Copyright
©The Author(s) 2025.
World J Hepatol. Feb 27, 2025; 17(2): 99046
Published online Feb 27, 2025. doi: 10.4254/wjh.v17.i2.99046
Published online Feb 27, 2025. doi: 10.4254/wjh.v17.i2.99046
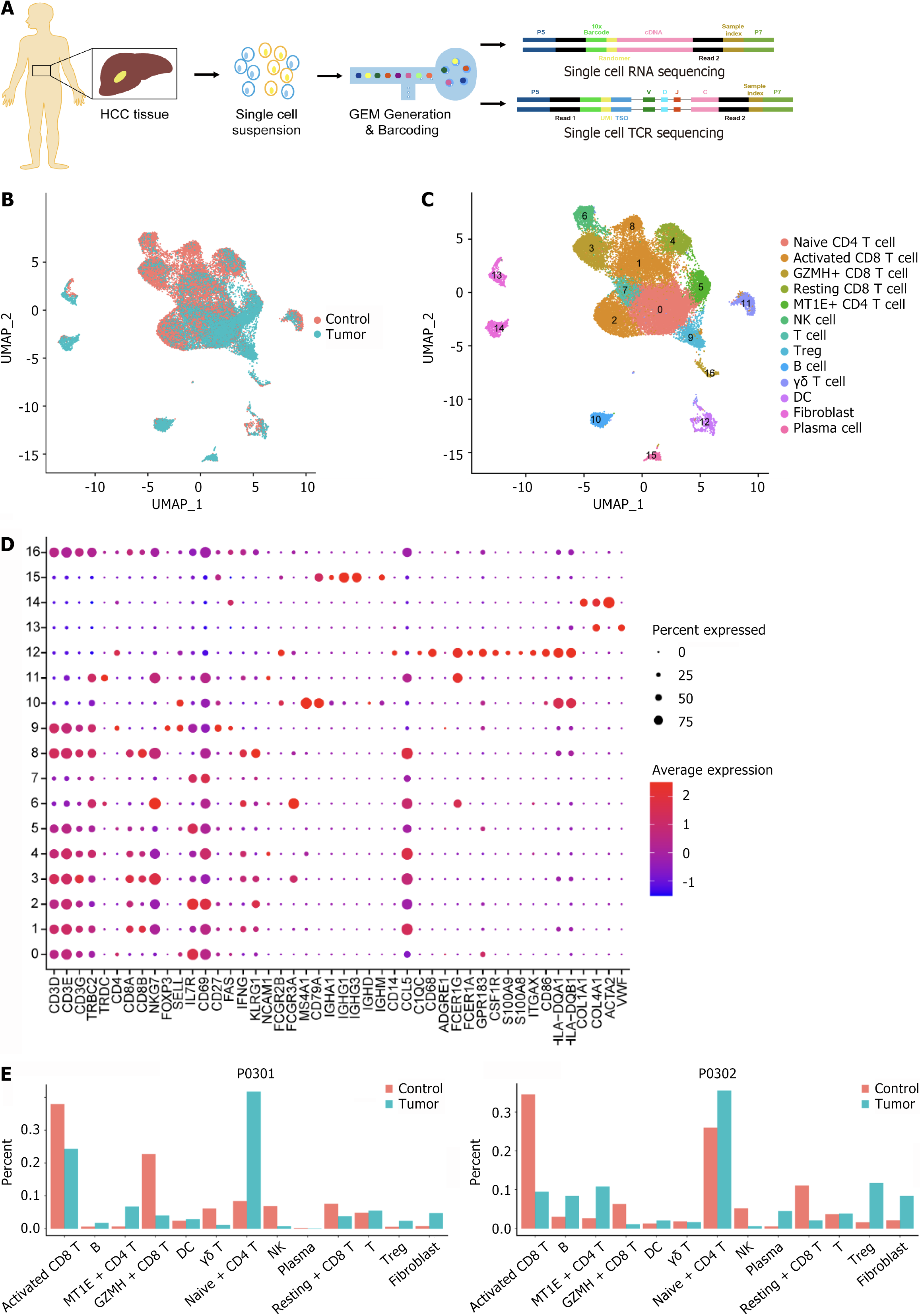
Figure 1 Single-cell transcriptome clustering of all cells from hepatocellular carcinoma patients.
A: Schematic diagram of the single-cell RNA sequencing experimental design; B: Uniform manifold approximation and projections of the combined single-cell RNA sequencing profiles of immune cells from control and tumor samples are plotted and colored according to samples; C: Cells are plotted and colored according to cell clusters. Each point corresponds to a single cell colored according to the cell cluster; D: A bubble plot of typical marker genes in each cluster is displayed. Darker colors indicate higher average expression; larger bubbles indicate a greater percentage of genes; E: Histogram of the abundance distribution of each cell subpopulation in the hepatocellular carcinoma and control groups. HCC: Hepatocellular carcinoma; TCR: T-cell receptor; UMAP: Uniform manifold approximation and projections; GZMH: Granzyme H; MT1E: Metallothionein 1E; NK: Natural killer; Treg: Regulatory T cells; DC: Dendritic cell.
- Citation: Gu XY, Gu SL, Chen ZY, Tong JL, Li XY, Dong H, Zhang CY, Qian WX, Ma XC, Yi CH, Yi YX. Uncovering immune cell heterogeneity in hepatocellular carcinoma by combining single-cell RNA sequencing with T-cell receptor sequencing. World J Hepatol 2025; 17(2): 99046
- URL: https://www.wjgnet.com/1948-5182/full/v17/i2/99046.htm
- DOI: https://dx.doi.org/10.4254/wjh.v17.i2.99046