pathways. HR: Hazard ratio; AUC: Area under the curve.
- Citation: Zhang HL, Zhao R, Wang D, Mohd Sapudin SN, Yahaya BH, Harun MSR, Zhang ZW, Song ZJ, Liu YT, Doblin S, Lu P. Candida albicans and colorectal cancer: A paradoxical role revealed through metabolite profiling and prognostic modeling. World J Clin Oncol 2025; 16(4): 104182
- URL: https://www.wjgnet.com/2218-4333/full/v16/i4/104182.htm
- DOI: https://dx.doi.org/10.5306/wjco.v16.i4.104182