Copyright
©2011 Baishideng Publishing Group Co.
World J Gastroenterol. Apr 7, 2011; 17(13): 1772-1778
Published online Apr 7, 2011. doi: 10.3748/wjg.v17.i13.1772
Published online Apr 7, 2011. doi: 10.3748/wjg.v17.i13.1772
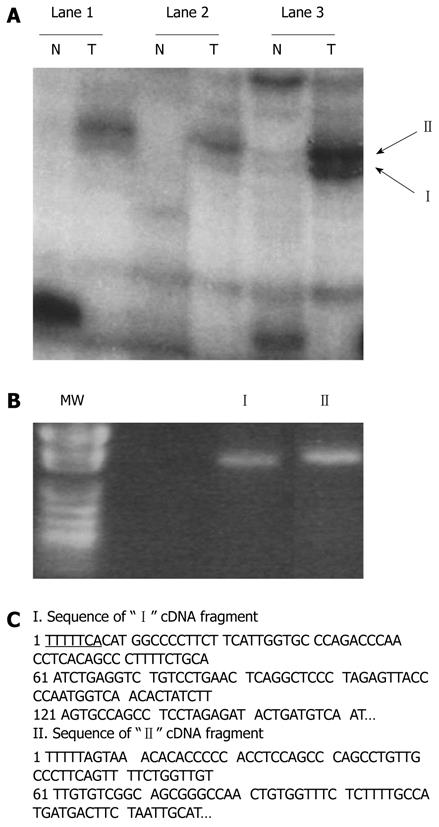
Figure 1 Identification of differentially displayed cDNA fragments from paired hepatocellular carcinoma cells (T) and nontumorous liver tissues (N).
A: Differentially displayed PCR products (“I” and “II”) amplified with AP-10 primer and T12MA. Lanes 1-3 denote the three B6C3 F1 mouse samples, respectively; B: Recovered “I” and “II” from the dried DNA sequencing gel reamplified by PCR. MW lane shows the pUC118 DNA fragments cut by HapII (a restriction enzyme) as molecular weight markers; C: Nucleotide sequences of the bands shown in A. Flanking sequences of T12MA primers are underlined. The two insert-containing fragments were sequenced and identified as gene fragments of GLNS and GSTM3 in comparison with those of nucleotide in GenBank.
- Citation: Li YS, Liu M, Nakata Y, Tang HB. β-catenin accumulation in nuclei of hepatocellular carcinoma cells up-regulates glutathione-s-transferase M3 mRNA. World J Gastroenterol 2011; 17(13): 1772-1778
- URL: https://www.wjgnet.com/1007-9327/full/v17/i13/1772.htm
- DOI: https://dx.doi.org/10.3748/wjg.v17.i13.1772